INTRODUCTION
Successful reproduction and fruit set of most flowering plants, including valuable food crops and fruit trees that make up 35% of global food production, are dependent on animal pollination, primarily bee pollination (Klein et al., 2007). Pollinating insects can increase fruit or seed quality or quantity of many major crops (Klein et al., 2007). Yields of insect-pollinated crops are often managed by bringing hives of honeybees (Apis mellifera) to the fields to increase pollination. However, wild bee species (Bombus spp.) are also vital to our cropping systems. For over 40 globally important crops, wild pollinators improved pollination efficiency and increased fruit set compared to honeybees alone (Garibaldi et al., 2013). Floral strips can be used to increase plant biodiversity within and surrounding crop fields by providing diverse nutritional resources for wild bee species (Bombus spp.) and to increase pollination in agroecosystems as long as potential negative consequences of insecticide contact are considered (Garibaldi et al., 2013, 2021; Nicholls and Altieri, 2013; Blaauw and Isaacs, 2014). Better understanding pollination cues of major crops could help inform future decisions regarding conservation of wild pollinator abundance and diversity.
Flowers have adapted diverse advertising strategies to attract bees, including flower size, shape, scent, color, and pattern, which often correspond to morphological and sensory characteristics of pollinators. Bees can associate visual cues such as flower color and shape with nectar and pollen rewards and will use these cues to optimize foraging times (Spaethe et al., 2001). Bees are sensitive to ultraviolet (UV), blue, and green wavelengths, and their detection of large flowers (about 15 mm in diameter or larger) is dependent on high contrast between flower and green foliage background color (Spaethe et al., 2001). For detection of small flowers (around 8 mm in diameter or less), wild bees fly closer to the ground and use a different neuronal channel focused on green contrast only. The switch between neuronal channels is best exemplified by search times for yellow and UV-absorbing, white flowers, since yellow flowers have a higher color contrast to the background but about two-thirds the amount of green contrast. Spaethe et al. (2001) found that bumblebees (B. terrestris) took 22% longer to detect large, UV-absorbing white flowers than large, yellow flowers but 72% longer to identify small, yellow flowers than small, UV-absorbing white flowers due to using color contrast to identify large flowers and green contrast to identify small flowers. While flower detection of small and large flowers both involve color contrast, honeybees and wild bee species appear to ignore flower brightness during foraging (Vorobyev and Brandt, 1997; Spaethe et al., 2001).
While flower color is vital in attracting pollinators from a distance, flower color and UV patterns are important to bee orientation and landing (Papiorek et al., 2015). Many plants have adapted to display floral guides, often referred to as nectar guides, that are color and UV absorbance and reflectance flower patterns that direct pollinators to nectar and pollen to increase pollination success (Orbán and Plowright, 2013). In radially symmetric flowers with UV floral guides, generally apical parts of flower petals have UV reflecting carotenoids and the central flower parts have UV absorbing flavonoids creating a “bullseye” pattern (Thompson et al., 1972; Harborne and Smith, 1978). In addition to UV absorbing flavonoids, another type of UV absorbing pigment, dearomatized isoprenylated phloroglucinols, have been identified in some species on the anthers and ovarian wall that perform both a visual and defensive function, such as being a deterrent to insect feeding (Gronquist et al., 2001). While most radially symmetric flowers have radially symmetric floral guides, most bilaterally symmetric flowers have bilaterally symmetric floral guides (Dafni and Kevan, 1996). Whether radially or bilaterally symmetric, these floral guides orient pollinators toward the center of the flower where nectar and pollen are produced.
Peanut (Arachis hypogaea L.) is an annual, self-pollinated crop that is grown worldwide on about 25.9 million ha as an oil, food and feed crop (FAOSTAT, 2019). Like most crops, the predominant goal for peanut breeding is increased yield either directly through selection of genes that increase yield or indirectly such as through improved yield retention via increased biotic and abiotic stress tolerance and/or resistance. Wild Arachis species are important as donors of strong, durable resistances to cultivated peanut (Stalker, 2017). Previous reports found high bee visitation to peanut flowers and that visitation by large bee species that trip flowers increased peanut yields in some U.S. varieties (Girardeau and Leuck, 1967; Rashad et al., 1979). A more recent report found bees did not increase yield in newer peanut varieties grown in north Queensland, Australia, likely due to low bee visitation, and suggested that selection for other desirable traits may have resulted in development of varieties less attractive to flower-tripping bees (Blanche et al., 2006). There is no recent study testing the effect of bee pollination on peanut yield of U.S. cultivars. Most Arachis species, including cultivated peanut, are self-pollinating species and are not exclusively dependent on bee pollination for seed set. However, peanut has been documented as attractive to honeybees, many wild bee species, and other insect pollinators with reported cross-pollination rates between 1 to 10% (Culp et al., 1968; Knauft et al., 1992), despite lacking a nectary, meaning insect visitation is likely for pollen foraging (Vogel, 1997; Hammons and Leuck, 1966; Leuck and Hammons, 1969; Blache et al., 2005; USDA, 2015). However, there have been no studies in Arachis evaluating the nutritional quality and quantity of pollen as well as no study documenting the presence or absence of nectar using a nectar extraction method. Without these studies, it is unknown whether pollinators are benefitting from Arachis flowers or being manipulated into exerting time and energy to foraging reward-less flowers by false nectar guides, which are floral guides that attract pollinators to flowers that have no nectar nor quality pollen resource (Slater and Calder, 1988; Lunau and Webster, 2017). If Arachis species are found to have nutritious pollen, high-density flowering species such as A. glabrata, A. repens, and A. pintoi that are used as summer legume forage and cover crops and as urban lawns, could be beneficial for increasing plant biodiversity to provide habitats for wild bees in urban landscapes (Anderson et al., 2015; Kröning et al., 2019).
Despite the documentation of bee-pollination in Arachis and the extensive morphological characterization of Arachis that has been performed, UV reflectance pattern diversity in cultivated peanut and other species in the Arachis genus has not been documented. Since Arachis species generally develop yellow and orange flowers, and yellow flowers are more likely than other colored flowers to contain UV floral guides (Horovitz and Cohen, 1972; Guldberg and Atsatt, 1975; Primack, 1982), it is likely that peanut species have UV reflectance patterns. Furthermore, given the wide pool of genetic and phenotypic variation in the Arachis genus (Kochert et al., 1996; Moretzsohn et al., 2005, 2013), different species may have distinct flower UV reflectance patterns. This study documented flower UV absorbance and UV reflectance levels and patterns as well as absorption levels at visible-light wavelengths, including at blue and green wavelengths, in cultivated peanut, diploid Arachis species, wild Arachis species-derived allotetraploid interspecific hybrids that are cross-compatible to peanut, and BC1F3 hybrids derived from crosses between cultivated peanut and allotetraploids. Our results indicate that UV floral guides of various sizes are present in wild Arachis species, allotetraploids, and A. hypogaea genotypes.
