Introduction
Fall armyworm (FAW), Spodoptera frugiperda (J. E. Smith) (Lepidoptera: Noctuidae), is a major defoliating pest in the Americas and has recently become an economically devastating, invasive pest in Sub-Saharan Africa (Sparks, 1986; Goergen et al., 2016), as well as in Asia. FAW feeds on more than 80 economically important crops, including peanut. A thirteen billion dollar loss to FAW infestation was reported for maize, sorghum, rice, and sugarcane in Africa from 2016 to 2017 (Abrahams et al., 2017). Without control, FAW is a major defoliator that can cause high yield reductions, seriously affecting production in developing countries with limited access to pesticides and safety equipment to apply them properly (Cock et al., 2017; Day et al., 2017; Bateman et al., 2018; CABI, 2020). Producing FAW resistant cultivars will reduce reliance on pesticides, increase food security, and promote sustainable agriculture in these at-risk countries. In addition, resistant cultivars are needed to mitigate the insecticide resistance reported in FAW populations (Yu, 1991).
Genetic sources with strong FAW resistance are limited in cultivated peanut due to its narrow genetic base (Stalker et al., 2016). Peanut is an allotetraploid species (AABB; 2n=4x=40), and evidence suggests that it arose from a natural polyploidization event involving hybridization of the diploid species A. ipaensis (BB; 2n=2x=20) and A. duranensis (AA; 2n=2x=20) (Bertioli et al., 2016). The difference in ploidy levels between cultivated peanut and its wild relatives results in crossing barriers and constricts genetic diversity in peanut. Moderate levels of host resistance to FAW have been found in a few peanut cultivars such as ‘Southeastern Runner 56-15′ (Hammons 1970, Leuck and Skinner 1971), ‘Florunner’ and ‘Tifton 8′ (Todd et al., 1991). On the other hand, more than 80 wild peanut relatives have diverse levels of resistance against a wide range of peanut insect pests and diseases (Stalker et al., 2016; Stalker, 2017). Wild peanut relatives, such as Arachis cardenasii, already have proved to be useful sources of resistance to multiple pests including late leaf spot, early leaf spot, root-knot nematode, Cylindrocladium black rot, and Sclerotinia blight (Simpson and Starr, 2001; Stalker and Mozingo, 2001; Gowda et al., 2002; Simpson et al., 2003; Tallury et al., 2014). Previous studies have identified wild peanut relatives such as A. cardenasii, A. correntina, and A. villosa as potential donors of strong FAW antibiosis and antixenosis (Painter, 1951; Kogan and Ortman, 1978; Lynch et al., 1981; Ortega et al., 2016). These Arachis species displayed antibiosis through high FAW mortality rates and inhibition of FAW development and displayed antixenosis due to FAW non-preference to feed on these materials (Lynch et al., 1981). In addition, A. ipaensis showed antibiosis through a high FAW mortality rate (Yang et al., 1993). However, further work has not resulted in FAW resistance being introgressed from these wild relative species into a cultivated background useful for peanut breeders. This study identified FAW resistance in newly created allotetraploids that are cross compatible to cultivated, allotetraploid peanut. Resistant allotetraploid lines identified in this study can be used in breeding programs in the United States and shared with breeding programs in East and West Africa to introgress FAW resistance into elite cultivars. The long-term goal of this study is to create FAW resistant peanut cultivars that can protect yields in the United States and increase yields in regions with limited access to pesticides in Africa.
Materials and Methods
Plant Materials
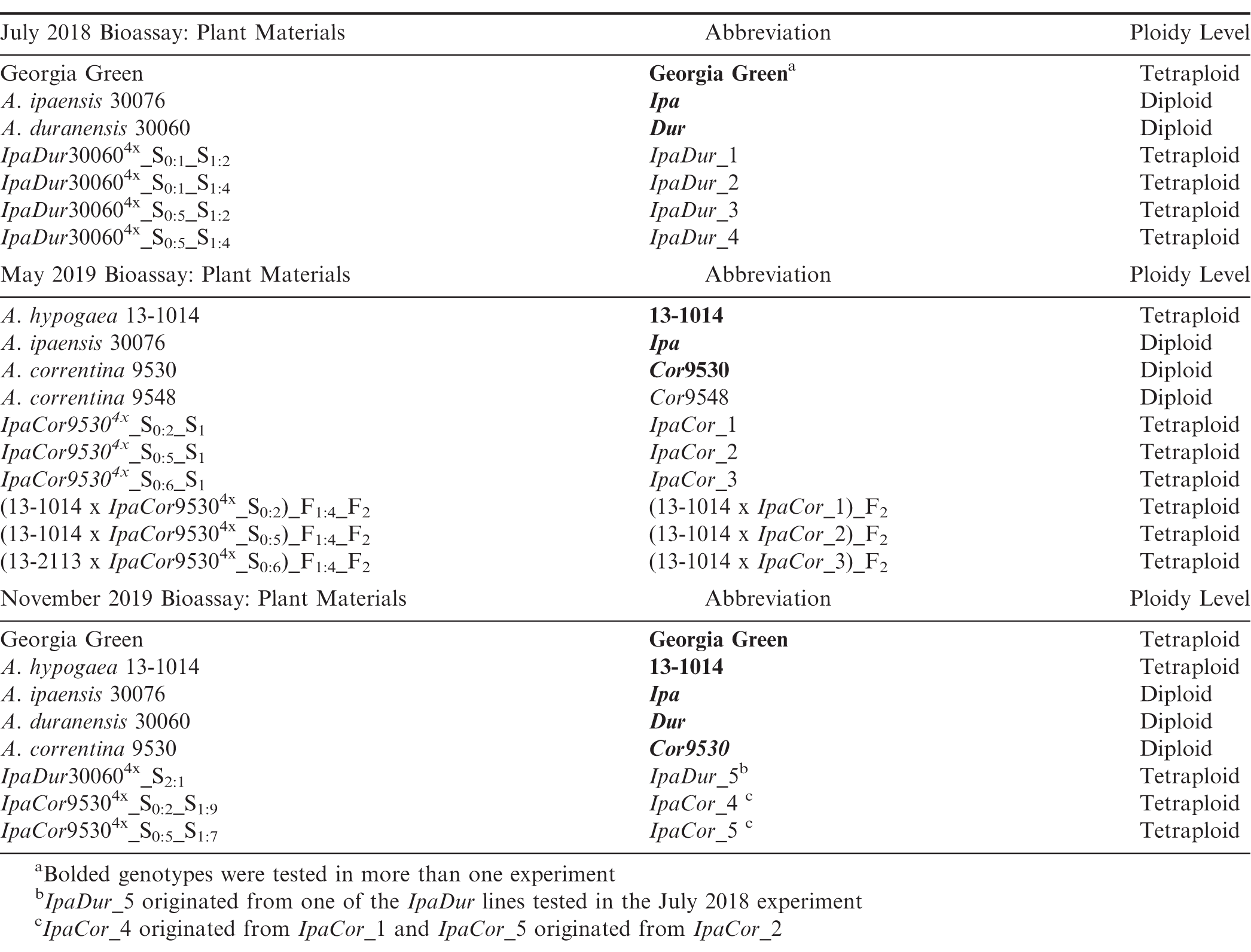
Wild Arachis species, A. ipaensis Krapov. and W.C. Gregory (PI 468322, GKBSPSc 30076; abbrev.: Ipa), A. duranensis Krapov. and W.C. Gregory (PI 468197, GKBSPSc 30060; abbrev.: Dur), A. correntina (Burkart) Krapov. and W.C. Gregory (PI 262808, GKP 9530; abbrev.: Cor9530) and (PI 262881, GKP 9548; abbrev.: Cor9548) were grown and the diploid hybrids generated in 2016 at North Carolina State University (NCSU). IpaDur4x and IpaCor95304x allotetraploids were created from the diploid hybrids by colchicine treatment of F1 hybrid cuttings at the University of Georgia (UGA) Tifton Campus. The susceptible A. hypogaea controls included ‘Georgia Green’ (Branch, 1996) and ‘13-1014,’ a breeding line selected from [(C1805-617-2 x ‘Florida-07′ (Gorbet and Tillman, 2009)) x ‘Georgia-06G’ (Branch, 2007)]. C1805-617-2 is a selection from ‘Tifguard’ (Holbrook et al., 2008) x ‘Florida-07′.
