Introduction
Water stress (WS) can affect the structural attributes of membrane lipids, nucleic acids and proteins (Vertucci and Leopold, 1987). WS also enhances accumulation of sugars and free amino acids and decreases leaf turgidity, chlorophyll, ascorbic acid and soluble proteins in leafy vegetables (Guehl et al., 1993; Irigoyen et al., 1992; Ishitani et al., 1995; Keller and Ludlow, 1993; Nguyen and Lamant, 1988). Plants subjected to water stress show changes in physiological and biochemical processes reflected in altered enzyme activities and exhibit enhanced activities of proteases, and increased concentration of low-molecular weight. nitrogenous compounds (Jharna et al., 2001; McCue and Hanson, 1990; Smirhoff and Colombe, 1989). Investigations of osmoregulation at the whole plant level indicated that free amino acids and proteins may play a significant role in response to water deficits and create a dynamic adjustment in nitrogen metabolism (Stewart and Larher, 1980) contributing to osmotic adjustment (Morris et al., 1990; Jacobson et al., 1992; Delauney and Verma, 1993). Metabolic changes in response to water stress include reduction in photosynthetic activity (Ritchie et al., 1990) and in protein synthesis (Mason et al., 1988) and accumulation of organic acids such as malate, citrate and lactate accompanied by accumulation of proline, sugars and betaine (Aspinall and Paleg, 1981; Bohnert et al., 1995; Shinozaki and Yamaguchi, 1996; Tabaeizadeh, 1998). Exposure of plants to low water potential often leads to loss of cell turgor, and plants undergo osmotic adjustments by the rapid accumulation of abscisic acid (ABA) and osmoprotectants (Bray, 1997; Grumet and Hanson, 1986; Skriver and Mundy, 1990). Dehydrins and Rab-related proteins are typically accumulated (Zhu, 2002) in response to low temperature, drought, salinity, or extra cellular freezing. These proteins are believed to be associated with desiccation tolerance, and their major function is to rescue the metabolic processes that are required for survival and regrowth under drought conditions. Our previous studies using dehydrin antibodies showed that peanut seed produces dehydrins following water stress, and the amount of dehydrins increase with increased stress period (Mazhar and Basha, 2002). Pre-harvest aflatoxin contamination is a major problem in semi-arid tropics where drought stress aggravates the Aspergillus infection and aflatoxin contamination in peanut (Cole et al., 1989; Sanders et al., 1993; Turner et al., 2000). Aspergillus thrive in extremely dry conditions and invade seeds under high temperatures causing severe damage in peanut seeds (Widstorm et al., 2003). Drought stress is known to predispose peanut to aflatoxin contamination (Blankenship et al., 1984; Cole et al., 1982 and 1985; Hill et al., 1983; Wilson and Stansell, 1983) possibly due to loss of the phytoalexin producing capacity of the peanut seed (Wooton and Strange, 1985; Dorner et al., 1989) and altered seed composition. Occurrence of aflatoxin contamination depends upon maturity status, condition of the seed and duration of stress. This would suggest that, susceptibility of peanuts to aflatoxin contamination is inconsistent and vary depending upon genotype, environment, maturity and severity of stress. Exposure of peanut plants to water stress for 5 to 20 d showed significant increases in leaf amino acids, soluble sugars, proteins and polypeptide composition (Ali-Ahmad and Basha, 1998; Amaregouda et al., 1994; Patil and Patil, 1993; Purushotham et al.,, 1998; Sharma et al.,, 1990). The genetic mechanisms leading to drought induced aflatoxin contamination are still unclear (Holbrook and Stalker, 2003). Depending upon the developmental stage or the external stimuli applied, the drought induced genes are classified as dehydrin (dehydration-induced), RAB (responsive to ABA), or LEA (late embryogenesis abundant) genes. Although genes induced by drought have been identified in a wide range of plant species (Iuchi et al., 1996; Oliver et al., 1998; Shinozaki and Yamaguchi, 1996), molecular basis of plant's tolerance to water stress remains far from being completely understood (Cellier et al., 1998; Ingram and Bartels, 1996). Proteomics has proved to be a powerful tool for identification of protein and mechanisms involved in drought response and tolerance (Hajhaeidri 2005; Kawasaki et al., 2000; Riccardi et al., 1998; Salekeh et al., 2002 a and b) and has been used to identify stress induced proteins in several crops that may play a role in food and nutrition (Haravarth et al., 2005). Several stress-induced genes have been isolated which play major role in understanding the molecular mechanism of drought. The ABA responsive gene expression modulated during seed development (Choi et al., 1987; Galau et al., 1987; Gomez et al., 1988; Heikkila et al., 1984) have been isolated in response to plant dehydration (Bray, 1997; Close et al., 1989; Guerrero and Mullet, 1986) or low temperature (Close, 1997; Shinozaki and Yamaguchi, 1996). Zegzouti et al. (1997) have isolated a novel ethylene-responsive cDNA, designated ER5 by differential display which also showed strong homology to LEA protein family. In peanut, several stress induced genes such as dehydration-induced cDNA encoding a putative 9 cis-epoxycartenoid dioxygenase (Wan and Li, 2005) and phospholipase D (Guo et al., 2006) have been isolated. All these genes have been found useful in germplasm screening for drought tolerance. Luo et al. (2005 a and b) have generated 1825 Expressed Sequence Tag (EST) libraries using leaf and seed tissue of two peanut lines, and found that 52.8% of the ESTs from leaf tissue and 78.6% of the ESTs from the pod tissue have homology to genes of known function. The major seed storage protein genes of arachin, conarachin and conglutin were sequenced and full-length cDNAs that encode isoforms of five arachin and six conglutin precursors were obtained (Yan et al., 2005). Earlier, we have identified the isoforms of conglutin as methionine rich protein subunits (Basha and Pancholy, 1981). Several peanut seed and leaf derived ESTs were used to validate their function through microarray and real-time PCR, however, their association with drought and disease tolerance traits are not yet confirmed (Cushman and Bohnert, 2000; Keon et al., 2003; Luo et al., 2003, 2005 a and b; Seki et al., 2001). Improvements in drought and disease tolerance characteristics are critical in peanut breeding programs aimed at reducing aflatoxin contamination. However, limited information is available on peanut seed components, especially the seed proteins and their role in maintaining drought tolerance in peanut. The primary objective of this research was to identify the proteins/polypeptides affected by drought stress for their possible use as markers for evaluating peanut germplasm to select drought-tolerant genotypes. Here, we report the seed polypeptides that are affected by water stress, and show that these polypeptides are differentially expressed among peanut genotypes under water stress. These proteins may be useful as potential markers for screening peanut genotypes to identify drought tolerant genotypes.