Materials and Methods
Plant Materials
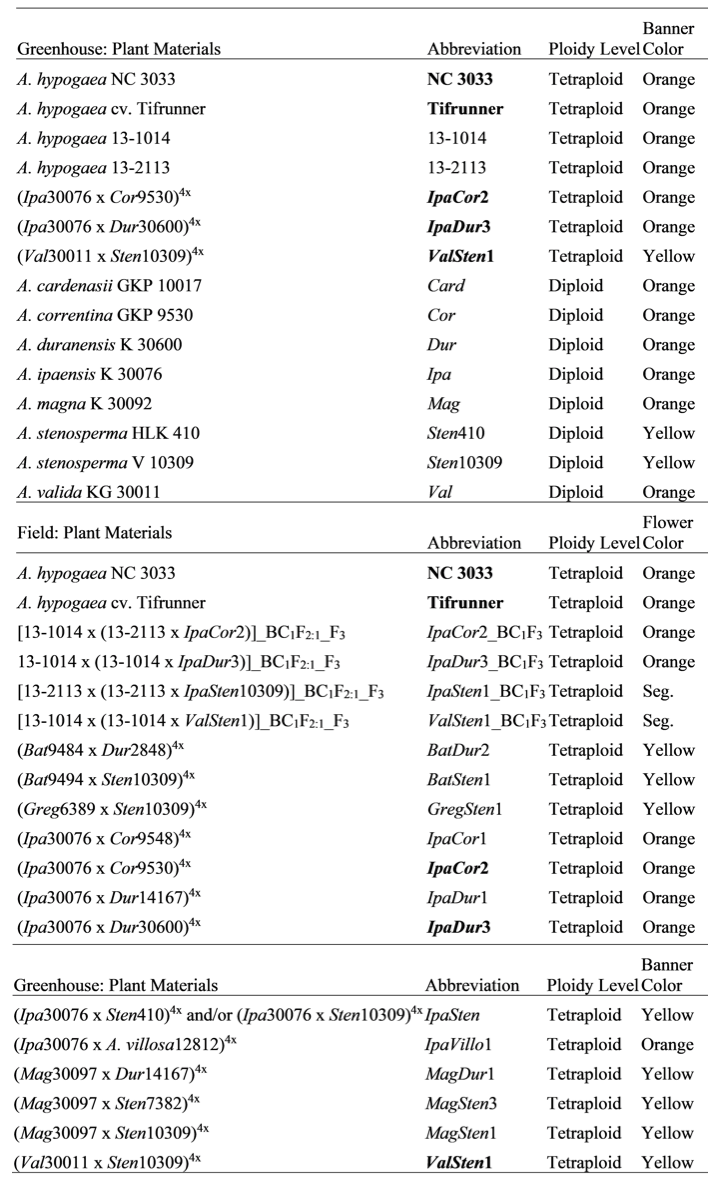
Plant materials tested in this study and their nomenclature are shown in Table 1. Diploid, wild Arachis species, A. correntina, A. duranensis, and A. ipaensis were used to generate the diploid hybrids, IpaCor2 and IpaDur3 in 2016 at North Carolina State University (NCSU). Wild Arachis species, A. batizocoi Krapov. and W.C. Gregory (PI 298639, K 9484, abbrev.: Bat9484), A. correntina (PI 262881, GKP 9548, abbrev.: Cor9548), A. duranensis Krapov. and W.C. Gregory (V 14167, abbrev.: Dur14167 and SeSn 2848, abbrev.: Dur2848), A. gregoryi A. Gripp, C.E. Simpson, and J.F.M. Valls (PI 476116, VSGr 6389, abbrev.: Greg6389), A. magna Krapov., W.C. Gregory, and C.E. Simpson (PI 468340, K 30097, abbrev.: Mag30097), A. stenosperma Krapov. and W.C. Gregory (V 10309, abbrev.: Sten10309 and PI 338280, HLK 410, abbrev.: Sten410), A. valida Krapov. and W.C. Gregory (PI 468154, GK 30011, abbrev.: Val30011), and A. villosa Benth. (V 12812, abbrev.: Villo12812) were used to create diploid hybrids BatDur2, BatSten1, GregSten1, IpaCor1, IpaDur1, IpaSten, IpaVillo1, MagDur1, MagSten3, MagSten1, and ValSten1 at the University of Georgia (UGA) Athens Campus. All allotetraploids were derived from the diploid hybrids by colchicine treatment of F1 hybrid cuttings at the UGA Athens Campus, except for IpaDur3 and IpaCor2, which were generated at the UGA Tifton Campus. IpaCor2, IpaDur3, IpaSten, and ValSten1 allotetraploids, used as pollen donors, were crossed to cultivated peanut breeding lines, A. hypogaea ‘13-2113’ or ‘13-1014’ and subsequently backcrossed to these breeding lines (Table 1). Breeding lines 13-2113 and 13-1014 were selected from [(C1805-617-2 x ‘Florida-07’ (Gorbet and Tillman, 2009)) x ‘Georgia-06G’ (Branch, 2007)], in which C1805-617-2 was a selection from ‘Tifguard’ (Holbrook et al., 2008) x Florida-07. Line 13-2113 was selected with the ADSNP124 (A09 6720287) marker (Chu et al., 2016) to have an A09 A. cardenasii introgression that confers nematode resistance.
Genetic materials tested and their abbreviations, ploidy level, and visible light flower banner color, in which “Orange” refers to banners with a yellow center and orange edge, yellow refers to completely yellow banners, and seg. refers to BC1F3 genotypes that are segregating for banner color, in which plants had flowers with either orange or yellow banners. Bolded genotypes were grown in the greenhouse and field.
UV reflectance evaluation was performed on plant materials grown in two different environments, field and greenhouse, in which two A. hypogaea genotypes and three allotetraploids were grown in both environments. The greenhouse materials included two A. hypogaea genotypes ‘NC 3033’ (Beute et al., 1976) and ‘Tifrunner’ (Holbrook and Culbreath, 2007), three allotetraploids, and eight wild Arachis species. The field grown materials included the A. hypogaea genotypes, four BC1F3 lines derived from four unique allotetraploids, and 13 allotetraploids. The Ipa30076Sten410 and Ipa30076Sten10309 seeds were accidently mixed, but since both hybrids represent the same species cross, the data were retained in the analyses. However, it should be noted that IpaSten data may come from plants from either or both allotetraploids (Table 1).
For the field study, seeds from the 13 allotetraploid lines were coated in Vitavax PC on 5 May 2020 (Vitavax, Crompton, Middlebury, CT) and treated overnight in 0.5% Florel Growth Regulator (Lawn and Garden products Inc., Fresno, CA) on 9 cm diameter Whatman No. 1 filter paper (ThermoFisher Scientific) in 100 mm x 15 mm Petri dishes (ThermoFisher Scientific) to break dormancy. This process was repeated with seed from the two A. hypogaea genotypes, NC 3033 and Tifrunner, 13 days later to account for the slower growth of the allotetraploids as compared to the A. hypogaea genotypes. Twenty-four hours later, seeds were planted in #123 7.62 cm round x 11.43 cm deep Jiffy pots (Harris Seeds, Rochester, NY) filled with Promix growth medium (Premier Tech Horticulture, Quakertown, PA). On 2 June, all genotypes were transplanted from the greenhouse to the Black Shank farm in Tift County, GA, in a randomized complete block design with three replications of a single plant of each genotype. Due to limited field space, IpaCor2_BC1F3, IpaDur3_BC1F3, IpaSten1_ BC1F3, and ValSten1_ BC1F3 seeds were coated in Vitavax PC and directly seeded into plots at the Rigdon farm, located one mile from the Black Shank farm, on 1 June 2020. Both fields were tilled, deep-turned and bedded before planting. The field at the Black Shank farm was treated with a 14-day fungicide program of chlorothalonil (1.26 kg/ha) to control foliar diseases. In contrast, fungicides were withheld from the field management plan at the Rigdon Farm.
On 20 July, seeds of all genotypes for greenhouse cultivation were coated in Vitavax PC and treated overnight in 0.5% Florel Growth Regulator to break dormancy. Seeds were then planted in #123 7.62 cm round x 11.43 cm deep Jiffy Pots and transplanted approximately one month later into 121.92 cm round x 27.94 cm deep pots filled with Promix growth medium. Plants in the greenhouse were treated with 50% bifenazate to control for spider mites.
Ultraviolet photography
A Nikon D 5000 digital camera was used to take photographs in a greenhouse in natural light. The UV/IR cut color correcting hot mirror filter was removed from the camera by Kolari Vision (New Jersey, USA) to make it sensitive to UV, visible, and IR light. The camera was equipped with a Baader U-Venus-Filter 2" (350 nm) and an adapter Baader Hyperion DT Ring HDT54/52 (M54 to M52) # 2958052 (Mammendorf, Germany), which allowed transmission of UV light from 320 to 380 nm and blocked the rest of the spectral range from 200 to 320 nm and from 380 to 1,120 nm.
Flower Characterization
Between 78 and 92 days after transplanting, four to six flowers from three plants of each genotype (15 flowers per genotype total) at the Black Shank farm were collected in 15 ml falcon tubes (Corning CoStar, Corning, NY) containing a moist Kimwipe (Kimerly-Clark, Neenah, WI) to keep flowers from wilting during collection. On 109 days after planting, five flowers from three plants of each BC1F3 line were collected. Between 112 and 221 days after planting, five flowers from three plants of each genotype grown in the greenhouse were collected. All collections were conducted between 8:30 to 10:30 am. The greenhouse collection period was protracted mostly due to delayed flowering in some wild Arachis species. NC 3033, Tifrunner, and Ipa only had flowers collected from two plants, while Val only had flowers collected from one plant. Flowers were dissected and photographed; measurements of banner area (cm2), area of UV absorbance on banner (abaxial and adaxial) (cm2), left-wing area (cm2), and area of UV absorbance on left-wing (abaxial and adaxial) (cm2) were measured with Assess 2.0 (APS Press) (Fig. S1). Identical analysis was performed on the right-wing but was not reported due to high similarity to left-wing results. Each image was calibrated by using rulers photographed in each image. Measurements of flowers collected from the same plant were not independent observations since, plant, not flower, was the experimental unit. Therefore, the measurements for the flowers per plant were averaged to make one data point, meaning each genotype had three data points per tested parameter. Lastly, flowers collected from the greenhouse were scanned to obtain regular, non-UV, photographs of these materials.
The flower banners dissected for UV photography from plants grown in the greenhouse were used to measure banner pigmentation. The banners were first weighed before they were put into 2 ml tubes (Phenix Research, Swedesboro, NJ) with 4, 3.2 mm diameter, beads (BioSpec, Bartlesville, OK). The tubes were then submerged in liquid nitrogen and vortexed until the banner was a fine powder. The volume of 100% ethanol added to the pulverized banner was 10 times the banner weight. The tubes were vortexed for approximately 3 s and then spun down at 3000 rpm for 5 min. Two hundred ul of the supernatant was loaded into a spectrometry plate, with 100% ethanol blanks as controls. Readings were normalized to the 100% ethanol, and only readings at wavelengths of 380, 415, 450, 473 and 495 nm were above the blank ethanol. Banner pigmentation was not measured for NC 3033, Tifrunner, or Val.