Due to limitations of labor and pesticide-free plants required for bioassay, the resistance evaluation was performed in three experiments with Ipa and allotetraploids with Ipa as one parent being tested in all of the experiments (Table 1). The first experiment, aimed to test the level of resistance in IpaDur4x, was conducted in July 2018 and included seven genotypes as treatments: Ipa, Dur, Georgia Green, and four S2 generation IpaDur4x plants. The allotetraploids were each designated an arbitrary number to make distinguishing them easier (Table 1). IpaDur_1 and IpaDur_2 originated from the same IpaDur_S0:1 plant, while IpaDur_3 and IpaDur_ 4 originated from the same IpaDur_S0:5 plant. The second experiment was designed to test the level of resistance in IpaCor95304x; it was carried out in May 2019 and included 10 genotypes as treatments: Ipa, Cor9530, Cor9548, A. hypogaea 13-1014, three S1 generation IpaCor95304x plants, and three F1 plants made from a cross between A. hypogaea 13-1014 and three different S0IpaCor95304x plants (Table 1). Cor (PI 261870) was previously reported to have FAW resistance (Lynch et al., 1981), but was not included in this study due to lack of seed supply. The third experiment was conducted in November 2019 to directly compare the level of resistance detected in materials from the first two experiments. The third experiment included eight genotypes as treatments: Ipa, Dur, Cor9530, Georgia Green, A. hypogaea 13-1014, one IpaDur4x S2 plant, and two IpaCor95304x S1 plants (Table 1). IpaDur_5 was an S2 plant that originated from one of the IpaDur4x lines tested in the July 2018 experiment. IpaCor_4 originated from IpaCor_1 tested in the May 2019 experiment, while IpaCor_5 originated from IpaCor_2. In this last experiment, four genotypes were terminated early because the plants grew slowly in the short day length season and failed to produce enough leaves to complete the test. The earliest date of termination was 17 DAI (days after infestation) for genotype Dur.
Fall Armyworm Resistance Evaluation
For each of the three experiments, seeds were coated in Vitavax PC (Vitavax, Crompton, Middlebury, CT) and treated overnight in 0.5% Florel Growth Regulator (Lawn and Garden Products Inc., Fresno, CA) to break dormancy. Seeds were then planted in #123 7.62 cm round x 11.43 cm deep Jiffy Pots (Harris Seeds, Rochester, NY) and transplanted approximately one month later into 121.92 cm round x 27.94 cm deep pots filled with Promix growth medium (Premier Tech Horticulture, Quakertown, PA). Normal plant management was applied in the greenhouse except that insecticide treatments were withheld.
When plants were three months old, FAW eggs (corn strain) were obtained from USDA-ARS Corn Host Plant Resistance Research Unit, Mississippi State, MS. Thirty replications were tested for each genotype, in which one replication comprised one FAW larva fed on the excised leaflets from a genotype. Within 24 h of egg hatch, the neonate larvae were gently transferred with a soft tip paintbrush from the bag of FAW egg masses onto a fresh, newly emerged leaflet placed in a 100 mm x 15 mm Petri dish (ThermoFisher Scientific). Each Petri dish contained a sheet of 9 cm diameter Whatman No. 1 filter paper (ThermoFisher Scientific) supported by a cotton round (Equate Beauty) that was saturated with approximately 4 ml of deionized water. The day that the neonate larvae were transferred onto the leaflets was considered day 0 after infestation (0 DAI). Petri dishes were sealed with Parafilm® M laboratory film (Bemis Company, Inc.) for the first week to prevent the FAW from escaping. For the next three days, Petri dishes with dead FAW larva were replenished with another neonate larva. On the day a Petri dish was restarted with a new FAW larva, that day was considered 0 DAI for that particular plate. The Petri dishes were examined daily to relocate FAW larvae that may have moved off of the leaflets and onto or underneath the filter paper. Daily inspection ensured that FAW larvae would have the opportunity to feed on plant material, and that the filter paper and cotton pad could be moistened if they had become dry. Fresh leaflets were offered every other day for one week, then fresh leaves were offered daily to meet the need of increased larval feeding. The moistened cotton pad and filter paper were changed as the frass from FAW larva accumulated and contaminated the filter paper.
The following parameters were evaluated to study the effect of plant genotype on FAW growth and development: survival, larval weight, larval stage duration, pupation, pupal stage duration, moth emergence relative to pupation, and moth sex. Low survival, larval weight, pupation, and emergence as well as high larval stage duration and pupal stage duration indicated deceleration of normal FAW growth and development. Sex ratio was an indicator of genotype effect on FAW fitness in which a deviation from a 1:1 (male to female) indicated a genotypic effect of the host plants on FAW sexual dimorphism. Sex ratio was included, since diet and other environmental effects have been found to negatively affect adult insect, including FAW, sex ratios (Bull, 1981; Murúa and Virla, 2004). Biased sex ratio could be utilized as one of the valuable tactics for sustained pest management in multiple cropping systems. FAW survival was documented daily. Larval weight was recorded at 14 DAI. Larval stage duration documented the number of days from first-instar larva stage to pupal formation. Pupation and relative moth emergence described completed pupation and moth emergence relative to the number of pupa formed, respectively. Pupal stage duration denoted the number of days recorded from pupal development to moth emergence. Pupal weight was recorded on the day of complete pupal formation. Moth sex (male or female) was determined after emergence.
Genotyping the diploid species
Cor demonstrated morphological diversity; therefore, we genotyped the diploid species using the Affymetrix Axiom_Arachis2 SNP array (Clevenger et al., 2018; Korani et al. 2019) consisting of 47,000 features (ThermoFisher Scientific). DNA was extracted by the Qiagen DNeasy Plant mini kit (Qiagen, Germantown, MD). SNP calling was performed with Axiom Analysis Suite (Version 1.2). Genetic markers were grouped into six categories by the software depending on the quality and separation of markers 1) Monomorphic 2) PolyHighResolution 3) NoMinorHom 4) OfftargetVariant 5) CallRateBelowThreshold 6) Other. Only the 5,342 markers in the PolyHighResolution class were used for analysis since the grouping of samples was unambiguous and all the samples passed quality control.