Materials and Methods
Plant Material and Treatments
Peanut (Arachis hypogaea L.) genotypes with diverse drought tolerance characteristics (A.N.G.R. Agricultural University, Kadiri, India) were used in this study. Drought-tolerant (DT) and drought-susceptible (DS) designations are given to these genotypes based on extensive yield and field performance trials conducted across India under rain-fed conditions through the National Research Center for Groundnut (NRCG). However, so far no attempts have been made to determine their biochemical and drought-response characteristics. DT genotypes included: Kadiri-5, Vemana, K-1319, K-1375, JL-220, CSMG-84-1, ICGS-44, ICGS-76, TAG-24, TMV-2, TPT-2, GG-2, TIR-31, JUG-37, and ICR-25. DS genotypes included: Kadiri-3, Kadiri-4, K-1340, K-1341, TKG-19A, M-13, and JL-24. The seed were surface sterilized with 70% ethanol, washed, and planted in three gallon pots. Peanut plants were maintained in the greenhouse and after 120 d they were subjected to WS by withholding irrigation. Control plants were irrigated daily. All treatments were conducted in triplicate, and the experimental period was from May 2004 to September 2004. Six replicates were maintained for each experiment and for each genotype. Peanut pods (Brown to Black stages; Williams and Drexler, 1981) were collected from control and treatments at 7, 14 and 28 d after the stress. At least 15 pods were collected from each replicate in each genotype which yielded over 30 seeds. The average seed size range from 0.5 to 2.0 cm depending on the genotype. The pods were shelled, seed were freeze-dried and stored at −80 C for further analysis.
Soil Water Potential
The soil water potential was measured at weekly intervals using Quick Draw soil moisture probe (Model 2900F1, Soil moisture Equipment Corp., Santa Barbara, CA). The soil water potential was measured thrice for all the genotypes in all the six replicates, and the average values were taken. The average soil water potential was 10, 16, 28 and 38 cb at 7, 14, 21 and 28 d stress periods, respectively.
Protein Extraction
The seed were ground into a meal using a mortar and pestle. The meal was defatted with hexane (Basha and Pancholy, 1981). A portion (50 mg) of the defatted meal was homogenized (1 ml) in dissociation buffer [1.2% (w/v) Tris, 2% (w/v) SDS and 5% (v/v) 2- mercaptoethanol], heated in a boiling water bath for 3 min and used for one-dimensional gel electrophoresis.
Quantification of Proteins
The total protein content of the samples was determined as per the method of Bradford (1979). Bovine Serum Albumin (BSA) was used as the standard to quantify the total protein in the sample. Equal concentration (75 µg) of protein was loaded on each lane of SDS-PAGE.
One-dimensional Gel Electrophoresis
The denatured protein (75 µg) was subjected to SDS-gel electrophoresis (Laemmli, 1970) in 11 % (w/v) polyacrylamide gel under denaturing conditions. The gel was run at 30 mA of current and Bromophenol Blue was used as the tracking dye. Following electrophoresis the proteins were detected using Coomassie blue R-250 (0.25%).
Two-dimensional Gel Electrophoresis (2-d Page)
Defatted peanut meal was extracted with a buffer containing 9.3 M urea, 5 mM K2CO3, 0.5% (w/v) dithiothreitol and 2% (v/v) Nonidet P-40 and subjected to two-dimensional gel electrophoresis as described by Basha (1979). The above extraction buffer has been reported (Basha, 1979) to solublize more than 98% of the seed proteins and hence, the extract should represents all the seed protein population. The 2-D PAGE consisted of isoelectricfocusing in the first dimension and SDS-PAGE (12% polyacrylamide) in the second dimension. After electrophoresis, the gels were stained with Coommassie Blue R-250 (0.25%) to visualize the protein spots. Bromophenol Blue was included in the overlay buffer to determine the end point of the run.