Statistical Analysis
One-way analysis of variance (ANOVA) was performed using RStudio (RStudio, Inc.) to determine the effect of the two different environments (field environment collected between 18 August to 18 September 2020 and greenhouse environment collected from 9 November 2020 to 26 February 2021) on UV reflection in Arachis flowers according to the following parameters: area of UV absorption on the banner (abaxial and adaxial), area of UV absorption on the banner as a percentage of total banner area (abaxial and adaxial), area of UV absorption on the left-wing (abaxial and adaxial), area of UV absorption on the left-wing as a percentage of total left-wing area (abaxial and adaxial). ANOVA was also performed to determine the effect of the two different environments on banner area and left-wing area. Means of each parameter among the genotypes were separated based on the Tukey’s Test (α = 0.05) results with RStudio. Only genotypes grown in both the field and greenhouse were included in ANOVA and Tukey’s analysis, including NC 3033, Tifrunner, IpaCor2, IpaDur3, and ValSten1.
ANOVA was performed using RStudio (RStudio, Inc.) to determine the effect of genotype on flower size in Arachis and on UV reflection in Arachis flowers according to the following parameters: banner area, area of UV absorption on the banner (abaxial and adaxial), area of UV absorption on the banner as a percentage of total banner area (abaxial and adaxial), area of UV absorption on the adaxial as well as the abaxial side of the left-wing (abaxial and adaxial), area of UV absorption on the adaxial as well as abaxial left-wing as a percentage of total left-wing area (abaxial and adaxial). Means of each parameter among the genotypes were separated based on the Tukey’s Test (α = 0.05) results with RStudio. ANOVA and subsequent Tukey’s analysis were performed separately on materials from the greenhouse and materials from the field. ANOVA was also performed to determine the effect of genotype on banner pigment at wavelengths 380, 415, 450, 473 and 495 nm (the wavelengths that had readings above the blank ethanol control) followed by Tukey’s test to separate the means of each parameter among the genotypes.
Results
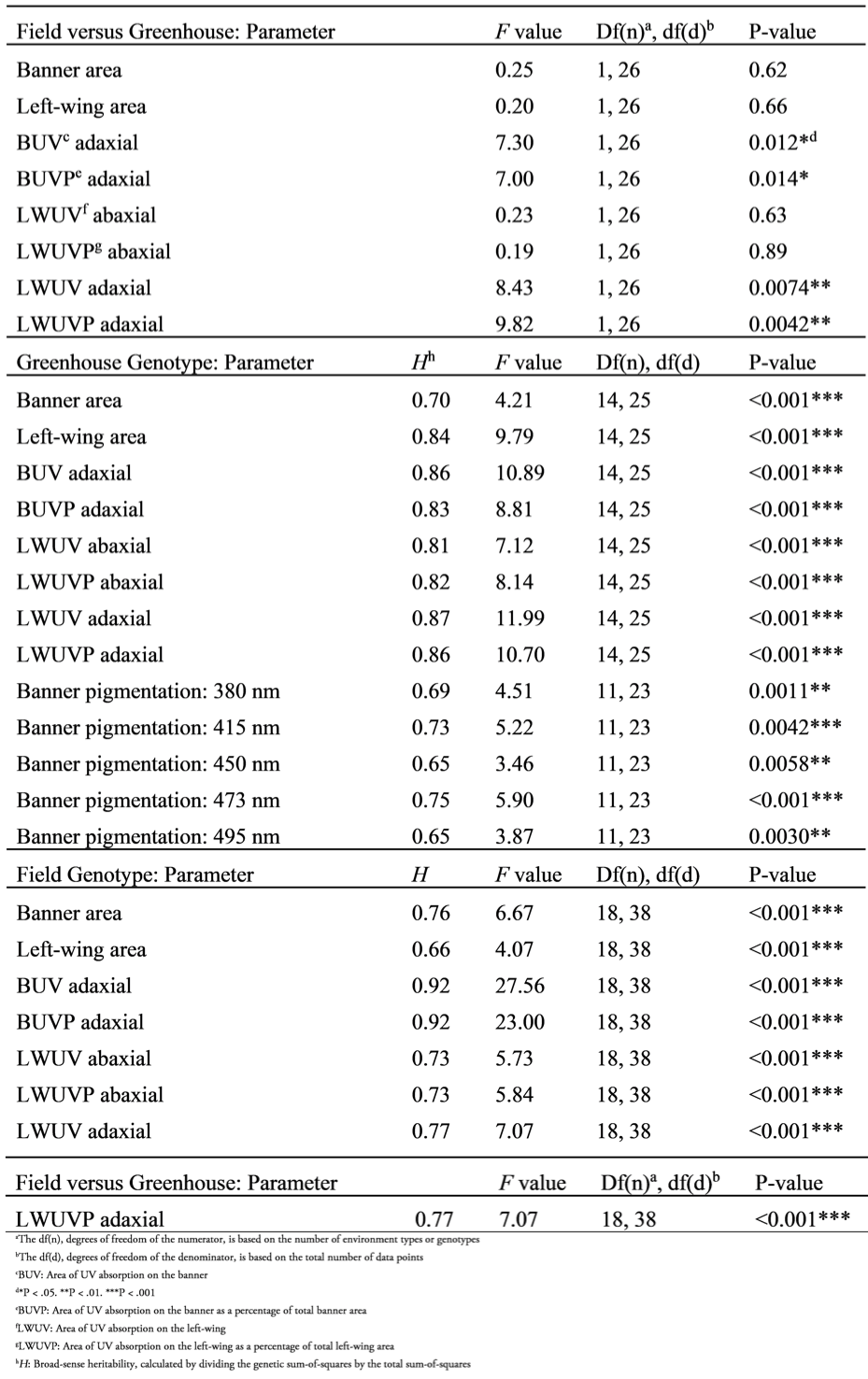
Field versus greenhouse environments had a significant effect on area of UV absorption on the banner (adaxial), area of UV absorption on the banner as a percentage of banner area (adaxial), area of UV absorption on the left-wing (adaxial), and area of UV absorption on the left-wing as a percentage of left-wing area (adaxial) (Table 2). No environmental effect was observed on banner area, left-wing area, area of UV absorption on the left-wing (abaxial), nor area of UV absorption on the left-wing as a percent of left-wing area (abaxial). In addition, no flowers collected from the field nor the greenhouse had UV absorption on the abaxial side of the banner. Flowers from the field had greater total area of UV absorption on the banner (adaxial) and area of UV absorption as a percent of banner area (adaxial) than flowers from the greenhouse (Fig. S2). Similarly, flowers from the field had greater total area of UV absorption on the left-wing (adaxial) and area of UV absorption as a percent of left-wing area (adaxial) than flowers from the greenhouse (Fig. S3). The UV absorption pattern on the adaxial side of the banner remained consistent within each genotype (Fig. S4). The A. hypogaea genotypes NC 3033 and Tifrunner had UV absorption areas on the banner that were shaped similarly to a vertical cross section of a mushroom while the allotetraploids had heart-shaped cross-sections whether grown in the field or greenhouse (Fig. S4).
ANOVA output testing the effect of the two different environments, on flower size in Arachis and UV absorption in Arachis flowers according to the following parameters: banner area, left-wing area, area of UV absorption on the banner (adaxial), area of UV absorption on the banner as a percentage of total banner area (adaxial), area of UV absorption on the left-wing (abaxial and adaxial), and area of UV absorption on the left-wing as a percentage of total left-wing area (abaxial and adaxial). The second and third sections of the table show ANOVA output testing the genotype effect of greenhouse plants and field plants, respectively, according to the aforementioned parameters. The second section also includes ANOVA output testing the genotype effect on banner pigmentation.
Genotype had a significant effect on banner area, left-wing area, and all parameters of UV reflection on Arachis flowers collected from the greenhouse (Table 2). IpaCor2 and Cor had the largest banner area at 2.11 and 2.23 cm2, respectively, which were numerically larger than the A. hypogaea genotypes but only statistically larger than Dur, Mag, and Sten10309, which had banner areas of 0.68, 0.96, and 0.97 cm2, respectively (Fig. S5A). The only genotype with significantly different banner area from A. hypogaea was Dur. IpaCor2 and Cor had the largest left-wing area at 0.51 and 0.56 cm2, respectively, and the latter was significantly greater than all four A. hypogaea genotypes (Fig. S5B). Only IpaCor2 and Cor were significantly different from at least one A. hypogaea genotype for left-wing area. All the genotypes had yellow wings, but only ValSten1, Sten410, and Sten10309 had completely yellow banners (Fig. S6). The remaining genotypes had banners with a yellow center and orange edge, though the intensity of the orange varied. All genotypes except Ipa, Mag, and two of the three Cor plants had thin red stripes originating from where the hypanthium and banner connect and spread outwards on their banners.