Statistical Analysis
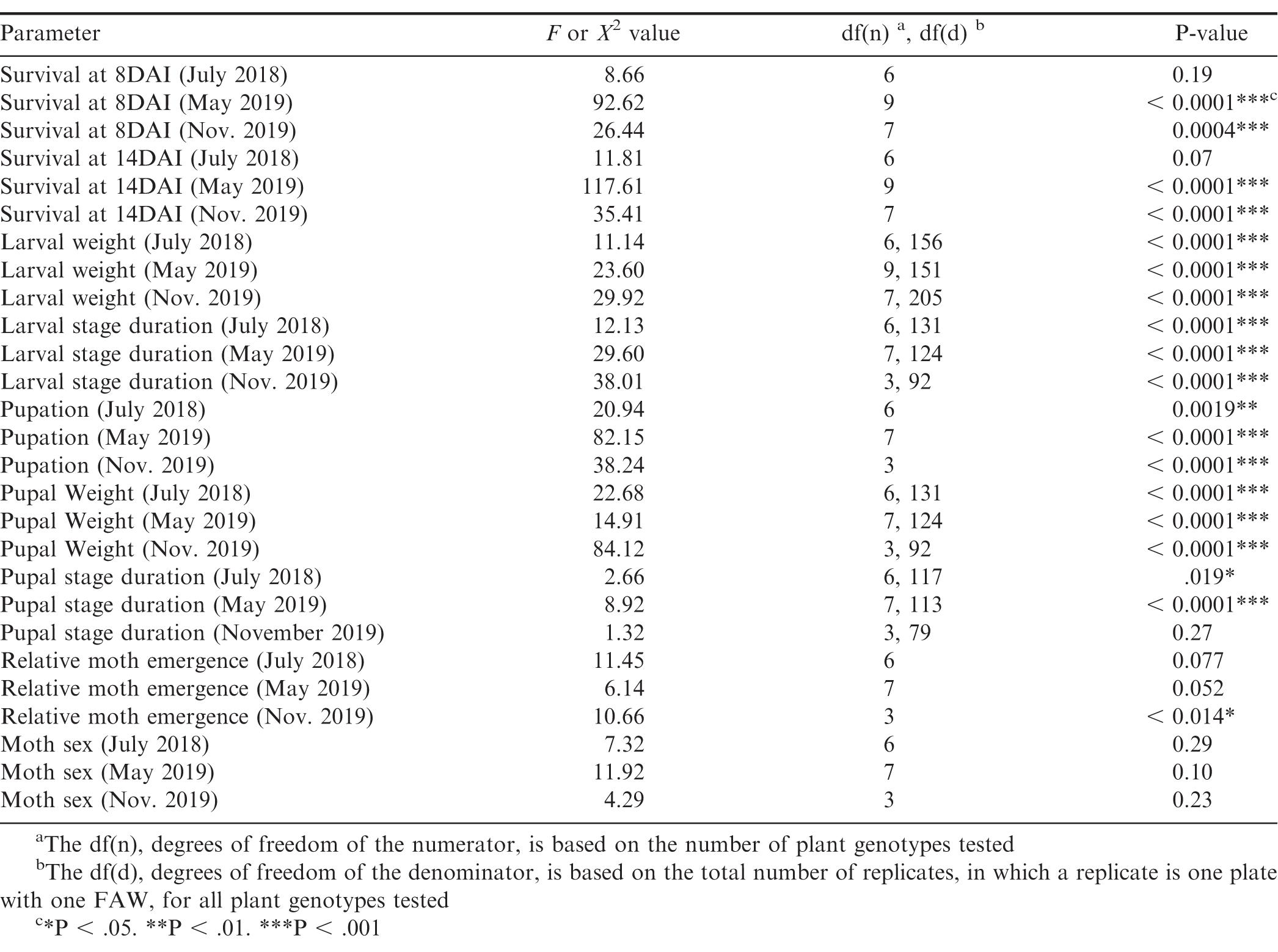
One-way analysis of variance (ANOVA) was performed using JMP software (SAS Institute) to determine the genotype effect on FAW growth and development according to the following parameters: larval weight, larval stage duration, pupal stage duration, and pupal weight. Chi-squared tests were performed with JMP software to determine the genotype effect on the following categorical parameters used to assess FAW development: survival at 8 DAI and 14 DAI (dead or alive), pupation (pupated or did not pupate), relative moth emergence (emerged or did not emerge), and moth sex (male or female). For relative moth emergence, data (emerged or did not emerge) was only included for FAW that pupated. Means of each parameter among the treatments were separated based on the Tukey’s Test (α = 0.05) results with JMP software. Tukey’s Test for binomial data was not supported with JMP software, so RStudio (RStudio, Inc.) was used by running genotype effect on survival at 8 DAI and 14 DAI, pupation and relative moth emergence as a generalized linear model and then Tukey’s Test (α = 0.05) was performed on the model results. Genotypes with less than four replications for a measurement were excluded from statistical analysis. Therefore, larval stage duration, pupation, pupal weight, pupal stage duration, relative moth emergence, and moth analysis only included eight genotypes for the May 2019 experiment with Ipa and IpaCor_3 excluded and only four genotypes for the November 2019 experiments with Cor, Dur, Ipa, and IpaDur_5 excluded due to being terminated before pupal formation. The data for both the May and November 2019 experiments were still valid, since they were maintained according to experimental design and the controls were continued.
Results
Survival
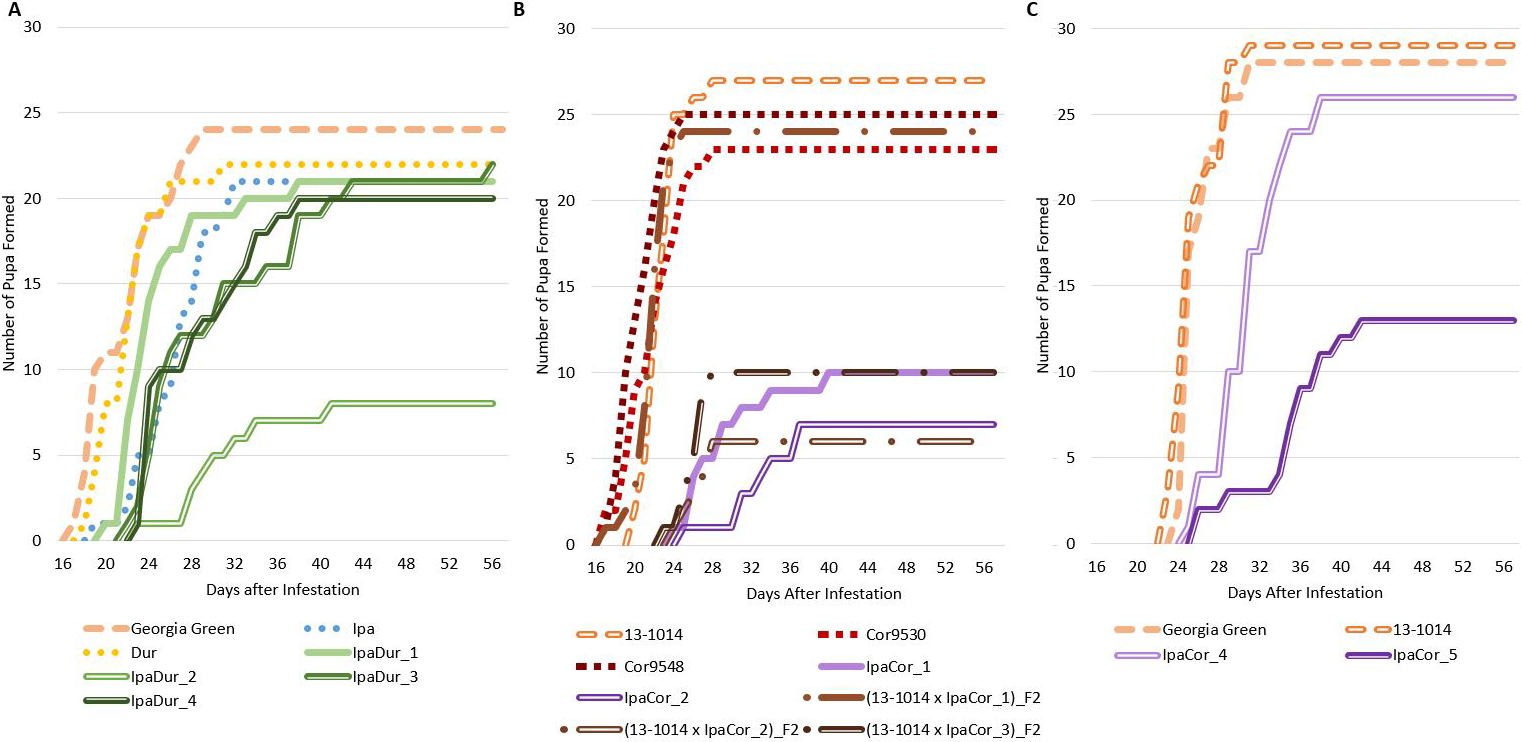
Significant genotypic effect on FAW survival at 8 DAI and 14 DAI was found for the May 2019 experiment and the November 2019 experiment, but not for the July 2018 experiment (Table 2). However, Tukey-Kramer significant differences between genotypes were not found for neither 8 DAI nor 14 DAI for all three experiments (Table S1). In all three experiments, the cultivated control genotypes had the highest FAW survival (Fig. 1). For the July 2018 experiment (Fig. 1A), the allotetraploid IpaDur_2 had the greatest FAW mortality. Only 30% (9 of the 30) of FAW fed exclusively on IpaDur_2 survived to the end of the experiment and completed their life cycle. The larval survival curves on Ipa and Dur were below that of the susceptible cultivated peanut control.