Protein Sequencing and Characterization
The proteins showing significant suppression in response to water stress were cut out from the 2D- PAGE gel and transferred into 2 ml Eppendorf tubes. The samples were digested with trypsin, subjected to Liquid Chromatography and the individual peptides were sequenced using Mass Spectrophotometer at the Protein Core Facility, University of Florida, Gainesville, FL. Amino acid sequences were compared with the sequences in non-redundant public sequence database of the NCBI by using BLAST program (Altschul et al., 1990).
Results
Qualitative and quantitative change in peanut seed proteins due to water stress was determined by comparing polypeptide profiles of control and stress treatments, and among peanut genotypes.
Effect of Water Stress on Peanut Seed Polypeptide Composition
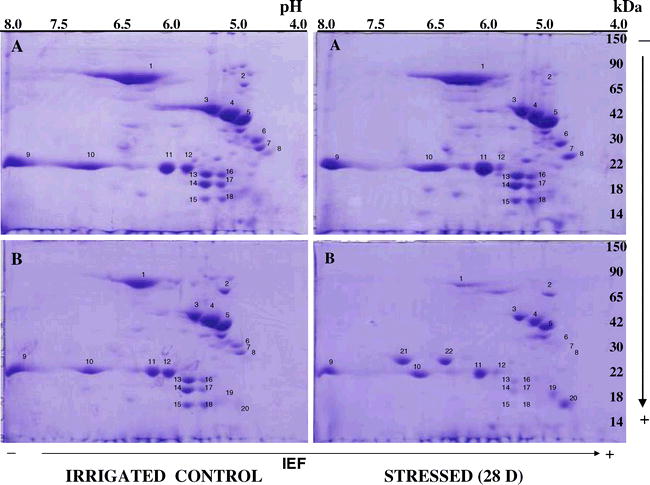
WS had a variable effect on peanut seed protein content and composition. These responses included over-expression, suppression and appearance of new proteins in water-stressed seed compared to the irrigated control seed. WS induced only minor changes in seed polypeptide composition of certain genotypes while in others it caused significant changes in seed polypeptide composition (Fig. 1). WS affected polypeptides with apparent molecular weight (Mr) around 70, 35, 25, 20, 18 and 14 kDa. Most of the changes in seed polypeptide composition occurred between 0 and 7 d of WS, and additional periods (14 and 28 d) of stress caused only limited changes in seed polypeptide composition. The nature and characteristics of drought-responsive polypeptides was further determined by two-dimensional gel electrophoresis (2-D PAGE). The 2-D PAGE resolves proteins based on their isoelectric point (pI) and molecular weight (Fig. 2). Each protein spot on 2-D gel was numerically labeled for easy identification. We have identified over 22 protein spots on 2-D gel that are differentially expressed in DT and DS genotypes. The 2-D PAGE data clearly showed that WS had a significant effect on DS genotype M-13 (Fig. 2B) and minimal effect on DT genotype Vemana (Fig 2A). For example, in DT genotype Vemana, only three polypeptides # 7, 8, 12 (pI 4.0 and 6.0; molecular weight 22 and 32 kDa were affected (Fig 2A), while in DS genotype M-13, more than 16 polypeptides (protein spots with isoelectric points between 4.5 to 6.5 pH and molecular weights between 14 to 75 kDa were affected (Fig. 2B). In addition, we have found that in DS genotype, M-13, two proteins (#19 and #20) with isoelectric points between 4.5 and 5.0 pH and molecular weights between 15 and 19 kDa were over expressed and two proteins (#21 and #22 with isoelectric points between 6.25 and 7.25 pH and molecular weights between 22 and 25 kDa) were newly synthesized. Thus, the electrophoretic data clearly demonstrated that peanut genotypes respond differentially to WS by undergoing varying levels of changes in seed polypeptide composition. Overall, WS had the following effects on peanut seed polypeptide content and composition: a) decrease in certain polypeptide levels, b) increase in certain polypeptide levels and c) appearance of new polypeptides. The observed differences in seed polypeptide content and composition between the DT and DS genotypes could be attributed to differences in gene expression resulting in down-regulation, up-regulation or de novo synthesis of proteins. These results are consistent with our earlier report (Jain et al., 2001) on peanut where we have shown up-regulation and down-regulation of several mRNA transcripts following WS, and identified 43 drought-responsive transcripts affected due to WS.