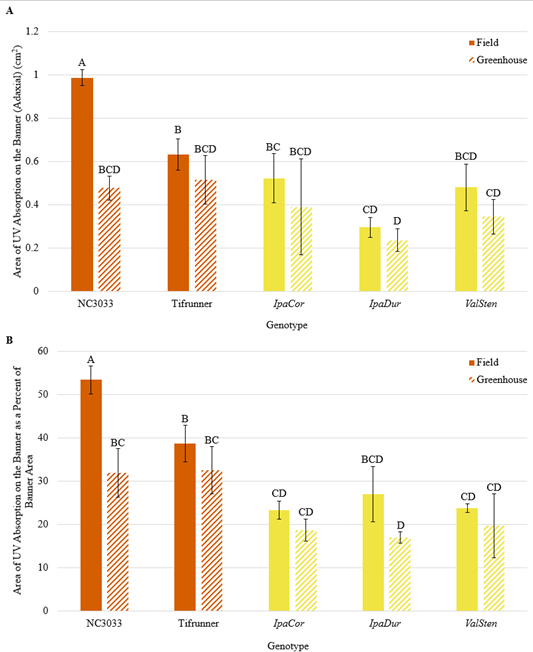
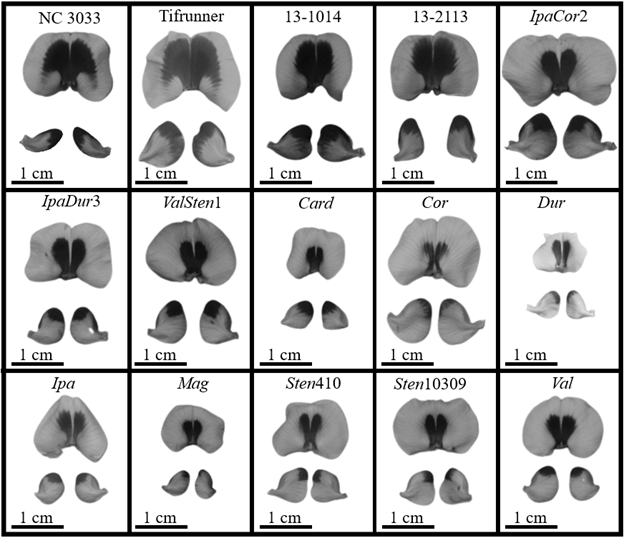
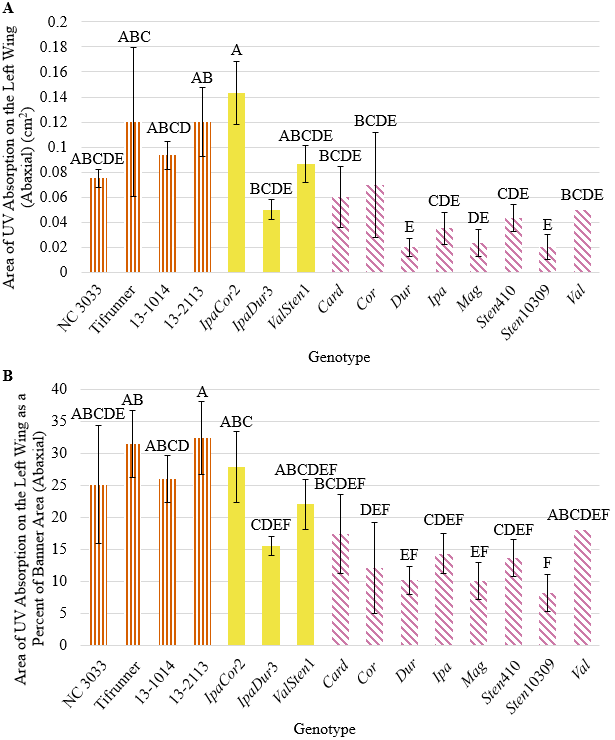
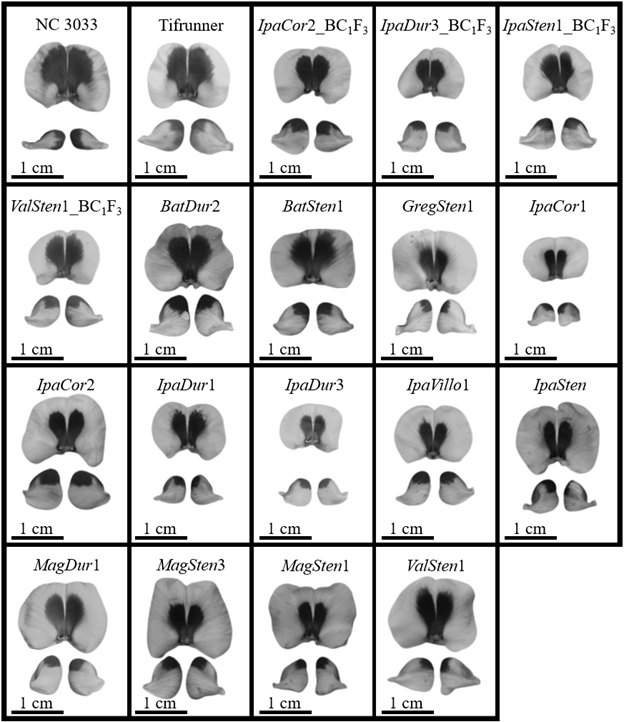
Numerically, the four A. hypogaea genotypes had the largest total area of UV absorption on the adaxial side of the banner as well as the largest UV absorption area as a percent of banner area (Fig. 1). 13-2113 had the largest total area of UV absorption at 0.56 cm2 and Tifrunner had the largest UV absorption as a percent of banner area at 32.5%. The total area of UV absorption of the four A. hypogaea genotypes was not significantly larger than the allotetraploids with the exception of Tifrunner and 13-2113 being significantly larger than IpaDur3; however, they were significantly larger than all the diploid, wild Arachis species except Val. All the wild Arachis species had total UV absorption areas smaller than half of the total UV absorption areas of the A. hypogaea genotypes. The areas of UV absorption as a percentage of banner area of the four A. hypogaea genotypes were not significantly larger than the allotetraploids IpaCor2 and ValSten1, but most were significantly larger than IpaDur3. Similar to total UV absorption, the areas of UV absorption as a percent of total banner area of the A. hypogaea genotypes were significantly larger than most wild Arachis species. As previously mentioned, NC 3033 and Tifrunner have mushroom-shaped UV absorption areas with wings on the widest section of the UV absorption area that extended downwards towards the hypanthium (Fig. 2). These characteristic wings were absent in 13-1014 and 13-2113, which had a more intermediate UV pattern between the mushroom shape and the heart shape of the allotetraploids and some wild species (Fig. 2).
While the A. hypoagaea genotypes had larger total UV absorption on the abaxial side of the left-wing than all the wild Arachis species, IpaCor2 had the largest UV absorption area of 0.14 cm2 (Fig. 3A). When accounting for left-wing area, 13-2113 and Tifrunner had the largest percent of UV absorption at 32.37 and 31.44%, respectively, and then IpaCor2 had the next largest at 27.84% (Fig. 3B). The percent UV absorption of the four A. hypogaea genotypes was larger than that of IpaDur3, ValSten1, and all the wild Arachis species, although only 13-2113 and Tifrunner were significantly larger than the majority of the wild Arachis species. NC 3033 had very narrow wings, distinctive from the rest of the genotypes, with UV absorption on the distal tips of the wings; the distinctive wing shape of NC 3033 makes it difficult to compare the UV absorption shape with the other genotypes (Fig. 2). Tifrunner and 13-1014 had UV absorption areas that ran vertically along the entire distal-most edge of the left- and right-wing, while the other genotypes had UV absorption just on the upper tip of the distal-most edge (Fig. 2).
Figure 3. (A) Area of UV absorption on the left-wing (abaxial) and (B) area of UV absorption as a percentage of left-wing area (abaxial) of A. hypogaea genotypes (orange, vertical stripes), allotetraploids (solid, yellow), and wild Arachis species (horizontal, pink stripes) collected from the greenhouse.