Unlike the other two experiments, the May 2019 experiment (Fig. 1B) had a high mortality of FAW larvae just four days after the onset of the experiment. This is likely due to a trained entomologist performing the July 2018 and November 2019 experiments. However, all allotetraploids had survival curves below the susceptible control. Both accessions of A. correntina were similar to the susceptible control, although Cor9530 lost five more FAW larvae (17% more) than Cor9548.
In the November 2019 experiment (Fig. 1C), Ipa, Dur, Cor, and IpaDur_5 had to be terminated early due to insufficient plant material to sustain the FAW feeding. However, FAW larvae fed on susceptible controls showed almost identical survival curves as the previous two experiments. The FAW larva survival curve on Cor9530 closely followed survival curves of the susceptible controls. IpaCor_4, a progeny line from IpaCor_1 that showed high FAW mortality in the May 2019 experiment (Fig. 1B), also had a survival curve similar to the controls. Ipa and Dur had similar survival curves to the July 2018 experiment (Fig. 1C) until the treatments were terminated. IpaCor_5 (Fig. 1C), a progeny line from IpaCor_2 (Fig. 1B) that had high FAW mortality in the May 2019 experiment, had the greatest FAW mortality in the November 2019 experiment.
Larval Weight
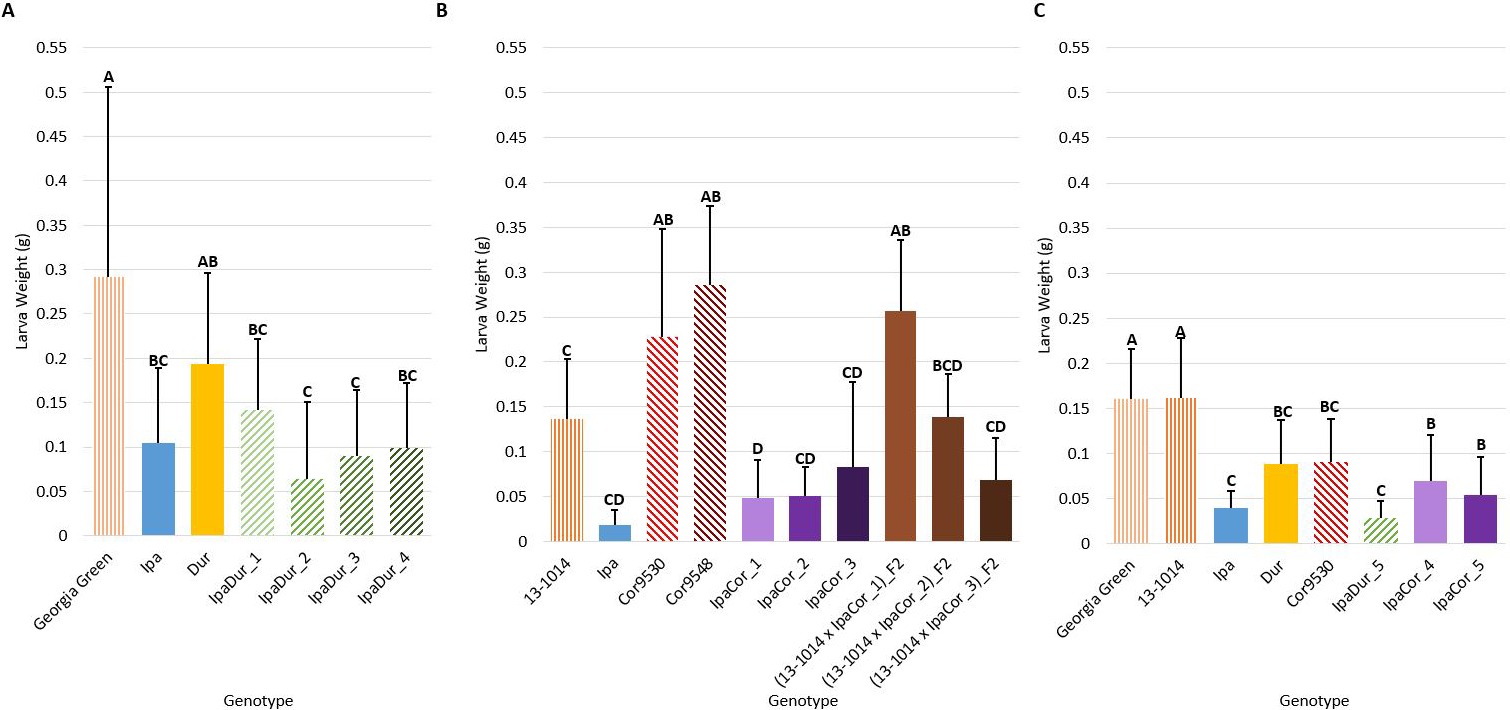
Genotype had a significant effect on larval weight in all of the experiments (Table 2). All five IpaDur4x allotetraploid plants significantly reduced FAW larval weight compared to the cultivated peanut genotypes included as experimental controls (Fig. 2). For the July 2018 experiment, the allotetraploid IpaDur_2 that had the greatest FAW mortality also had the numerically lowest larval weight (Figs. 1A, 2A). As for IpaCor95304x, three out of five tested allotetraploid plants (IpaCor_1, IpaCor_4, and IpaCor_5) significantly suppressed larval weight (Figs. 2B, C). The other two IpaCor95304x plants (IpaCor_2 and IpaCor_3) also reduced larval weight, yet the level of reduction did not show statistical significance when compared to the cultivated peanut controls (Fig. 2B). This is partly due to high mortality of FAW larvae in the May 2019 experiment (Fig. 1B), which led to small sample size, and affected statistical power for larval weight data (Fig. 2B). For example, while the larval weight on IpaCor_1 was 35% of the control and found to be significantly different, the Ipa larval weight was only 13.5% of the control but not significantly different due to the small sample size caused by mortality of FAW larvae on Ipa leaves. When the two allotetraploid lines were tested side-by-side in the November 2019 experiment (Fig. 2C), IpaDur_5 demonstrated a stronger level of FAW resistance by reducing FAW larval weight to a greater extent than IpaCor9530_4 and IpaCor9530_5.
Larva weight on day 14 after feeding on different peanut leaflets. (A) July 2018 Experiment. (B) May 2019 Experiment. (C) November 2019 Experiment. Error bars represent standard error. Tukey’s HSD significance levels were calculated within each experiment, so these significance groupings cannot be compared between the three experiments. Genotypes within an experiment with the same Tukey’s HSD letter are not significantly different (α = 0.05).
Diploid parents of both allotetraploid lines were included in this study to determine the source of FAW resistance. Compared to the cultivated peanut control, Ipa significantly suppressed FAW weight gain in the July 2018 and November 2019 experiments (Figs. 2A, C). Dur also significantly reduced the weight of FAW larvae but to a numerically lesser extent than that of Ipa in the November 2019 experiment. Conflicting results were found between the May 2019 (Fig. 2B) and November 2019 (Fig. 2C) experiments for Cor9530. A significant, positive effect on larval weight was observed in the May 2019 experiment (Fig. 2B) and a significant suppressive effect was found in the November 2019 experiment (Fig. 2C). Cor9548 was tested only in one experiment (Fig. 2B) and it was found to increase larval weight gain. A high level of genetic heterogeneity was found in Cor9530 (Table S2). Among the three genotyped Cor9530 plants, 1,259 out of 5,342 total markers (23.5%) were found to be polymorphic. On the contrary, only 23 (0.4%) and 27 (0.5%) polymorphic markers were found with Ipa and Dur, respectively. Therefore, the conflicting feeding response on Cor9530 could be due to the genetic diversity within this accession. In the May 2019 experiment (Fig. 2B), three 13-1014 x IpaCor F2 plants were found to be segregating for larval weight. The (13-1014 x IpaCor_1)_F2 plant demonstrated a significantly greater increase in larval weight compared to the cultivated control. The (13-1014 x IpaCor_2)_F2 plant also showed a similar effect on larval weight compared to the cultivated control. The (13-1014 x IpaCor_3)_F2 plant demonstrated high suppression of FAW growth similar to the IpaCor95304x.
