Introduction
According to the published literature on the crossability of the species in the genus Arachis, none of the species outside section Arachis will hybridize with the cultivated peanut (Arachis hypogaea L.) without the aid of in vitro methodology. In many instances when hybrids are produced between A. hypogaea and wild species outside the section Arachis, hybrids are completely sterile or, in other words, genetic dead-ends (Stalker and Simpson, 1995). With the development of embryo rescue and tissue culture techniques, hybrids between A. hypogaea and A. glabrata Benth., (section Rhizomatosae) have been reported (Mallikarjuna and Sastri 1985, 2002; Shen et al., 1995) Arachis chiquitana Krapov., W.C Gregory and C.E. Simpson, is a wild species collected from Chiquitos province of Santa Cruz in Bolivia, and although it was previously placed in section Erectoides by Gregory and Gregory (1979), it is currently placed in section Procumbentes (Krapovickas and Gregory, 1994).
Aflatoxin contamination of peanut, caused by the Aspergillus flavus group of fungi, is one of the most important constraints to quality peanut production in the semi-arid rainfed areas of the world. This is particularly significant in relation to public health and international trade (Waliyar et al., 1994). In addition to peanuts, aflatoxins are found in many agricultural crops such as corn, cotton, wheat, and rice and consumption of the toxin by humans can lead to liver cancer. Househam and Hunt (1991) also reported a striking association between exposure to aflatoxin in children and both stunting and low weight gain. In West Africa, people are chronically exposed to high levels of aflatoxin starting in the utero and continuing throughout life (Gong et al., 2002). Genetic resistance should be one of the major components in a strategy to manage aflatoxin contamination in peanut, but high levels of genetic resistance to aflatoxin have not been identified in cultivated peanuts. Ghewande et al., (1989) reported accessions of A. duranensis Krapov. and W.C. Gregory and A. cardenasii Krapov. And W.C. Gregory as being highly resistant to in vitro seed colonization to Aspergillus flavus. Xue et al., (2004) confirmed resistance in three A. duranensis and two A. cardenasii accessions after testing 36 accessions of the two species. Arachis chiquitana is one of the three wild species identified as resistant to Aspergillus flavus Link ex. Fries colonization and subsequent aflatoxin production (Thakur et al, 2000). Pande and Rao (2001) also identified A. chiquitana as one of the wild species resistant to late leaf spot (LLS), a fungal disease caused by Phaeoisariopsis personata (Berk. & M.A. Curtis) Deighton. Hence, developing strategies to cross A. hypogaea with A. chiquitana not only broadens the genetic base of cultivated peanut but will also potentially bring in other desirable genes.
Materials and Methods
Seeds of A. chiquitana with 2n = 2x = 20, (ICG 11560; collector number 36025; PI 476004; Fig. 1A) from section Procumbentes were obtained from the genetic resources division of ICRISAT and grown in a glasshouse. Seeds of A. hypogaea cv. ICGS 44 (2n = 4x = 40) were also grown and maintained in the glasshouse. Cross pollinations using A. hypogaea as the female parent and A. chiquitana as the pollen donor were carried out before 10:00 a.m. Emasculations of anthers from flowers were carried out the previous evening. Application of 75 mg/L of gibberellic acid (GA) at the base of pollinated pistils was mandatory to obtain pods from cross pollinations. A total of 493 pollinations were carried out and 167 pods were harvested between 25 and 30 days after pollination. Pods were then surface sterilized and ovules were extracted under sterile conditions. Ovules which were more than 4.0 to 5.0 mm long were dissected and the embryos (immature seeds) were cultured directly on the semisolid growth medium (Fig. 1E). The growth medium consisted of MS (Murashige and Skoog, 1962) basal medium with 3% sucrose plus napthalene acetic acid (NAA; 0.1 mg/L) and benzylamino purine (BAP; 1.0 mg/L). Ovules less than 4.0 mm in size were not used in the experiment.