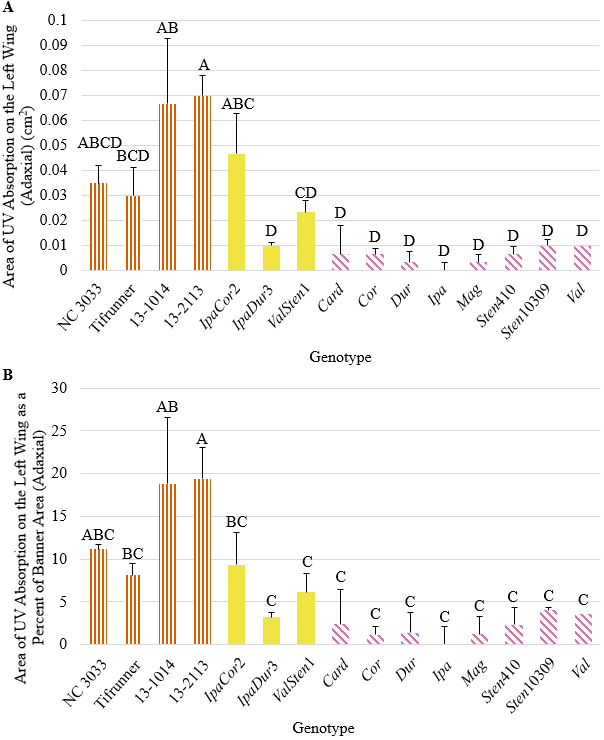
All genotypes had areas of UV absorption on the adaxial side of the left-wing less than half the area of UV absorption on the abaxial side of the wing, except for the breeding lines 13-1014 and 13-2113, in which the former had a mean adaxial UV absorption area of about three-fourths that of the abaxial UV absorption area (Fig. 4). 13-1014, 13-2113, and IpaCor2 had significantly larger total area of UV absorption than all the other allotetraploids and wild Arachis species, but when accounting for wing area, only 13-1014 and 13-2113 had significantly larger percentages of UV as compared to IpaDur3, ValSten1, and all the wild Arachis species. While smaller, UV absorption area on the adaxial side of the wings was located in the same position as on the abaxial side of the wings, in the upper tip of the wings (Fig. S7). For 13-1014 and 13-2113, adaxial UV absorption did not run the entire distal-most edge like on the abaxial side but was present only in the upper region of the wing. Once dissected, the wings laid flat for the UV photographs, except the wings of NC 3033, which curve inwards towards the keel and reproductive organs (Fig. S7).
Figure 4. (A) Area of UV absorption on the left-wing (adaxial) and (B) area of UV absorption as a percentage of left-wing area (adaxial) of A. hypogaea genotypes (orange, vertical stripes), allotetraploids (solid, yellow), and wild Arachis species (horizontal, pink stripes) collected from the greenhouse.
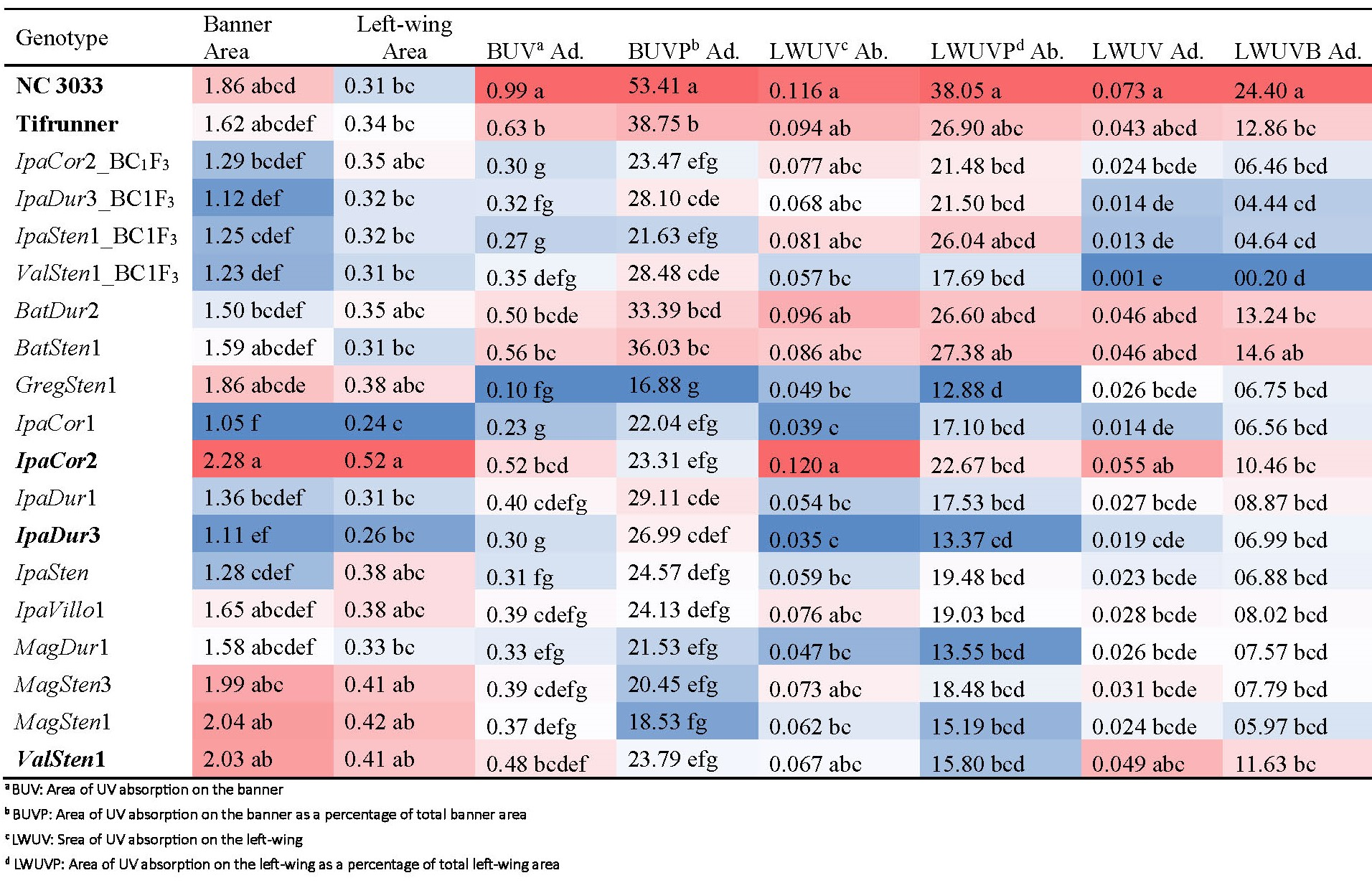
Genotype had a significant effect on banner area, left-wing area, and all parameters of UV reflection on Arachis flowers collected from the field (Table 2). As with the greenhouse collected flowers, IpaCor2 had the largest banner of 2.28 cm2 but was only numerically larger than NC 3033 and Tifrunner, which had banner areas of 1.86 and 1.62 cm2, respectively (Table 3). The next three genotypes, MagSten1, ValSten1, and MagSten3 with the largest banners all were progenies of Sten10309 or Sten7382. Despite IpaCor2 having the largest banner area, IpaCor1 had the smallest at 1.05 cm2. The BC1F3 lines had similar average banner sizes ranging from 1.29 to 1.12 cm2 for IpaCor2_BC1F3 and IpaDur3_BC1F3, respectively. IpaCor2 had a significantly larger left-wing area of 0.52 cm2 as compared to NC 3033 and Tifrunner, which had areas of 0.31 and 0.34 cm2, respectively. Similar to banner area, MagSten1, MagSten3, and ValSten1 had the next largest left-wing area after IpaCor2, and IpaCor1 had the smallest banner area.
Genotypes grown in the field and their means for banner area, left-wing area, area of UV absorption on the banner (adaxial) and on the left-wing (abaxial and adaxial), and area of UV absorption on the banner as a percentage of total banner area (adaxial) and on the left-wing as a percent of total left-wing area (adaxial and abaxial) and their Tukey’s HSD level (α = 0.05). Bolded genotypes were also grown in the greenhouse. Conditional formatting was applied within each column in Excel adding a color gradient to the values, in which red values are larger and blue values are smaller, to visualize the range of values more easily.
A. hypogaea had significantly larger total UV absorption areas on the adaxial side of their banners of 0.99 cm2 for NC 3033 and 0.63 cm2 for Tifrunner as compared to the BC1F3 genotypes and the majority of the allotetraploids (Table 3). When banner area was accounted for, the percent of UV absorption of 53.41% for NC 3033 and 38.75% for Tifrunner were significantly greater than for all the BC1F3 genotypes and allotetraploids, except Tifrunner was similar to BatDur2 and BatSten1. GregSten1 had the smallest total UV absorption on the adaxial side of the banner at 0.10 cm2 and the smallest percent of UV absorption at 16.88%. BC1F3 individuals derived from the four allotetraploids and 13-2113 and 13-1014 had UV absorption areas on the adaxial side of the banner shaped similarly to flowers of 13-2113 and 13-1014 collected from the greenhouse (Fig. 2, 5). While variations in size were found, the allotetraploids all had centralized, heart-shaped UV absorption areas that were narrowest proximal to the hypanthium (Fig. 5).
IpaCor2 and NC 3033 had total UV absorption areas of 0.120 and 0.116 cm2, respectively, on the abaxial side of the left-wing that were significantly larger than most of the allotetraploids, but not significantly larger than three of the four BC1F3 genotypes (Table 3). IpaDur3 and IpaCor1 had the smallest areas of UV absorption on their left-wings at 0.035 and 0.039 cm2, respectively. When accounting for left-wing area, NC 3033 and Tifrunner had the highest percent of UV absorption at 38.05 and 26.90%, respectively, and the percent of NC 3033 was significantly larger than IpaCor2. Only BatSten1 and BatDur2 had similar percentages of UV absorption on the abaxial side of the left-wing as compared to NC 3033. As was found with flowers collected from the allotetraploids in the greenhouse, the UV absorption of the BC1F3 flowers and allotetraploid flowers was confined to the distal tip of the wings.