Larval Stage Duration
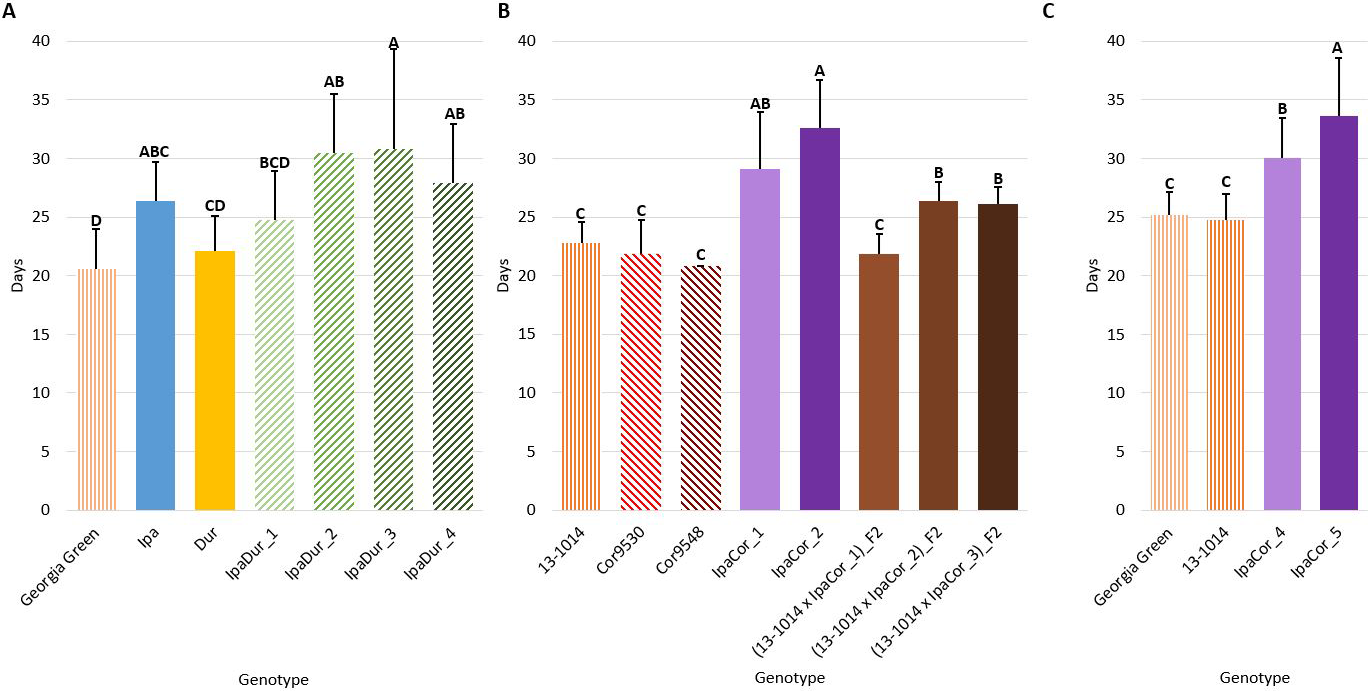
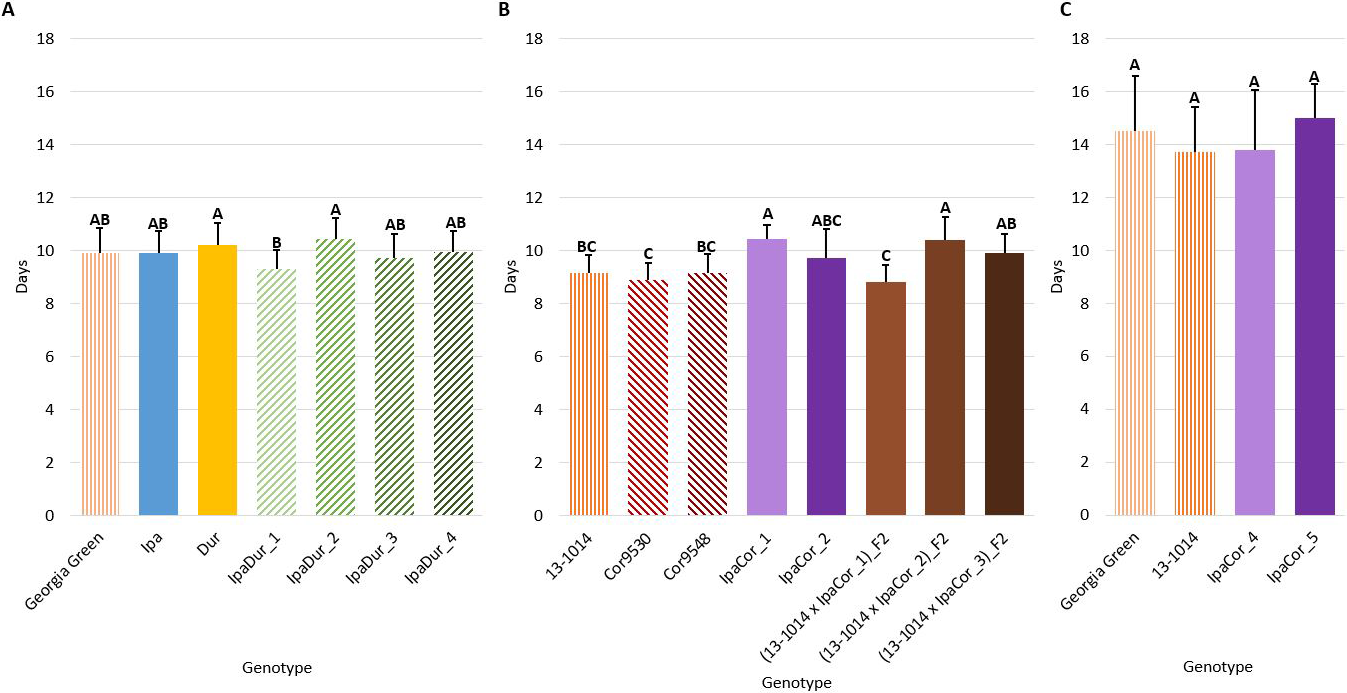
Genotype had a significant effect on larval stage duration in all of the experiments (Table 2). In all three experiments, the cultivated controls had the shortest larval stage duration (Fig. 3). Compared to the cultivated control, Ipa significantly reduced larval growth and development by extending larval stage duration in the July 2018 experiment (Fig. 3A). Likewise, all allotetraploids significantly extended larval stage duration, except for IpaDur_1 in the July 2018 experiment (Fig. 3A). IpaCor_5, the allotetraploid with the greatest FAW mortality in the November 2019 experiment (Fig. 1B), significantly outperformed IpaCor_4 by greatly extending larval stage duration (Fig. 4B). While Ipa significantly extended larval stage duration, Dur, Cor9530, and Cor9548 did not differ significantly from the cultivated controls.
Larval stage duration. (A) July 2018 Experiment. (B) May 2019 Experiment. (C) November 2019 Experiment. Error bars represent standard error. Tukey’s HSD significance levels were calculated within each experiment, so these significance groupings cannot be compared between the three experiments. Genotypes within an experiment with the same Tukey’s HSD letter are not significantly different (α = 0.05).
Pupation
Genotype had a significant effect on pupation in all of the experiments (Table 2). For the July 2018 and May 2019 experiments, Tukey-Kramer significant differences between genotypes were not found; however, 13-1014 had significantly more pupated FAW than IpaCor_5 (Table S1). Across all the experiments, the cultivated controls had the highest number of FAW to pupate (Fig. 4). For the July 2018 experiment, IpaDur_2, which had the highest FAW mortality (Fig. 1A), also had the fewest FAW to pupate (Fig. 4A). For the May 2019 experiment (Fig. 4B), pupation was heavily influenced by FAW mortality early in the experiment. Despite this artifact, all allotetraploids had a lower number of FAW to pupate than the cultivated control. Both Cor9530 and Cor9548 had a similar number of FAW to pupate as compared to the cultivated control. Line (13-1014 x IpaCor_1)_F2 also had a similar number of FAW to pupate as compared to the cultivated controls, while (13-1014 x IpaCor_2)_F2 and (13-1014 x IpaCor_3)_F2 had much lower numbers of FAW to pupate as compared to the cultivated controls. For the November 2019 experiment, IpaCor_4 had a similar number of FAW to pupate as the cultivated controls (Fig. 4C), while IpaCor_5 had significantly less pupa form as compared to 13-1014 (Table S1).