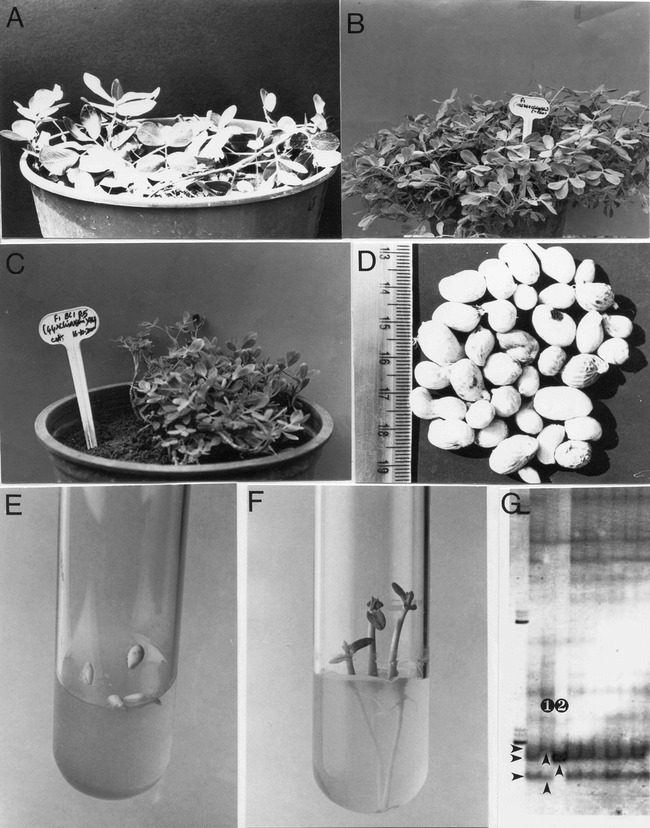
Crossability between A. hypogaea and A. chiquitana.
A. Plant of the wild species A. chiquitana.
B. F1 hybrid plant from the cross A. hypogaea × A. chiquitana.
C. BC1 hybrid plant of the cross A. hypogaea × A. chiquitana.
D. BC2 pods showing mostly single seeded and a few double seeded pods.
E. Large but immature embryos from the cross A. hypogaea × A. chiquitana on germination medium.
F. Germinating seedlings from the cross A. hypogaea × A. chiquitana.
G. SSR-3F1, fragment length 290 base pairs: analysis of the parents and the hybrids. From left to right: 1st lane: Molecular weight marker; 2nd lane: F1 hybrid, hybrid bands have been marked with arrows; 3rd lane: Arachis hypogaea, the female parent, marked no. 1; 4th lane: A. chiquitana, the male parent, marked no. 2; Lanes 5 to 10: F1 hybrids, hybrid bands have been marked with arrows.
Embryos germinated and gave rise to seedlings (Fig. 1F) with individual or multiple shoots. Shoots were rooted in vitro on rooting medium, consisting of ½ MS basal salts (plus sucrose), NAA (2.0 mg/L), indoleacetic acid (IAA; 1.0 mg/L). After 15 d on rooting medium, shoots were transferred to ½ MS basal medium (without growth regulators). Healthy roots developed within 3 wk of culture. Shoots with well-developed roots were transferred to sand and acclimatized under controlled conditions at 24 C and RH of 72–75%. A month of acclimatization was sufficient to transfer the plants to a glasshouse.