All genotypes had areas of UV absorption on the adaxial side of the left-wing less than that on the abaxial side (Table 3). NC 3033 had the greatest area of UV absorption on the adaxial side of the left-wing at 0.073 cm2, significantly greater than all the BC1F3 genotypes and most of the allotetraploids, but not significantly greater than the peanut cultivar, Tifrunner, which had an area of 0.043 cm2. When accounting for left-wing area, NC 3033 had the highest percent of UV absorption on the adaxial side of the left-wing at 24.40%, significantly greater than all other genotypes except BatSten1, which had an UV absorption of 14.6%. The three genotypes with the smallest total and percent of UV absorption on the adaxial side of the wings were IpaDur3_BC1F3, IpaSten1_BC1F3, and ValSten1_BC1F3. Furthermore, ValSten1_BC1F3 had almost no UV absorption on the abaxial side of the left-wing, with a mean UV absorption area of 0.001 cm2. While smaller, UV absorption area on the adaxial side of the wings was located in the same place as on the abaxial side of the wings, in the upper tip of the wings (Fig. S8).
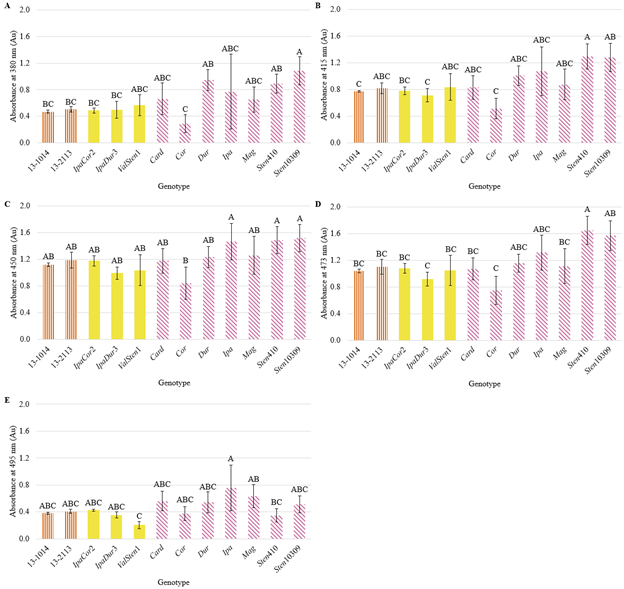
Significant genotypic effects on banner pigment at the following wavelengths were found: 380, 415, 450, 473, and 495 nm (Table 2). The peanut breeding lines and the allotetraploids, IpaCor2, IpaDur3, and ValSten1, had the same level of absorbance at all wavelengths (Fig. 6). Most wild Arachis species were not significantly different from the peanut breeding lines, except for Sten410, which had significantly higher absorbance at 415 (violet) and 473 nm (cyan), and Sten10309, which had significantly higher absorption at 380 nm (ultraviolet-A). Cor had the lowest absorbance at all wavelengths except for 495 nm (cyan/green), in which ValSten1 had the lowest absorbance. Sten410 and/or Sten10309 had the highest absorbance at all wavelengths except for 495 nm, in which Ipa had the highest absorbance.
Variation among genotypes for flower size and UV reflection in Arachis flowers according to all measured parameters was relatively more important than variation within genotypes given by the relatively high broad-sense heritability values, since error in the models was due to within-genotype variation (Table 2). For flowers grown in the greenhouse, UV reflection parameters had broad-sense heritability values that ranged from 0.81 for total UV absorbance on abaxial side of the left-wing to 0.87 for total UV absorbance on adaxial side of the left-wing. For flowers grown in the field, UV reflection parameters had broad-sense heritability that ranged from 0.73 for total and percent UV absorbance on abaxial side of the left-wing to 0.92 for total and percent UV absorbance on the banner. Among-genotype variation was also more important for determining banner pigmentation at 380, 415, 450, 473, and 495 nm than within-genotypes as indicated by broad-sense heritability values ranging between 0.62 to 0.74 (Table 2).
Discussion
Arachis species, including peanut, may provide pollen for wild bee species; however, visual cues that bees can associate with pollen rewards, such as contrasting color and UV absorption and reflectance on flowers, have not been characterized in Arachis. Since yellow flowers are more likely than other colored flowers to contain UV floral guides (Horovitz and Cohen 1972; Guldberg and Atsatt 1975; Primack, 1982) and Arachis species generally have yellow or orange and yellow flowers, we hypothesized that peanut and Arachis species would have UV reflectance patterns. This study found that all genotypes, including four A. hypogaea genotypes, eight wild Arachis species, 19 unique allotetraploids, and four BC1F3 lines, had UV floral guides. These UV floral guides were bilaterally symmetrical consistent with previous findings that bilaterally symmetric flowers have bilaterally symmetric floral guides (Dafni and Kevan, 1996). All genotypes had UV absorption located centrally on the banner with UV absorption extending towards the junction of banner and hypanthium and had UV absorption on the distal tips of the abaxial and adaxial sides of the wings.
While all studied genotypes had UV floral guides, the size and shape of these floral guides varied. In general, A. hypogaea genotypes had greater total UV absorption and UV absorption as a percent of banner area than the wild Arachis species. This same trend occurred for UV absorption on the abaxial side of the wings. This complements the finding of Horth et al. (2014), who found that mean UV floral guide size was smaller in wild populations as compared to cultivated populations of Rudbeckia hirta. It is unknown if larger UV absorption on the adaxial side of the banner and abaxial side of the wings increases or decreases Arachis flower conspicuousness for detection by bees or if it affects bee landing and orientation after detection. In other species, when UV floral guides are completely obscured in both radially and bilaterally symmetric flowers, bee visitation decreased (Peter and Johnson, 2008; Klomberg et al., 2019). However, the effect of altering the size of UV absorption and reflectance without completely eliminating the UV pattern on bee visitation frequency has not been extensively studied, especially in bilaterally symmetrical flowers. In radially symmetrical flowers, increasing the area of UV absorbance of Hypoxis camerooniana flowers did not affect bee visitation, while increasing the area of UV absorbance of R. hirta, R. flugida, and Argentina anserina flowers did increase bee visitation (Horth et al., 2014; Rae et al., 2013; Klomberg et al., 2019).
Increased UV absorption on the banner of cultivated peanut flowers as compared to wild Arachis species flowers may in part be a consequence of polyploidy or an evolved trait. Allotetraploidization of Arachis species increases leaf photosynthetic pigment production (Leal-Bertioli et al., 2017), but its effect on flower pigment production has not been studied. In this study, IpaDur3, IpaCor2, and ValSten1 and the diploid, wild Arachis species from which they were derived were tested in the greenhouse environment. In all comparisons, the allotetraploids had greater mean total UV absorption and UV absorption as a percent of banner and greater mean total UV absorption and UV absorption as a percent of left-wing (adaxial and abaxial) than the wild Arachis species from which they were derived, except IpaDur3 which had a smaller UV absorption area as a percent of banner area than Dur. Though these increases in UV area were only significant for IpaCor2 as compared to Ipa and Cor for UV on the adaxial and abaxial side of the left-wing, this trend suggests polyploidization may increase flower UV pigment production. A follow-up study with greater genotypic replication may yield significant differences between additional allotetraploids and diploid, wild Arachis species to further test if polyploidization increases flower UV pigment production.
Larger UV absorption areas in A. hypogaea could be an evolved trait. All Arachis genotypes had “large” flowers, greater than 8 mm in diameter as defined by Spaethe et al. (2001), meaning bees likely use the color-contrast neuronal channel visualizing blue, green, and UV wavelengths to detect Arachis flowers. Therefore, flowers with a greater contrast to a foliage background color should be most conspicuous to bees. A small area of UV absorption surrounded by a large area of UV reflectance characteristic of wild Arachis species may stand out more in UV-absorbing foliage as compared to flowers with a large area of UV absorption surrounded by a small area of UV reflectance more characteristic of the observed A. hypogaea genotypes (Fig. S9). Further studies elucidating the relationship between UV absorption and UV reflection area with bee flower detection and visitation on bilaterally symmetric flowers are needed.
A study to determine the effect of UV absorption area on bee visitation in Arachis could use UV-absorbing chemicals such as butyl methoxydibenzoylmethane and ethylhexyl methoxycinnamate to manipulate the size of UV absorbance to various degrees in a few genotypes similar to the study performed by Klomberg et al. (2019). Using unscented, UV-absorbing chemicals would keep other flower cues consistent, such as flower shape and size, while only altering UV reflectance area. If increased UV absorption area correlated with increased bee visitation, larger UV area could have led to greater fitness in early cultivated peanut fields by promoting outcrossing and lineage recombination between landraces. In contrast, if decreased UV absorption area correlates with decreased bee visitation, it is possible that selection for other desirable traits may have inadvertently resulted in landraces less attractive to bees, similar to the suggestion of Blanche et al. (2006) that modern peanut varieties may have inadvertently been selected to be less attractive to flower tripping bees. However, further research is needed to elucidate the relationship between UV absorption area and bee attractiveness in Arachis.