Pupal Weight
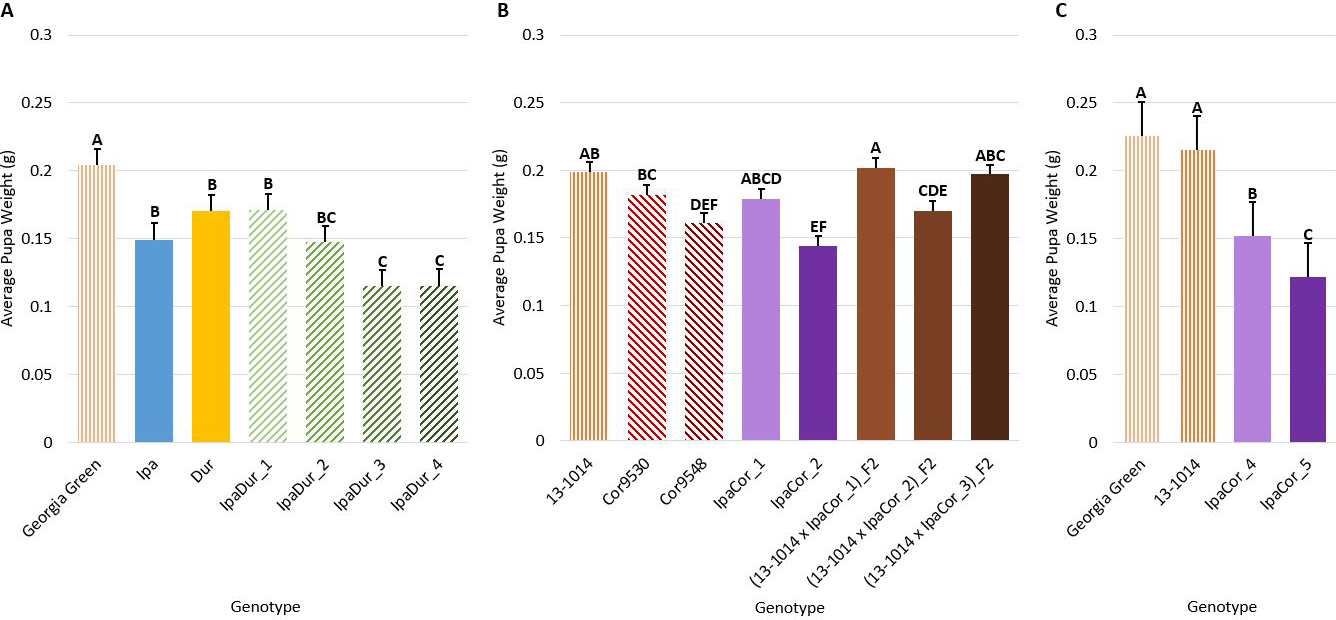
Genotype had a significant effect on average pupal weight across all experiments (Table 2). All allotetraploids, except IpaCor_1 in the May 2019 experiment, had significantly lower pupal weight than the cultivated controls (Fig. 5). IpaDur_3 and IpaDur_4 had significantly lower pupal weight than IpaDur_1 (Figs. 5A, C). IpaCor_2 had the lowest pupal weight of all the allotetraploids in the May 2019 experiment (Fig. 5B). IpaCor_5, the progeny line of IpaCor_2, had the lowest pupal weight in the November 2019 experiment (Fig. 5C). Ipa, Dur, and Cor9548 all had significantly lower pupal weight than the respective cultivated control. However, Cor9530 was not statistically different from the cultivated control in the May 2019 experiment (Fig. 5B).
Pupal weight. (A) July 2018 Experiment. (B) May 2019 Experiment. (C) November 2019 Experiment. Error bars represent standard error. Tukey’s HSD significance levels were calculated within each experiment, so these significance groupings cannot be compared between the three experiments. Genotypes within an experiment with the same Tukey’s HSD letter are not significantly different (α = 0.05).
Pupal Stage Duration
Genotype had a significant effect on pupal stage duration in the July 2018 experiment and the May 2019 experiment, but not in the November 2019 experiment (Table 2). Across all three experiments, Ipa, Dur, Cor9530, and Cor9548 were not significantly different from the cultivated controls (Fig. 6). However, IpaCor_1 and (13-1014 x IpaCor_2)_F2 had significantly longer pupal stage duration than the cultivated control, indicating these materials impeded pupal stage development (Fig. 6B).
Pupal stage duration. (A) July 2018 Experiment. (B) May 2019 Experiment. (C) November 2019 Experiment. Error bars represent standard error. Tukey’s HSD significance levels were calculated within each experiment, so these significance groupings cannot be compared between the three experiments. Genotypes within an experiment with the same Tukey’s HSD letter are not significantly different (α = 0.05).
Relative Moth Emergence
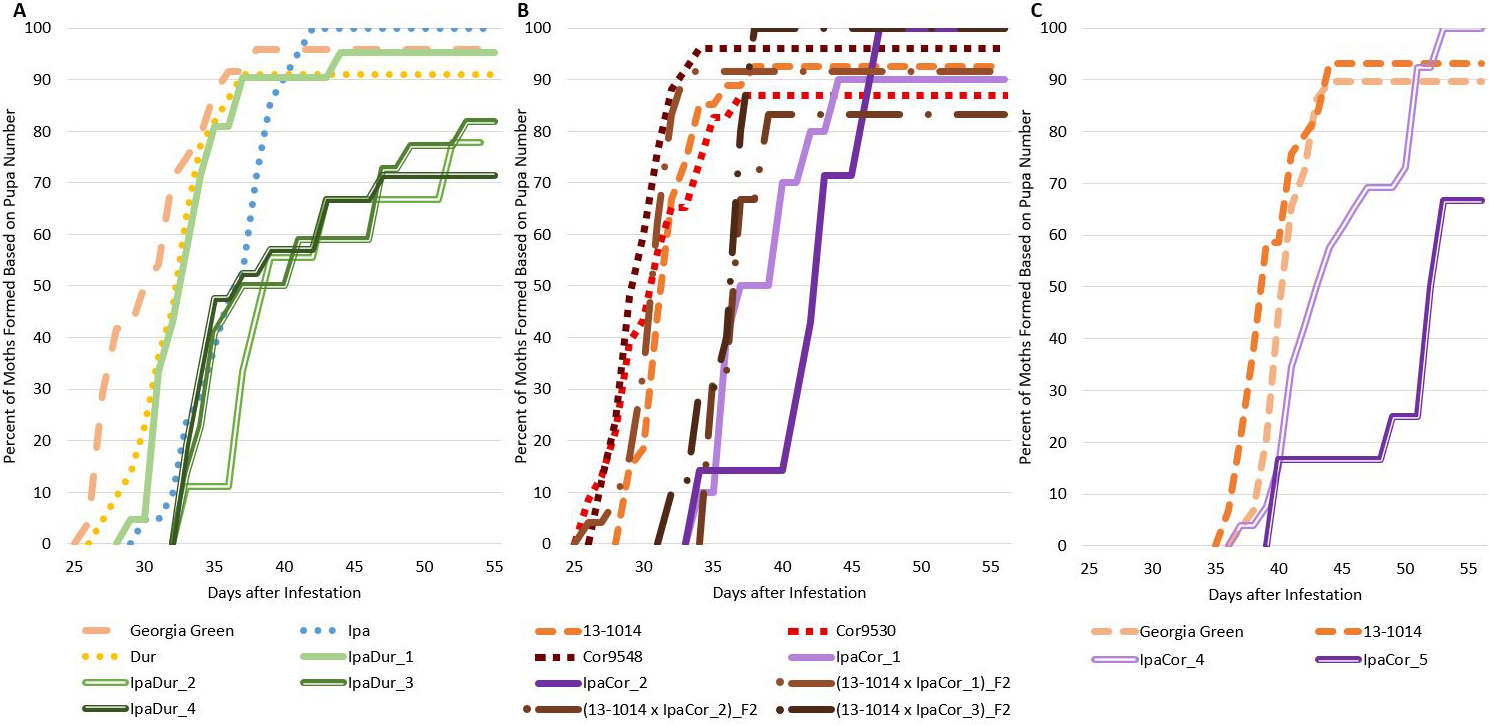
Genotype had a significant effect on relative moth emergence in the November 2019 experiment but not the July 2018 or May 2019 experiments (Table 2), and Tukey-Kramer significant differences between genotypes for relative moth emergence were not found for any of the three experiments (Table S1). All of the allotetraploids, except for IpaCor_2 and IpaCor_4 (Fig. 7B, C), supported less relative moth emergence than the cultivated controls (Fig. 7). Allotetraploids had a high number of aborted pupae with the highest being 38.5% (5 out of 13 pupa) for IpaCor_5, then 25% (5 out of 20 pupa) for IpaDur_4, and 18.9% (4 out of 22) for IpaDur_3 (Fig. 7A, C). In comparison, Georgia Green had 4.2% (1 out of 24) and 7.1% (2 out of 28) aborted pupa in the July 2018 and November 2019 experiments, respectively (Fig. 7A, C). Arachis hypogea 13-1014 had 7.1% (2 out of 28) and 6.9% (2 out of 29) aborted pupa in the July 2018 and November 2019 experiments, respectively (Fig. 7A, C). The overall low number of 8 moths formed on IpaCor_5, a progeny line of IpaCor_2, was due to early FAW mortality and high pupa abortion (Figs. 1B, 7B). All larvae fed on Ipa, Dur, Cor9530, and Cor9548 showed similar relative moth emergence as compared to the cultivated controls. Also, larvae fed on Cor9530 showed similar relative moth emergence as compared to the larvae fed on Cor9548 (Fig. 7B). Lastly, the (13-1014 x IpaCor)_F2 plants all had similar relative moth emergence when compared to the control (Fig. 7B).