Differential Response of Peanut Genotypes to Water Stress
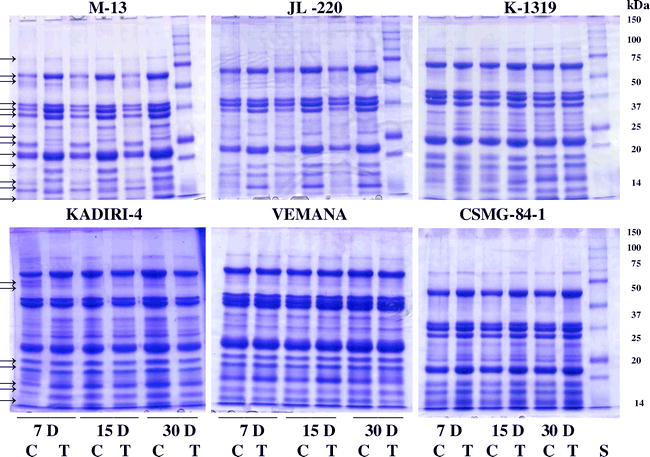
Genetic differences in the response of peanut genotypes with varying drought tolerance characteristics such as DT and DS to WS was determined by subjecting them to different periods of WS. The results showed that peanut genotypes respond differentially to WS depending on genotype and duration of stress. WS had major impact on polypeptide composition of certain peanut genotypes while it had minor effect on others (Fig. 3). For example, seed polypeptide composition of genotypes M-13 and JL-220 was significantly altered due to WS while it had little effect on genotypes Vemana and CSMG-84-1 (Fig. 3). Likewise, in other genotypes such as K-1319 and Kadiri-4 only moderate changes were observed in seed polypeptide composition due to WS (Fig. 3). In M-13, most of the polypeptides (molecular weight between 10 to 100 kDa) were affected due to WS while in Vemana no major changes were observed in seed polypeptide composition. These responses may be the result of differential gene expression as observed by Jain et al., 2001; Devaiah et al., 2007), who reported that WS resulted in up-regulation, down-regulation and expression of new transcripts in peanut. It should be noted that the genotypes viz. Vemana, CSMG-84-1, Kadiri-4, TMV-2, etc., showing limited changes in polypeptide composition are categorized as DT while genotypes M-13, JL-220, JL-24, TKG-19A, etc., are placed under DS category based on their field performance and yield under rain-fed conditions (ANGR Agricultural University, Kadiri; National Research Council for Groundnut, Junagadh). Results of this study showing minimal changes in seed protein composition of DT genotypes is also consistent with our earlier report (Jain et al., 2001) where we had observed expression of drought-responsive transcripts (PTRD-1, PTRD-2, PTRD-6, PTRD-10, PTRD-11, PTRD-16, PTRD-17) for a longer period (2 to 3 weeks) in DT line 72a (PI 145681) compared to DS genotype (1 week in Florunner). These data suggest that the DT genotypes maintain their seed protein levels by regulating gene expression for longer period than the DS genotypes.
Identification of Differentially Expressed Proteins
The proteins that are suppressed due to water stress were sequenced and characterized. We have found that seven proteins (# 1, 13, 14, 15, 16, 17, 18) that were significantly suppressed in DS genotype M-13 have maintained their expression in DT genotype throughout the water stress treatment period for 28 d. Protein #1 was slightly over expressed in response to water stress in DT while it was suppressed in DS genotype. The MASCOT analysis shows the sequence similarity of Protein # 1 with allergen protein Ara h1 (P43238). The expression of proteins # 13 to #18 were not affected in DT genotype, while they were significantly suppressed in DS upon water stress. The proteins #13 and 16 are found to be isomer of arachin Ahy-3 (AY722687), proteins # 14 and 17 are isomers of 2S protein 1 of Arachis hypogaea (AY 722689) and protein # 15 and 18 the isomers of conglutin (AY848699). The sequences of these polypeptides were deposited in EMBL database (P43238, Q647G9, Q6PSU2, and Q647H2). These proteins are known to be the major seed storage proteins in legumes. Our results indicated that in DT (Vemana), expression levels of these proteins were maintained during water stress while in DS (M-13) they were significantly suppressed.
Grouping of Peanut Genotypes
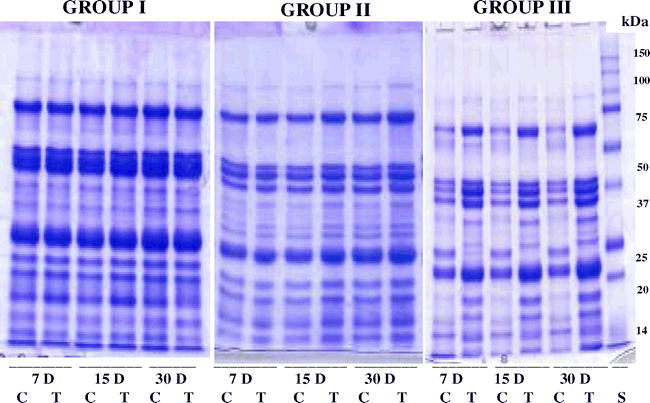
Electrophoretic evaluation of more than 30 peanut genotypes revealed differential protein expression following WS. Comparison of level of changes in seed polypeptide composition with that of drought-tolerance level revealed a negative correlation between them. Thus, DT genotypes showed little change in seed polypeptide composition while DS genotypes showed major change in seed polypeptide composition. The less drought-tolerant/drought-susceptible genotypes showed moderate changes in seed polypeptide content and composition. Based on this observation peanut genotypes are arranged into three groups: Group I = includes the genotypes showing limited (<2) changes in seed polypeptide composition, Group II = includes genotypes showing moderate (2 to 6) changes in seed polypeptide composition and Group III= includes the genotypes showing major (> 6) changes in seed polypeptide composition (Fig. 4). Thus, Group I includes most DT genotypes while Group II includes less drought-tolerant/drought-susceptible and Group III includes most DS genotypes.