DNA was extracted from young, folded leaflets with Qiagen miniprep kits (Qiagen, Valencia, CA) and amplified by means of 30 pmol primer, 5 ng template DNA, 4 mM MgCL2, 0.2 mM dNTPs, 1 U Taq polymerase with 1 × reaction buffer in a total reaction volume of 20 µL. Reaction conditions were 94 C for 2 min, 35 cycles of 94 C for 45 sec, empirically defined annealing temperature of 60 C for 1 min, 72 C for 90 sec, then a final extension of 10 min at 72 C. Amplification products were visualized on non-denaturing 9% 29:1 (w/w) polyacrylamide gel followed by silver staining. Silver staining consisted of 3 min in H2O, 20 min in 0.1% (w/v) CTAB, 15 min in 0.3% ammonium solution, 15 min in a solution of 1 M NaOH, 0.1% silver nitrate and a few drops of 25% ammonium solution, and a rinse in H2O, and followed by development in a 1.5% NaC solution with 0.02% by volume formaldehyde solution. The SSR procedure and primers are described in detail in Ferguson et al. (2004). Amplification products were noted as present or absent. The four SSR primers used to distinguish the parents and the hybrids, were as follows:
-
pPGPseq3F1 Forward primer—AGCGATCAATCGGTTTCAAG, fragment length—290 bp. Reverse primer—GAAACGAAACGAAGACCGAA.
-
pPGPseq4D4 Forward primer-CGGCTGTTAGGTAATCAGTTCA, fragment length-187 bp. Reverse primer-TCAACAGGAATAGCTGCACG.
-
pPGPseq2A6 Forward primer-GCTTCTTCGTTGTTGCCTTC, fragment length-249 bp. Reverse primer-TGCCAGTTGTTCATAGCTTCA.
-
pPGPseq4H11 Forward primer-ATCACCATCAGAACGATCCC, fragment length-269 bp. Reverse primer-TTGTAGCCTTCTGGCGAGT.
The ploidy of the derivatives was determined by pollen diameter analysis. Three classes of pollen diameter were observed. Diploids had a diameter of 25 to 27 µM, triploids were 25 to 29 µM, and tetraploids had a diameter of 45 to 47 µM (Singsit and Ozias-Akins, 1992). Pollen diameter of A. hypogaea was between 45 to 47 µM and that of A. chiquitana was between 25 to 29 µM. Pollen in triploids which had undergone 2n restitution had 43 to 45 µM (and apparently fertile) was comparable to pollen grains of tetraploid plants.
Nineteen seeds were surface sterilized and were uniformly wounded by pricking with a sterile needle to allow the invasion of A. flavus spores. Seeds were spray inoculated with A. flavus spore suspension (1 × 106 spores/mL). Petri dishes with sprayed seeds were placed in high humidity and incubated at 25 C. Individual seeds were scored for A. flavus colonization, using a rating scale of 1 to 4 as described by Thakur et al. (2000).
Results
Pods from cross pollinations using A. chiquitana as the pollen parent appeared similar to A. hypogaea but were mostly single seeded. Thirty-four percent of the pollinations formed pods from the cross A. hypogaea × A. chiquitana. Aborted and dried seeds were obtained if harvested beyond 30 d. Forty-seven percent of the ovules (immature seeds) were large (> 4.0 × 2.5 mm), rest of the seeds less than 4.0 mm long were discarded. A total of 25 large ovules were dissected and embryos were cultured (Fig. 1E). Approximately 30 d was required for the embryos to respond to culture and develop into healthy seedlings (Fig. 1F) or form multiple shoots. If the hybrid pods with large ovules were allowed to dry, then they did not germinate in vivo or in vitro. Eleven hybrid plants were obtained (Fig. 1B). One plant did not reach the flowering stage, in spite of having normal vegetative growth. The hybrid plants had intermediate morphology with the leaf shape resembling that of A. chiquitana. All the 10 plants flowered profusely and pollen fertility ranged from 5 to 24%.
Hybrid plants were backcrossed using A. hypogaea as the pollen parent. It was possible to obtain mature BC1 pods from one hybrid plant, whereas the remaining plants did not set mature seeds. BC1 pods were mostly single seeded and abnormal in their shape. Twelve BC1 hybrid plants were obtained (Fig. 1C), from which two remained in the juvenile stage with little vegetative growth. Four of the plants had good vegetative growth but did not reach the flowering stage. Six hybrid plants flowered after propagation for at least 180 d. Pollen fertility in the BC1 hybrids ranged from 18 to 24%, and were tetraploids (2n = 4x = 40) which was confirmed by pollen diameter analysis. Mature BC2 seeds were obtained, but in small number, ranging from one to 11 seeds. Most of the pods were single seeded, and the few double seeded pods produced severe constriction between the seeds (Fig. 1D).