For large flowers (> 8 mm), UV is not the only wavelength bees utilize to identify flowers. Blue and green color contrast with foliage in Arachis would increase flower conspicuousness as well. All genotypes tested for banner pigment had significant levels of absorption at wavelengths of 380, 415, 450, 473 and 495 nm, but not for wavelengths in the green interval. Levinson et al. (2021) reported a similar result for banner pigmentation of peanut breeding line 13-2113 and four allotetraploids, IpaCor2, IpaDur3, IpaSten10309, and ValSten1. UV absorption detection at 380 nm was consistent with the finding of UV absorption in the banners in the UV photographs. Flowers from peanut breeding lines, allotetraploids, and wild Arachis species absorb blue wavelengths (450, 473 and 495 nm) and reflect green wavelengths, which would contrast with Arachis foliage that absorbs primarily green wavelengths. In a future study, it would be interesting to test two additional camera filters, one that only allows blue and one that only allows green wavelengths to pass through, and to stack UV, blue, and green wavelength photographs of Arachis flowers still attached to the plant to better understand the contrast of all wavelengths between Arachis flowers and foliage.
For flower detection by bees, it is critical to consider flower visibility beyond just flower color. First, flower size affects flower detection time, in which flowers under 8 mm in diameter take wild bees a longer time to find (Spaethe et al., 2001). All germplasm in this study had flowers with diameters greater than 8 mm and their banner areas were generally similar. Therefore, banner size differences may not cause differences in bee detection of flowers of different Arachis species. Another aspect of flower visibility is the growth habit of the plant coupled with flower morphology. Wild Arachis species and allotetraploids derived from wild Arachis species have prostrate growth, in which branches grow flat along the ground (Stalker et al., 2016). This growth habit coupled with the long hypanthium area characteristic of wild Arachis species and allotetraploids allow flowers to rise above the foliage of the plant and is likely easier to detect by bees (Stalker et al., 2016; Levinson et al., 2021). In contrast, domesticated peanut has been selected for more compact growth, with shorter branches that curve upwards and flowers have a shorter hypanthium than wild Arachis species (Stalker et al., 2016; Levinson et al., 2021). Thus, flowers are more likely to be at least partially hidden from view by plant foliage.
Flower production intensity and duration of different genotypes must also be considered when determining their attractiveness to bees. Positive alleles for flowering precocity have been identified in some allotetraploids (Foncéka et al., 2012), and allotetraploids with significantly higher flower counts over the growing season as compared to cultivated peanut have been identified (Levinson et al., 2021). Furthermore, Levinson et al. (2021) found that allotetraploids made with perennial species such as A. correntina and A. stenosperma do not have a decline in flower production. However, cultivated peanut had a decline in flower production from its maximum of about 20 flowers/day from 52 to 66 days after transplanting to only 5 flowers/day at 87 days after transplanting. This is a large difference compared to the flower production of IpaCor allotetraploids that produced over 40 flowers/day from 52 to 108 days after transplanting (Levinson et al., 2021). In general, cultivated peanuts may have fewer flowers that may be harder to detect than some allotetraploid and diploid, wild Arachis species; however, studies including pollinator counts are needed to determine differences in bee visitation between wild Arachis species, allotetraploids, and cultivated peanut. Such studies may indicate some wild Arachis species and allotetraploids as good candidates for floral strips in agricultural fields and for ornamental use in urban landscapes if Arachis species are found to have abundant protein-, lipid-, and micronutrient-rich pollen.
Another finding of the study was a significant effect of the two different environments (field environment from 18 August to 18 September 2020 and greenhouse environment from 9 November 2020 to 26 February 2021) on the area of UV absorption on the banner, area of UV absorption on the banner as a percentage of banner area, area of UV absorption on the left-wing (adaxial), and area of UV absorption on the left-wing as a percentage of left-wing area (adaxial). However, no environmental effect was observed on banner area, left-wing area, area of UV absorption on the left-wing (abaxial), nor area of UV absorption on the left-wing as a percent of left-wing area. Therefore, while flower size remained similar between genotypes in the two different environments, variation of UV absorption on the banner and on the adaxial side of the wings can be displayed. This same trend of larger UV absorbing areas on flower petals grown in the field as compared to those grown in the greenhouse was also observed in Brassica rapa, in which Sasaki and Takahashi (2002) hypothesized this difference was due to enhancement of flavonoid biosynthesis caused by greater UV-B exposure in the field as compared to the greenhouse. Due to differences in both collection timing and environment in this study, the cause of UV absorbance variation in Arachis in these different environments could be due to UV-B exposure or other factors such as light quality and quantity, temperature, plant developmental stage, among others, that have been documented to affect flavonoid production in various plant tissues (Ghasemzadeh et al., 2010; Hemm et al., 2004). A study keeping all environmental factors constant while only varying UV-B exposure is needed to determine if increased UV-B light exposure increases production of UV-absorbing flavonoids in flowers.
Conclusions
Honeybees and wild bee species provide essential pollination services to improve yields for many fruit, nut, vegetable, and seed crops, yet poor nutrition from habitat loss, large monocultures, and insecticide exposure can result in honeybee colony declines. Pollen abundance and diversity are important to the growth and development of all bees by providing protein and essential amino acids while nectar provides carbohydrates, vitamins, minerals, and amino acids. Bridging the knowledge gap on crop nectar and pollen nutrition and on what contributes to flower attractiveness of various crops, including attractiveness caused by area and pattern of UV absorption, can lead to better decision-making regarding honeybee health and wild pollinator conservation near and within crop fields. Peanut has been documented to be attractive to honeybee and wild bee pollinators but little more of its relationship to bee pollinators is known. This study is one step towards narrowing this knowledge gap, and it found that UV floral guides of various sizes were present in wild, diploid Arachis species, synthetic allotetraploids, and A. hypogaea. The implications of the presence and size of these floral guides on bee visitation are dependent on the pollen quality and quantity in peanut and Arachis species. If future studies find Arachis species to be a source of high-quality pollen, then high-density flowering Arachis species and allotetraploids could be evaluated for promoting wild bee abundance and diversity in urban landscapes, especially those species already used as water-efficient lawns and as ornamental plants.
Acknowledgements
This work was supported by the National Science Foundation (grant # MCB-1543922) and by the AFRI NIFA Fellowships Grant Program: Predoctoral Fellowships project accession no. 1019105 from the USDA National Institute of Food and Agriculture. The authors thank Dr. David Bertioli of the Department of Crop and Soil Sciences, University of Georgia, for manuscript edits.
Literature Cited
Anderson B.D., Knox G.W., Blount A.R., Mackowiak C.L., and Gilman E.F.. 2015. Ornamental groundcover characteristics of rhizoma peanut (Arachis glabrata Benth.): Shade affects height but not cover. HortScience. 50(7):952–956.
Beute M.K., Wynne J.C., and Emery D.A.. 1976. Registration of NC 3033 peanut germplasm.(Reg. No. GP9). Crop Sci. 16:887.
Blaauw B.R., and Isaacs R.. 2014. Flower plantings increase wild bee abundance and the pollination services provided to a pollination-dependent crop. J. Appl. Ecol. 51:890–898.
Blanche K.R., Hughes M., Ludwig J.A., and Cunningham S.A.. 2006. Do flower-tripping beesenhance yields in peanut varieties grown in north Queensland? Aust. J. Exp. Agric. 46:1529–1534.
Branch W.D. 2007. Registration of 'Georgia-06G' peanut. J. Plant Reg. 1:120.
Chu Y., Gill R., Clevenger J., Timper P., Holbrook C.C., and Ozias-Akins P.. 2016. Identification of rare recombinants leads to tightly linked markers for nematode resistance in peanut. Peanut Sci. 43(2):88–93.
Culp T.W., Bailey W.K., and Hammons R.O.. 1968. Natural hybridization of peanuts, Arachis hypogaea L., in Virginia. Crop Sci. 8:109–111.
Dafni A., and Kevan P.. 1996. Floral symmetry and nectar guides: Ontogenetic constraints from floral development, colour pattern rules and functional significance. Bot. J. Linn. 120:371–377.
FAO (Food and Agriculture Organization of the United Nations) 2019. Website http://faostat.fao.org/[accessed08 March 2021].
Foncéka D., Tossim H.-A., Rivallan R., Vignes H., Faye I., Ndoye O., Moretzsohn M.C., et al. 2012. Fostered and left behind alleles in peanut: Interspecific QTL mapping reveals footprints of domestication and useful natural variation for breeding. BMC Plant Biol. 12:26.
Garibaldi L.A., Schulte L.A., Jodar D.N.N., Carella D.S.G., and Kremen C.. 2021. Time to integrate pollinator science into soybean production. Trends Ecol. Evol. 36:573–575.