Moth Sex
Genotype did not have a significant effect on moth sex in the July 2018, May 2019, nor November 2019 experiment (Table 2).
Discussion
The wild peanut-derived allotetraploid genotypes showed promise as sources for FAW resistance for peanut breeding programs, since all IpaDur4x and IpaCor95304x allotetraploids significantly reduced larval weight (except IpaCor_2 and IpaCor_3), increased pupal stage duration (except IpaDur_1), and decreased pupal weight (except IpaCor_1) as compared to the controls. All allotetraploid genotypes showed lower survival and pupation than the cultivated genotypes evaluated as well as lower relative moth emergence (except IpaCor_2 and IpaCor_4). The most promising allotetraploid line, IpaDur_2, had the lowest larval weight and the greatest FAW mortality in the July 2018 experiment. This was further supported by IpaDur_5, which significantly reduced larval and pupal weight in the November 2019 experiment. IpaCor_2 and its progeny IpaCor_5 were shown as a promising line as they both suppressed FAW growth in the May and November 2019 experiments. Since segregation for FAW resistance in the allotetraploids was found, further selections for FAW resistance in individual lines will be necessary for effective utilization of these materials in a breeding program.
Primarily Ipa, but also Dur, were shown to be donor sources for FAW resistance, by reducing larval and pupal weight, survival and pupation. The conclusion that Ipa is valuable as a source for FAW resistance confirms a previous report (Yang et al., 1993). However, a 79% mortality rate at 8 DAI was reported for A. ipaensis in a similar bioassay with five wild Arachis species and A. hypogaea (Yang et al., 1993). The bioassay did differ in the use of terminal buds from field-grown plants rather than expanded leaves from greenhouse-grown plants. Two of our three experiments showed generally higher survival for Ipa with 8 DAI mortality rates of 16.7%, 80%, and 20% for the July 2018, May 2019, and November 2019 experiments, respectively. The high mortality of FAW fed on Ipa leaves in the May 2019 experiment may be an artifact of this experiment having overall higher FAW mortality than the other two experiments (Fig. 1), influenced by a trained entomologist performing the July 2018 and November 2019 experiments. Overall, the mean Ipa mortality rate was 38.9% across the three experiments. However, the low larval weight of Ipa supports the previous conclusions (Yang et al., 1993). Another FAW bioassay study (Lynch et al., 1981) tested 14 wild Arachis species, not including Ipa and Dur, and recommended Cor (PI 261870) as a source for FAW resistance. Cor (PI 261870) impeded FAW survival and development and also showed antixenosis in a free-choice preference test. We tested Cor9530 and Cor9548 instead of Cor (PI 261870), because Cor (PI 261870) seeds were not available. Unlike the previously reported results, conflicting results were found regarding the FAW resistance in Cor9530 plants tested in these experiments. Genotyping detected a high level of SNP variation within Cor9530, suggesting this diploid species accession is highly heterogeneous which may explain the contradictory bioassay results of FAW resistance within this species. In addition, IpaCor_1, IpaCor_2, and IpaCor_3 were derived from crosses between Ipa and different Cor9530 individuals, so genetic difference between these lines can be ascribed to parental heterogeneity or genetic segregation in the allotetraploids.
This study builds upon previous reports by focusing on allotetraploids that are cross compatible to A. hypogaea and are therefore valuable to breeding programs, instead of focusing solely on diploid, wild Arachis species. In the May 2019 experiment, varied levels of FAW resistance were detected among the three tested F2 lines from 13-1014 x IpaCor95304x cross. The (13-1014 x IpaCor_1)_F2 plant had an even greater larval weight than A. hypogaea 13-1014, while the (13-1014 x IpaCor_3)_F2 plant was similar to the allotetraploids, suggesting the FAW resistance from IpaCor95304x segregated in the F2 populations. An expanded study of a population derived from the tested (13-1014 x IpaCor)_F2 materials is needed to determine the inheritance pattern and conduct genetic mapping of FAW resistance QTL.
The major limitation of this study is that all three experiments were confined to in vitro bioassays using excised peanut leaflets. Due to the required labor for these bioassays that included daily examination of each plate for 44 days per experiment, only a limited number of genotypes could be tested. A progression to this study would be to confirm these results in a field trial and then to map FAW resistance QTL in a mapping population. However, it is difficult to maintain a peanut field with FAW infestation in the southeastern U.S. because of the abundance of natural insect enemies in the peanut fields throughout the growing season and the sporadic nature of FAW as a pest on peanut plants. However, a field study could be performed in collaboration with partners in Africa, where FAW is a newly invasive pest and FAW pressures are high.
Conclusions
This study built upon previous reports by testing FAW resistance in the unique allotetraploids instead of just wild, diploid Arachis species alone. The allotetraploids examined in this study showed strong FAW resistance, making them more useful than peanut cultivars previously found to have only moderate levels of FAW resistance. Furthermore, these allotetraploids are cross-compatible with peanut cultivars, making this resistance accessible for peanut breeders. These FAW resistant allotetraploids will be shared with breeding programs in the United States and Africa so that FAW resistance can be introgressed into elite peanut cultivars to effectively reduce the impact of the FAW (as an invasive pest) on peanut production in countries where growers have limited access to pesticides. At the same time, FAW resistant peanut cultivars can protect yields in the United States, increase yields in regions with limited use of pesticides, decrease reliance on pesticides, and promote organic peanut production and sustainable agriculture in general.
Acknowledgements
This work was supported by the National Science Foundation (grant # MCB-1543922) and by the AFRI NIFA Fellowships Grant Program: Predoctoral Fellowships project accession no. 1019105 from the USDA National Institute of Food and Agriculture. The authors would like to express their appreciation to Susan Wolf of USDA-ARS, Mississippi State, Mississippi, for providing the FAW eggs and their appreciation to Micah Levinson of the Department of Horticulture, University of Georgia, Georgia, for assisting in data collection.