Discussion
Water stress alters plant gene expression leading to the induction of specific gene/s (Romo et al., 2001) accompanied by increased transcription and thus an increase of the corresponding proteins. In this study we have evaluated peanut genotypes for water stress-regulated proteins in order to identify the seed proteins affected by WS, and to understand the mechanisms associated with peanut plant acclimation and adaptation to drought. Identification of drought-responsive proteins would be useful to distinguish DT and DS peanut genotypes, and to understand the biochemical and molecular events responsible for drought-induced aflatoxin contamination of peanut. Some of the peanut genotypes used in this study were developed (ANGR Agri. Univ., Kadiri; National Research Center for Groundnut and ICRISAT, India) for drought tolerance, and are cultivated under rain-fed conditions in the arid and semi-arid regions of India. For example, the genotypes viz. Vemana, CSMG-84-1, Kadiri-4, and TMV-2 have been developed and released as DT cultivars for Andhra Pradesh, India. In addition, these genotypes are also being widely used in several breeding programs at Kadiri, Tirupathi, and Hyderabad, India for identifying peanut genotypes with high water use efficiency. Inclusion of these genotypes with proven drought-tolerance characteristics was helpful in identifying drought-responsive proteins as well as demonstrating that WS had limited effect on their seed protein composition. The genotypes, M-13, JL-220, JL-24 and TKG-19A which are classified as DS showed significant changes in seed polypeptide composition following WS, reflecting their DS trait. Peanut seed proteins have been grouped into arachin, conarachin / non-arachin (conglutin) based on their solubility (Basha and Pancholy, 1981). Our results clearly revealed that seed polypeptide composition, specifically arachin and conglutin proteins of DT genotypes were least affected while that of DS genotypes, significantly altered due to WS. Our data showed that proteins # 1 and 13 to18 were significantly suppressed in DS while they were unaffected in DT during water stress. Based on amino acid sequence data, protein # 1 is identified as Ara h1, the arachin subunit (Kolarich and Altman, 2000). Earlier, Basha and Pancholy (1981 a and b) have isolated the conglutin subunits and found two of them (# 15) to be rich in methionine (>4%). The MRP (methionine-rich protein) is a 120 kDa protein and consists of six polypeptides with molecular weights ranging from 15.5 to 20 kDa and pI between 4.5 and 5.5 pH. Later, the gene sequences of these conglutin subunits were identified by Yan et al., (2005). Our results show that the storage proteins, particularly the MRP subunits were highly suppressed in DS genotypes compared to DT genotypes due to WS. This data would suggest that the observed differential responses in subset of arachin and conglutin proteins could be employed to distinguish DT and DS peanut genotypes. Previously, our studies using Differential Display Reverse Transcriptase technique has enabled us to identify peanut transcripts affected by water stress (Jain et al., 2001). The observations are consistent with our earlier findings (Jain et al., 2001) which showed prolonged gene expression in DT genotypes compared to DS genotypes upon imposition of various levels of WS. In our previous studies, we have found transcripts that are severely affected (up-regulated, down-regulated or newly produced) due to water stress and are termed as Peanut Transcripts Responsive to Drought (PTRD). The down-regulated transcripts were studied by Northern Dot-Blot hybridization in drought-tolerant and drought-susceptible peanut lines. Results showed differential expression of these transcripts in the drought-tolerant and drought-susceptible lines indicating that these three transcripts (PTRD-1, PTRD-2 and PTRD-16) can be used as molecular tools to screen peanut germplasm for identifying drought-tolerant lines. Once the genotypes are classified, it is necessary to work with a large number of genotypes differing in their drought tolerance level to isolate differentially expressed transcripts induced upon water stress to address various stress related problems in peanut. Such markers would be of great value in the breeding programs aimed at identifying DT genotypes for developing drought- and aflatoxin-tolerant peanut genotypes. Although several molecular markers are currently available in peanut (Burrow and Blake, 1998), these have limited polymorphism in peanut and proved inadequate for identifying the drought specific peanut genotypes. Therefore, identification of drought responsive proteins would be useful to determine drought tolerance characteristics of peanut genotypes. Further studies are needed to clone and characterize the genes expressed due to drought stress and use them as probe to study the expression levels across genotypes.
Acknowledgements
This research was supported by a grant from the USAID, Peanut CRSP, Grant #FAM 51.
Literature Cited
Ali-Ahmad M. and Basha S. M. 1998 Effect of water stress on composition of peanut leaves. Peanut Sci 25 : 31 – 34 .
Altschul S. F. , Gish W. , Miller W. , Myers E. W. , and Lipman D. J. 1990 Basic local alignment search tool. J. Mol. Biol 215 : 403 – 410 .
Amaregouda A. , Chetti M. B. , Ealimath P. M. , and Gowda G. V. C. 1994 Intolerance of Anti-transpirants on osmoregulants in summer groundnut (Arachis hypogaea L.). Ann. Plant Physiol 8 : 114 – 120 .
Aspinall D. and Paleg L. G. 1981 Proline accumulation - Physiological aspects. 205 – 240 In Paleg L. G. and Aspinall D. Physiology and Biochemistry of Drought Resistance in Plants New York Academic Press .
Basha S. M. 1979 Identification of cultivar differences in seed polypeptide composition of peanuts (Arachis hypogaea L.) by two-dimensional gel electrophoresis. Plant Physiol 63 : 301 – 306 .
Basha S. M. and Pancholy S. K. 1981 Identification of methionine-rich polypeptides in peanut seed. J Agric Food Chem 29 : 331 – 335 .
Basha S. M. and Pancholy S. K. 1981 Changes in the polypeptide composition of maturing seeds from four peanut (Arachis hypogea L.) cultivars. Peanut Sci 8 : 6 – 10 .