Four SSR markers showed polymorphism between the two parents. SSR 3F1 (Fig. 1G) showed heterozygosity between the two parents. Two unique bands were consistently seen in A. hypogaea, whereas A. chiquitana had one distinct band. All seven F1 hybrids used in the study showed the presence of bands from both the parents.
To check if it was possible to transfer resistance to A. flavus colonization, seeds were subjected to A. flavus colonization following the method described by Thakur et al. (2000). Large variation was observed with respect to fungal colonization. Individual seeds were scored for surface colonization, and the results showed six seeds which had the rating of 1 (resistant), and 12 seeds had the rating of 4 (susceptible). The other seeds showed a rating of 2 or 3. Since the method to test for the presence of aflatoxin is destructive to the seed (Devi et al., 2000), seeds were not sacrificed to test for the presence of aflatoxin. Once large numbers of seeds are produced, the logical next step will be to test seeds for A. flavus colonization and aflatoxin production.
Discussion
Wild species from section Arachis have been used in the improvement of peanut because they are compatible with cultivated peanut (Mallikarjuna et al., 2004). Although there have been various attempts in the past to cross wild species from other sections of Arachis, the attempts have not been successful because of barriers to crossability. Even if the barriers are overcome by the use of various in vivo or in vitro techniques, many times the hybrid plants are male and female sterile or they do not mature into the reproductive phase (Stalker and Simpson, 1995). Tissue culture techniques first developed to recover aborting embryos from crosses between A. hypogaea and A. glabrata (Mallikarjuna and Sastri, 2002), a wild species belonging to section Rhizomatosae, can be applied to recover hybrid embryos from other intersectional crosses (Mallikarjuna, 2002) as is evident from the present study. Crosses between A. hypogaea and A. paraguariensis (ICG 8130; PI 337350 KCF 11462) and A. hypogaea and A. appressipila (ICG8945; PI 468149; GK 30003) were obtained by the use of tissue culture techniques. The hybrids had profuse vegetative growth but did not reach reproductive stage (Mallikarjuna, unpubl. data), and were considered genetic dead-ends. The hybrids obtained in the present study as well as with other A. hypogaea intersectional hybrids indicate that a generalized statement cannot be made that all hybrids involving wild species outside section Arachis are genetic dead-ends.
It would be of interest to pursue crosses involving A. chiquitana because it is one of the few wild species that can be crossed with A. hypogaea to potentially transfer resistance to A. flavus. The other species that have resistance to A. flavus are A. pusilla and A. triseminata ( Thakur et al., 2000), A. duranensis and A. cardenesaii (Xue et. al., 2004), the latter two species are cross compatible with A. hypogaea.
The present study shows that the crossability barriers are not rigid and with the manipulation of embryo rescue techniques available for peanut, many more species from different sections may be successfully crossed with cultivated peanut.
Literature Cited
Ferguson M. E. , Burrow M. , Schulze S. R. , Bramel P. J. , Paterson A. H. , Kresovich S. , and Mitchell S. 2004 Microsatellite identification and characterization in peanut (A. hypogaea L.). Theor. Appl. Genet 108 : 1064 – 1070 .
Ghewande H. P. , Nagaraj G. , and Reddy P. S. 1989 Aflatoxin research at the National Research Center for Groundnut. 237 – 243 In McDonald D. and Mehan V. K. eds. Aflatoxin Contamination of Groundnut: Proc. Intl. Workshop, 6–9 Oct. 1987, ICRISAT, Patancheru, A.P., India
Gong Y. Y. , Cardwell K. , Hounsa A. , Egal S. , Turner P. C. , Hall A. J. , and Wild C. P. 2002 Dietary aflatoxin exposure and impared growth in young children from Benin and Togo: Cross sectional study. British Medical J 325 : 20 – 21 .