Literature Cited
Abrahams, P., Beale, T., Cock, M., Corniani, N., Day, R., Godwin, J., Murphy, S., Richards, G., and Bos. J. 2017. Fall armyworm status, Impacts and control options in Africa: Preliminary evidence note. United Kingdom’s Department for International Development. Retrieved 21 Dec 2019, from https://www.cabi.org/isc/FullTextPDF/2018/20187200430.pdf
Bateman, M.L., Day, R.K., Luke, B., Edgington, S., Kuhlmann, U., and Cock. M.J.W. 2018. Assessment of potential biopesticide options for managing fall armyworm (Spodoptera frugiperda) in Africa. J. Appl. Entomo. 142: 805– 819.
Bertioli, D.J., Cannon, S.B., Froenicke, L., Huang, G., Farmer, A.D., Cannon, E.K., Liu, X., Gao, D., Clevenger, J., Dash, S., Ren, L., Moretzsohn, M.C., Shirasawa, K., Huang, W., Vidigal, B., Abernathy, B., Chu, Y., Niederhuth, C.E., Umale, P., Araújo, A.C., Kozik, A., DoKim, K. Burow, M.D., Varshney, R.K., Wang, X., Zhang, X., Barkley, N., Guimarães, P.M., Isobe, S., Guo, B., Liao, B., Stalker, H.T., Schmitz, R.J., Scheffler, B.E., Leal-Bertioli, S.C., Xun, X., Jackson, S.A., Michelmore, R., and Ozias-Akins. P. 2016. The genome sequences of Arachis duranensis and Arachis ipaensis, the diploid ancestors of cultivated peanut. Nat. Genet. 48: 438– 446.
Branch, W.D. 1996. Registration of ’Georgia Green’ peanut. Crop Sci. 36: 806.
Branch, W.D. 2007. Registration of ’Georgia-06G’ peanut. J. Plant Regist. 1: 120.
Bull, J.J. 1981. Sex ratio evolution when fitness varies. Heredity 46(1): 9– 26.
CABI. 2020. Invasive Species Compendium: Spodoptera frugiperda (fall armyworm). Retrieved 21 Dec 2019, from http://www.cabi.org/isc/datasheet/29810
Clevenger, J.P., Korani, W., Ozias-Akins, P., and Jackson. S. 2018. Haplotype-based genotyping in polyploids. Frontiers in Plant Science 9: 564.
Cock, M.J.W., Beseh, P.K., Buddie, A.G., Cafá, G., and Crozier. J. 2017. Molecular methods to detect Spodoptera frugiperda in Ghana, and implications for monitoring the spread of invasive species in developing countries. Sci. Rep. 7(1): 4103.
Day, R., Abrahams, P., Bateman, M., Beale, T., Clottey, V., Cock, M., Colmenarez, Y., Corniani, N., Early, R., Godwin, J., Gomez, J., Moreno, P.G., Murphy, S.T., Oppong-Mensah, B., Phiri, N., Pratt, C., Silvestri, S., and Witt. A. 2017. Fall armyworm: Impacts and implications for Africa. Outlooks on Pest Management 28(5): 196– 201.
Goergen, G., Kumar, P.L., Sankung, S.B., Togola, A., and Tamò. M. 2016. First report of outbreaks of the fall armyworm Spodoptera frugiperda (J E Smith) (Lepidoptera, Noctuidae), a new alien invasive pest in West and Central Africa. PLoS ONE 11(10): e0165632.
Gorbet, D.W. and Tillman. B.L. 2009. Registration of ‘Florida-07′ peanut. J. Plant Regist. 3: 14– 18.
Gowda, M.V.C., Motagi, B.N., Naidu, G.K., Diddimani, S.B., and Sheshagiri. R. 2002. GPBD 4: A Spanish bunch groundnut genotype resistant to rust and late leaf spot. International Arachis Newsletter 22: 29– 32.
Hammons, R.O. 1970. Registration of Southeastern Runner 56-15 peanuts. Crop Sci. 10(6): 727.
Holbrook, C.C., Timper, P., Culbreath, A.K., and Kvien. C.K. 2008. Registration of ‘Tifguard’ peanut. J. Plant Regist. 2: 92– 94.
Kogan, M. and Ortman. E.F. 1978. Antixenosis: A new term proposed to define Painter’s nonpreference modality of resistance. Bull. Entomol. Soc. Am. 24: 175– 176.
Korani, W., Clevenger, J.P., Chu, Y., and Ozias-Akins. P. 2019. Machine learning as an effective method for identifying true single nucleotide polymorphisms in polyploid plants. Plant Genome 12: 180023.
Leuck, D.B. and Skinner. J.L. 1971. Resistance in peanut foliage influencing fall armyworm control. J. Econ. Entomol. 64: 148– 150.
Lynch, R.E., Branch, W.D., and Garner. J.W. 1981. Resistance of Arachis species to the fall armyworm, Spodoptera frugiperda. Peanut Sci. 8: 106– 109.
Murúa, G. and Virla. E. 2004. Population parameters of Spodoptera frugiperda (Smith) (Lep.: Noctuidae) fed on corn and two predominant grasses in Tucuman (Argentina). Acta Zool. Mex. 20(1): 199– 210.
Ortega, M.A., All, J.N., Boerma, H.R., and Parrott. W.A. 2016. Pyramids of QTLs enhance host–plant resistance and Bt-mediated resistance to leaf-chewing insects in soybean. Theor. Appl. Genet. 129: 703– 715.
Painter, R.H. 1951. Insect resistance in crop plants. Soil Sci. 72: 481.
Simpson, C.E. and Starr. J.L. 2001. Registration of ‘COAN’ peanut. Crop Sci. 41: 918.
Simpson, C.E., Starr, J.L., Church, G.T., Burow, M.D., and Paterson. A.H. 2003. Registration of ‘Nema TAM’ peanut. Crop Sci. 33: 1418.
Sparks, A.N. 1986. Fall armyworm (Lepidoptera: Noctuidae): Potential for area-wide management. Fla. Entomol. 69(3): 603– 614.
Stalker, H.T. 2017. Utilizing wild species for peanut improvement. Crop Sci. 57: 1102– 1120.
Stalker, H.T. and Mozingo. L. 2001. Molecular markers of Arachis and marker-assisted selection. Peanut Sci. 28: 117– 123.
Stalker, H.T., Tallury, S.P., Seijo G.R., and Leal-Bertioli. S.C. 2016. Biology, speciation, and utilization of peanut species. In: Stalker, H.T. and R.F. Wilson,(Eds.), Peanuts: Genetics, Processing, and Utilization. Academic Press and AOCS Press, pp. 27-66.
Tallury, S.P., Isleib, T.G., Copeland, S.C., Rosas-Anderson, P., Balota, M., Singh, D., and Stalker. H.T. 2014. Registration of two multiple disease-resistant peanut germplasm lines derived from Arachis cardenasii Krapov. & Gregory, W.C. GKP 10017. J. of Plant Regist. 8(1): 86– 89.
Todd, J.W., Beach, R.M., and Branch. W.D. 1991. Resistance in eight peanut genotypes to foliar feeding of fall armyworm, velvetbean caterpillar and corn earworm. Peanut Sci. 18: 38– 40.
Yang, G., Espelie, K.E., Todd, J.W., Culbreath, A.K., Pittman, R.N., and Demski. J.W. 1993. Cuticular lipids from wild and cultivated peanuts and the relative resistance of these peanut species to fall armyworm and thrips. J. Agric. and Food Chem. 41: 814– 818.
Yu, S.J. 1991. Insecticide resistance in the fall armyworm, Spodoptera frugiperda (J. E. Smith). Pestic. Biochem. Phys. 39(1): 84– 91.
Notes
- First and last authors: Institute of Plant Breeding, Genetics and Genomics, University of Georgia, Tifton, GA 31793; Second, third, and last authors: Horticulture Department, University of Georgia, Tifton, GA 31793. Fourth author: Department of Crop and Soil Sciences, North Carolina State University, Raleigh, 27695. Fifth and sixth authors: USDA-Agricultural Research Service, Crop Genetics and Breeding Research Unit, Tifton, GA, 31793. Sixth author: USDA-Agricultural Research Service, Corn Host Plant Resistance Research Unit, Mississippi State, MS, 39762. [^] *Corresponding author Email: pozias@uga.edu
Author Affiliations