Blankenship P. D. , Cole R. J. , Sanders T. H. , and Hill R. A. 1984 Effect of geocarposphere temperature on pre-harvest colonization of drought-stressed peanut by Aspergillus flavus and subsequent aflatoxin contamination. Mycopathologia 85 : 69 – 74 .
Bohnert H. J. , Nelson D. E. , and Jensen R. G. Adaptations to environmental stress. Plant Cell 1995 7 : 1099 – 1111 .
Bradford M. 1976 A rapid and sensitive method for the quantification of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem 72 : 248 – 254 .
Bray E. A. 1997 Plant responses to water deficit. Trends in Plant Sci 2 : 48 – 54 .
Burrow M. D. and Blake T. K. 1998 Molecular tools for the study of complex traits. 13 – 29 In: Patterson A. H. Molecular dissection of complex traits Boca Raton, NY, USA CRC Press .
Cellier F. , Conejero G. , Breitler J. C. , and Case F. 1998 Molecular and physiological responses to water deficit in drought-tolerant and drought-sensitive lines of sunflower. Plant Physiol 116 : 319 – 328 .
Choi J. H. , Liu L. , Borkird C. , and Sung Z. R. 1987 Cloning of genes developmentally regulated during plant embryogenesis. Proc. Natl. Acad. Sci. U.S.A 84 : 1906 – 1910 .
Close T. J. , Kortt A. A. , and Chandler P. M. 1989 A cDNA-based comparison of dehydration-induced proteins (dehydrins) in barley and corn. Plant Mol. Biol 13 : 95 – 108 .
Close T. J. 1997 Dehydrins: a commonalty in the response of plants to dehydration and low temperature. Physiologia Plant 100 : 291 – 296 .
Cole R. J. , Hill R. A. , Blankenship P. D. , Sanders T. H. , and Garren K. H. 1982 Influences of irrigation and drought stress on invasion by Aspergillus flavus in corn kernels and peanut pods. Dev. Ind. Microbiol 23 : 229 – 236 .
Cole R. J. , Sanders T. H. , Hill R. A. , and Blankenship P. D. 1985 Geocarposphere temperatures that induce pre-harvest aflatoxin contamination of peanuts under drought-stress. Mycopathologia 91 : 41 – 46 .
Cole R. J. , Sanders T. H. , Hill R. A. , and Blankenship P. D. 1989 Environmental conditions required to induce pre-harvest concentration in groundnut. Summary of Six Years Research. 279 – 287 In: Proceedings of International Workshop on Aflatoxin Concentration in groundnut. International Crop Research Institute for Semi-Arid Tropics, Patancheru, India 6–9 Oct. 1987
Cushman J. C. and Bohnert H. J. 2000 Genomic approaches to plant stress tolerance. Curr. Opin. Plant Biol 3 : 117 – 124 .
Delauney A. J. and Verma D. P. S. 1993 Proline biosynthesis and osmoregulation in plants. Plant J 4 : 215 – 223 .
Devaiah K. M. , Bali Geetha , Athamaram T. N. , and Basha S. M. 2007 Identification of two new genes from drought tolerant peanut up-regulated in response to drought. Plant Growth Regul 52 : 249 – 258 .
Dorner J. W. , Cole R. J. , Sanders T. H. , and Blankenship P. D. 1989 Interrelationship of kernel water activity, soil temperature, maturity, and phytoalexin production in pre-harvest aflatoxin contamination of drought stressed peanuts. Mycopathologia 105 : 117 – 128 .
Galau G. A. , Bijaisoradat N. , and Hughes D. W. 1987 Accumulation kinetics of cotton late embryogenesis-abundant (LEA) mRNAs and storage protein mRNAs: Coordinate regulation during embryogenesis and role of abscisic acid. Develop. Biol 123 : 198 – 212 .
Gomez J. , Sanchez-Martinez D. , Steifel V. , Rigau J. , Puigdomench P. , and Pages M. 1988 A gene induced by the plant hormone abscisic acid in response to water stress encodes a glycine rich protein. Nature 334 : 262 – 264 .
Grumet R. and Hanson A. D. 1986 Genetic evidence for an osmoregulatory function of glycinebetanin accumulation in barley. Aus. J. Pl. Physiol 13 : 353 – 364 .
Guehl F. M. , Clement A. , Kaushal P. , and Aussenac A. 1993 Tree Physiol. 12 : 173 – 183 .
Guerrero F. and Mullet J. E. 1986 Increased abscisic acid biosynthesis during plant dehydration requires transcription. Plant Physiol 80 : 588 – 591 .
Guo B. Z. , Xu G. , Cao Y. G. , Holbrook C. C. , and Lynch R. E. 2006 Identification and characterization of phospholipase D and its association with drought susceptibilities in peanut (Arachis hypogaea). Planta 223 / 3 : 512 – 20 .
Hajhaeidri M. , Abdollahian-Noghabi M. , Askari H. , Hedari M. , Sadeghian S. Y. , Ober E. S. , and Hosseini Salekdeh G. N. 2005 Proteome analysis of sugar beet leaves under drought stress. 11. Proteomics 5 : 950 – 960 .
Haravarth S. , Szabo Z. , Janaky T. , Pauk J. , and Hajos G. 2005 Proteomics as an emerging tool for identification of stress-induced proteins in control and genetically modified wheat lines. Chromatographia Supplement 63 : S143 – 147 .