Garibaldi L.A., Steffan-Dewenter I., Winfree R., Aizen M.A., Bommarco R., Cunningham S.A., Kremen C., et al. 2013. Wild pollinators enhance fruit set of crops regardless of honey bee abundance. J. Sci. 339:1608–1611.
Ghasemzadeh A., Jaafar H.Z.E., and Rahmat A. 2010. Antioxidant activities, total phenolics and flavonoids content in two varieties of Malaysia young ginger (Zingiber officinale Roscoe). Molecules. 15:4324–4333.
Girardeau J.H., and Leuck D.B.. 1967. Effect of mechanical and bee tripping on yield of the peanut. J. Econ. Entomol. 60:1454–1455.
Gorbet D.W., and Tillman B.L.. 2009. Registration of ‘Florida-07’ peanut. J. Plant Regist. 3:14–18.
Gronquist M., Bezzerides A., Attygalle A., Meinwald J., Eisner M., and Eisner T.. 2001Attractive and defensive functions of the ultraviolet pigments of a flower (Hypericum calycinum). PNAS. 98:13745–13750.
Guldberg L.D., and Atsatt P.R.. 1975. Frequency of reflection and absorption of ultraviolet light in flowering plants. Am. Midl. Nat. 93:35–43.
Hammons R.O., and Leuck D.B.. 1966. Natural cross-pollination of the peanut, Arachis hypogaea L., in the presence of bees and thrips. Peanut Sci. 58(4):396.
Harborne J.B., and Smith D.M.. 1978. Anthochlors and other flavonoids as honey guides in the Compositae. Biochem. Syst. Ecol. 6:287–291.
Hemm M.R., Rider S.D., Ogas J., Murry D.J., and Chapple C.. 2010. Light induces phenylpropanoid metabolism in Arabidopsis roots. Plant J. 38(5):765–778.
Holbrook C.C., and Culbreath A.K.. 2007. Registration of ‘Tifrunner’ peanut. J. Plant Regist. 1:124.
Holbrook C.C., Timper P., Culbreath A.K., and Kvien C.K.. 2008. Registration of ‘Tifguard’ peanut. J. Plant Regist. 2:92–94.
Horth L., Campbell L., and Bray R.. 2014. Wild bees preferentially visit Rudbeckia flower heads with exaggerated ultraviolet absorbing floral guides. Biol. Open. 3:221–230.
Horovitz A., and Cohen Y.. 1972. Ultraviolet reflectance characteristics in flowers of crucifers. Am. J. Bot. 59:706–713.
Klein A.M., Vaissière B.E., Cane J.H., Steffan-Dewenter I., Cunningham S.A., Kremen C., Kremen C., and Tscharntke T.. 2007. Importance of pollinators in changing landscapes for world crops. Proc. Royal Soc. B. 274:303–313.
Klomberg Y., , Bartoš R.D., Mertens M., Tropek J.E.J., Fokam R., E.B., and S. Janeček. 2019. The role of ultraviolet reflectance and pattern in the pollination system of Hypoxis camerooniana (Hypoxidaceae). AoB Plants. 11(5):plz057.
Knauft D.A., Chiyembekeza A.J., and Gorbet D.W.. 1992. Possible reproductive factors contributing to outcrossing in peanut (Arachis hypogaea L.). Peanut Sci. 19:29–31.
Kochert G., Stalker H.T., Gimenes M., Galgaro L., Lopes C.R., and Moore K.. 1996. RFLP and cytogenetic evidence on the origin and evolution of allotetraploid domesticated peanut, Arachis hypogaea (Leguminosae). Am. J. Bot. 83:1282–1291.
Leuck D.B., and Hammons R.O.. 1969. Occurrence of atypical flowers and some associated bees (Apoidea) in the peanut, Arachis hypogaea L. J. Agron. 61:958–960.
Leal-Bertioli S.C.M., Moretzsohn M.C., Santos S.P., , Guimarães A.C.M., Bertioli P.M., D.J., and A.C.G. Araujo. 2017. Phenotypic effects of allotetraploidization of wild Arachis and their implications for peanut domestication. Am. J. Bot.104(3):379–388.
Levinson C.M., Chu Y., Luo X., Stalker H.T., Gao D., Holbrook C.C., and Ozias-Akins P.. 2021. Morphological and reproductive characterization of nascent allotetraploids cross-compatible to cultivated peanut (Arachis hypogaea L.). Genet. Resour. Crop Evol. 68:2883–2896.
Lunau K., and Wester P.. 2017. Mimicry and deception in pollinationIn Becard, G. (Ed.), Advances in Botanical Research. Academic Press, pp. 259-279.
Moretzsohn M.C., Gouvea E.G., Inglis P.W., Leal-Bertioli S.C.M., Valls J.F.M., and Bertioli D.J.. 2013. A study of the relationships of cultivated peanut (Arachis hypogaea) and its most closely related wild species using intron sequences and microsatellite markers. Ann. Bot. 111(1):113–126.
Moretzsohn M., Leoi L., , Guimarães K., Leal‐Bertioli P., Gimenes S., Martins M., W.S., et al. 2005. A microsatellite‐based, gene‐rich linkage map of the AA genome of Arachis (Fabaceae). Theor. Appl. Genet. 111:1060–1071.
Nicholls C.I., and Altieri M.A.. 2013. Plant biodiversity enhances bees and other insect pollinators in agroecosystems. A review. Agron. Sustain. Dev. 33:257–274.
Orbán L.L., and Plowright C.M.S.. 2013. The effect of flower‐like and non‐flower‐like visual properties on choice of unrewarding patterns by bumblebees. Naturwissenschaften. 100:621–631.
Papiorek S., Junker R.R., Alves ‐dos‐Santos, I., Melo, G.A.R., Amaral‐Neto, L.P., Sazima, M., Wolowski, et al. 2015. Bees, birds and yellow flowers: Pollinator‐dependent convergent evolution of UV patterns. Plant Biol. 18:46–55.
Peter C.I., and Johnson S.D.. 2008. Mimics and magnets: The importance of color and ecological facilitation in floral deception. J. Ecol. 89:1583–1595.
Primack R.B. 1982. Ultraviolet patterns in flowers, or flowers as viewed by insects. Arnoldia. 42:139–146.
Rae J.M., and Vamosi J.C.. 2013. Ultraviolet reflectance mediates pollinator visitation in Mimulus guttatus. Plant Species Biol. 28:177–184.
Rashad S.E., Ewies M.A., and El Rabie H.G.. 1979. Pollinators of peanut (Arachis hypogaea L.) and the effect of honeybees on its yield. In Proceedings of the IVth International Symposium on Pollination. Agricultural Experiment Station Special Miscellaneous Publication 1, pp. 227-230.
Sasaki K., and Takahashi T.. 2002. A flavonoid from Brassica rapa flower as the UV-absorbing nectar guide. Phytochemistry. 61(3):339–343.
Slater A.T., and Calder D.M.. 1988. The pollination biology of Dendrobium speciosum Smith: A case of false advertising? Australian Journal of Botany. 36:145–158.
Spaethe J., Tautz J., and Chittka L.. 2001. Visual constraints in foraging bumblebees: Flower size and color affect search time and flight behavior. PNAS. 98(7):3898–3903.
Stalker H.T. 2017. Utilizing wild species for peanut improvement. Crop Sci. 57:1102–1120.
Stalker H.T., Tallury S.P., Seijo G.R., and Leal-Bertioli S.C.. 2016Biology, speciation, andutilization of peanut species. In Stalker, H.T., and R.F. Wilson (Eds.), Peanuts: Genetics, processing, and utilization. Academic Press and AOCS Press, pp. 27–66.
Thompson W.R., Meinwald J., Aneshansley D., and Eisner T.. 1972. Flavonols: Pigments responsible for ultraviolet absorption in nectar guide of flower. J. Sci. 177:528–530.
United States Department of Agriculture (USDA). 2015. Attractiveness of agricultural crops to pollinating bees for the collection of nectar and/or pollen. USDA.
Vogel S. 1997. Remarkable nectaries: Structure, ecology, organophyletic perspectives I. Substitutive nectaries. Flora. 192:305–333.
Vorobyev M., and Brandt R.. 1997. How do insect pollinators discriminate colors? Isr.J. Plant Sci. 45:103–113.
Notes
-
Author Affiliations
- Institute of Plant Breeding, Genetics and Genomics, University of Georgia, Athens, GA, USA; [^]
- Horticulture Department, University of Georgia, Tifton, GA, USA; [^]
- Plant Pathology Department, University of Georgia, Athens, GA, USA; [^]
- Crop and Soil Sciences, North Carolina State University, Raleigh, NC, USA; [^]
- USDA- Agricultural Research Service, Crop Genetics and Breeding Research Unit, Tifton, GA, USA; [^]
- Center for Applied Genetic Technologies, University of Georgia, Athens, GA, USA. [^] Corresponding author’s E-mail: pozias@uga.edu