Gregory M. P. and Gregory W. C. 1979 Exotic germplasm of Arachis L. interspecific hybrids. J. Hered 70 : 185 – 193 .
Househam K. C. and Hunt H. K. 1991 Aflatoxin exposure and its relationship to kwashiorkor in African children. J. Tropical Pediatrics 37 : 300 – 302 .
Krapovikas A. and Gregory W. C. 1994 Taxonomia del genero Arachis (Leguminosae). Bonplandia 8 : 1 – 186 .
Mallikarjuna N. 2002 Gene introgression from A. glabrata into A. hypogaea, A. duranensis and A. diogoi. Euphytica 124 : 99 – 105 .
Mallikarjuna N. and Sastri D. C. 2002 Morphological, cytological and disease resistance studies of the intersectional hybrids between Arachis hypogaea and A. glabrata. Euphytica 126 : 161 – 167 .
Mallikarjuna N. , Suresh Pande , Jadhav D. R. , Sastri D. C. , and Rao J. N. 2004 Introgression of disease resistance genes from Arachis kempff-mercadoi into cultivated groundnut. Plant Breeding 123 : 573 – 576 .
Murashige T. and Skoog F. 1962 A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15 : 473 – 497 .
Pande S. and Rao J. N. 2001 Resistance of wild Arachis species to late leaf spot and rust in greenhouse trials. Plant Disease 85 : 851 – 855 .
Saghai-Maroof M. A. , Solinam K. M. , Jorgonsan R. A. , and Allard R. W. 1984 Ribosomal DNA spacer length polymorphism in barley. Mendelian inheritance, chromosomal location and population dynamics. Proc. Natl. Acad. Sci., (USA) 81 : 8014 – 8018 .
Stalker H. T. 1981 Hybrids in the genus Arachis between sections Erectoides and Arachis. Crop Sci 21 : 359 – 362 .
Stalker H. T. and Simpson C. E. 1995 Genetic Resources in Arachis. 14 – 53 In Pattee H. E. and Stalker H. T. eds. Advances in Peanut Science Amer. Peanut Res. Educ. Soc Stillwater, OK .
Thakur R. P. , Rao V. P. , Reddy S. V. , and Ferguson M. 2000 Evaluation of wild Arachis germplasm accessions for in vitro seed colonization and aflatoxin production by Aspergillus flavus. Intl Arachis Newsletter 20 : 44 – 46 .
Thirumala Devi K. , Mayo M. A. , Reddy G. , Reddy S. V. , Delfosse P. , and Reddy D. V. R. 2000 Production of polyclonal antibodies against ochratoxin A and its detection in chilies by ELISA. J. of Agric. and Food Chem 48 : 5079 – 5082 .
Waliyar F. , Ba A. , Hassan H. , Bonkoungou S. , and Bose J. P. 1994 Sources of resistance to Aspergillus flavus and aflatoxin contamination in groundnut in West Africa. Plant Disease 78 : 704 – 708 .
Williams J. G. K. , Livak A. R. , Rafalski J. A. , and Tingey S. V. 1990 DNA polymorphisms amplified by arbitary primers are useful as genetic markers. Nucleic Acids Res 18 : 6531 – 6535 .
Xue H. Q. , Isleib T. G. , Stalker H. T. , Payne G. A. , and Obrian G. 2005 Evaluation of Arachis species and Interspecific Tetraploid Lines for Resistance to Aflatoxin Production by Aspergillus flavus. Peanut Sci 31 : 134 – 141 .
Notes
- 1I thank Dr. R. Thakur and his group for screening the material for Aspergillus flavus colonization. I also acknowledge the technical assistance provided by Mr. Balakrishna and Mr. Satyanarayana in the crossing program. [^]
- 2International Crops Research Institute for Semi Arid Tropics (ICRISAT) Patancheru P.O. 502 324, Andhra Pradesh, India, (email: N.Mallikarjuna@CGIAR.ORG). [^]