Heikkila J. J. , Papp J. E. T. , Schultz G. A. , and Bewley J. D. 1984 Induction of heat shock protein messenger RNA in maize mesophylls by water stress, abscisic acid, and wounding. Plant Physiol 76 : 270 – 274 .
Hill R. A. , Blankenship P. D. , Cole R. J. , and Sanders T. H. 1983 Effect of soil moisture and temperature on pre-harvest invasion of peanuts by Aspergillus flavus group and subsequent aflatoxin contamination. Appl. Environ. Microbiol 45 : 628 – 933 .
Holbrook C. C. and Stalker H. T. 2003 Peanut breeding and genetic resources. Plant Breed. Rev 22 : 297 – 356 .
Ingram J. and Bartels D. 1996 The molecular basis of dehydration tolerance in plants. Ann. Rev Plant Mol Biol 47 : 377 – 403 .
Irigoyen J. , Emerich D. W. , and Sanchez Diaz M. 1992 Water stress induced changes in concentrations of proline and total soluble sugars in nodulating alfalfa (Medicago sativa) plants. Physiol. Planta 84 : 55 – 60 .
Ishitani M. , Nakamura T. , Han S. Y. , and Takabe T. 1995 Expression of the betaine aldehyde dehydrogenase gene in barley in response to osmotic stress and abscisic acid. Plant Mol. Biol 27 : 307 – 315 .
Iuchi S. , Yamaguchi-Schinozaki K. , Urao T. , Terao T. , and Shinozaki K. 1996 Novel drought-inducible genes in the highly drought-tolerant cowpea: cloning of cDNAs and analysis of the expression of the corresponding genes. Plant and Cell Physiol 37 : 1073 – 1082 .
Jacobsen K. S. , Sepelund M. , Saeboe-Lassen S. D. , Wayne A. , Glenn F. , and Larsen G. 1992 Late embryogenesis abundant genes encoding protein with different members of hydrophilic repeats are regulated differentially by abscisic acid and osmotic stress. Plant J 2 : 241 – 252 .
Jain A. K. , Basha S. M. , and Holbrook C. C. 2001 Identification of drought responsive transcripts in peanut (Arachis hypogaea L.). Electronic J. Biotechnology 4 : 59 – 67 .
Jharna D. E. , Chowdhury B. L. D. , Haque M. A. , Bhuiyan M. R. H. , and Husain M. M. 2001 Biochemical screening of some groundnut (Arachis hypogaea L.) genotypes for drought tolerance. J. Biol. Sci 1 : 1009 – 1011 .
Kawasaki S. , Miyake C. , Kohci T. , Fujii S. , Uchida M. , and Yokota A. 2000 Response of wild watermelon to drought stress: accumulation of an ArgE homologue and citrulline in leaves during water deficits. Plant Cell Physiol 102 : 1353 – 1354 .
Keller F. and M Ludlow M. 1993 Carbohydrate metabolism in drought- stressed leaves of pigeonpea. (Cajanus cajan). J. Exp. Bot 44 : 1351 – 1359 .
Keon J. , Curtis R. , Cabrera H. , and Hargreaves J. 2003 A genomic approach to crop pest and disease research. Pest Manage. Sci 59 : 43 – 148 .
Kolarich D. and Altmann F. 2000 N-glycan analysis by matrix-assisted laser desorption / ionization mass spectrometry of electrophoretically separated non-mammalian proteins: application to peanut allergen Ara h 1 and olive pollen allergen Ole e 1. Anal. Biochem 285 : 64 – 75 .
Laemmli U. K. 1970 Cleavage of structural proteins during the assembly of the head of T4. Nature 227 : 680 – 685 .
Luo M. , Bausher M. G. , Holbrook C. C. , Lee R. D. , Lynch R. E. , and Guo B. Z. 2003 Application of EST technology in functional genomics of Arachis hypogaea. L. Phytopathol 95 : 381 – 387 .
Luo M. , Dang P. , Guo B. Z. , He G. , Holbrook C. C. , Bausher M. G. , and Lee R. D. 2005 (a) Generation of Expressed Sequence Tags (ESTs) for gene discovery and Marker development in Cultivated Peanut. Crop Sci 45 : 346 – 353 .
Luo M. , Liang X. Q. , Dang P. , Holbrook C. C. , Bausher M. G. , Lee R. D. , and Guo B. Z. 2005 (b) Microarray-based screening of differentially expressed genes in peanut in response to Aspergillus parasiticus infection and drought stress. Plant Science 169 : 695 – 703 .
Mason H. S. , Guerreo F. D. , Boyer J. S. , and Mullet J. E. 1988 Proteins homologous to leaf glycoproteins are abundant in stem of dark grow soybean seedlings. Plant Mol. Biol 11 : 845 – 856 .
Mazhar H. and Basha S. M. 2002 Effect of desiccation of peanut (Arachis hypogea L.) seed protein Composition. Environ. Exp. Bot 47 : 67 – 75 .
McCue K. F. and Hanson A. D. 1990 Drought and salt tolerance: Towards understanding and application. Trends in Biotechnol 8 : 358 – 362 .
Morris P. C. , Kumar A. , Bowles D. J. , and Cuming A. C. 1990 Osmotic stress and abscisic acid induce expression of the wheat Em genes. Eur. J. Biochem 190 : 625 – 630 .
Nguyen A. and Lamant A. 1988 Pinitol and myo-inositol accumulation in water-stressed seedlings of maritime pine. Phytochemistry 27 : 3423 – 3427 .
Oliver M. J. , Wood A. J. , and O'Mahony P. 1998 To dryness and beyond - preparation for the dried state and rehydration in vegetative desiccation-tolerant plants. Plant Growth Regulation 24 : 193 – 201 .
Patil N. A. and Patil T. M. 1993 Physiological responses of groundnut genotypes to water stress. Legume Res 16 : 23 – 30 .
Purushotham M. G. , Patil V. , Reddy P. C. , Prasad T. G. , and Vajranabhaiah S. N. 1998 Development of in vitro PEG stress tolerant cell lines in two groundnut (Arachis hypogaea L.) genotypes. Indian J. Plant Physiol 3 : 49 – 51 .
Riccardi F. , Gazeau P. , de Vienne D. , and Zivy M. 1998 Protein changes in response to progressive water deficit in maize: quantitative variations and identification. Plant Physiol 117 : 1263 – 1253 .
Ritchie S. W. , Nguyen H. T. , and Holaday A. S. 1990 Leaf water content and gas exchange parameters of two wheat genotypes differing in drought resistance. Crop Sci 30 : 105 – 111 .
Romo S. , Labrador E. , and Dopico B. 2001 Water stress-regulated gene expression in Cicer arietinum seedlings and plants. Plant Physiol. Biochem 39 : 1017 – 1026 .
Sanders T. H. , Cole R. J. , Blankenship P. D. , and Dorner J. W. 1993 Aflatoxin concentration of peanuts from plants drought stressed in pod or root zones. Peanut Sci 20 : 5 – 8 .
Salekeh G. H. , Siopongco H. J. , Wade I. J. , Ghareyazie B. , and Bennett J. 2002 (a) A proteomics approach to analyzing drought- and salt-responsiveness in rice. Field Crops Res 76 : 199 – 219 .
Salekeh G. H. , Siopongco H. J. , Wade I. J. , Ghareyazie B. , and Bennett J. 2002 (b) A proteomics Analysis of rice leaves during drought stress and recovery. Proteomics 2 : 1131 – 1145 .
Seki M. , Narusaka M. , and Abe H. 2001 Monitoring the expression pattern of 1300 Arabidopsis genes under drought and cold stress by using full-length cDNA microarray. Plant Cell 13 : 61 – 72 .
Sharma K. S. , Sharma G. R. , and Sing G. 1990 Biochemical changes in groundnut seedlings grown under polyethylene glycol induced water stress. Environ. Ecol 8 : 854 – 856 .
Skriver K. and Mundy J. 1990 Gene expression in response to abscisic acid and osmotic stress. Plant Cell 2 : 503 – 512 .
Smirhoff N. and Colombe S. V. 1989 Drought influences the activity of enzymes of the chloroplast hydrogen peroxide scavenging system. J. Exptl. Bot 39 : 1097 – 1180 .
Shinozaki K. and Yamaguchi-Shinozaki K. 1996 Molecular responses to drought and cold stress. Curr. Opin. Biotech 7 : 161 – 167 .
Stewart G. R. and Larher F. 1980 A Comprehensive Teatise. 609 – 635 In Miflin B. J. Biochemistry of Plants, 5 NY Academic Press .
Tabaeizadeh Z. 1998 Drought-induced responses in plant cells. International Rev. Cytol 182 : 193 – 247 .
Turner P. C. , Mendy M. , White H. , Fortuin M. , Hall A. J. , and Wild C. P. 2000 Hepatitis B infection and aflatoxin biomarker levels in Gambian children. Trop. Med. Int. Health 5 : 837 – 841 .
Vertucci C. W. and Leopold A. C. 1987 The relationship between water binding and desiccation tolerance in tissues. Plant Physiol 85 : 232 – 238 .
Wan X. and Li L. 2005 Molecular cloning and characterization of a dehydration-inducible cDNA encoding a putative 9-cis-epoxycarotenoid dioxygenase in Arachis hygogaea. L. DNA Seq 16 / 3 : 217 – 23 .
Widstorm N. W. , Guo B. Z. , and Wilson D. M. 2003 Integration of crop management and genetics for control of pre-harvest aflatoxin contamination of corn. J. Toxicol. Toxin Rev 22 : 195 – 223 .
Williams E. J. and Drexler J. S. 1981 A non-destructive method for determining peanut pod maturity. Peanut Sci 8 : 134 – 137 .
Wilson D. M. and Stansell J. R. 1983 Effect of irrigation regimes on aflatoxin contamination of peanut pods. Peanut Sci 10 : 54 – 56 .
Wooton H. R. and Strange R. N. 1985 Circumstantial evidence for phytoalexin involvement in the resistance of peanuts to Aspergillus flavus. J. Gen. Microbiol 131 : 487 – 494 .
Yan Y. S. , Lin X. D. , Zhang Y. S. , Wang L. , Wu K. , and Huang S. Z. 2005 Isolation of peanut genes encoding arachins and conglutins by expressed sequence tags. Plant Sci 169 : 439 – 445 .
Zegzouti H. , Jones B. , Marty C. , Leliebre J. , Latche A. , Pech J. , and Bouzayen M. 1997 Plant Mol. Biol 27 : 587 – 595 .
Zhu J. K. 2002 Salt and drought stress signal transduction in plants. Ann. Rev. Plant Biol 53 : 247 – 273 .