INTRODUCTION
Climate change and limited water are significant challenges to the future of peanut production. In the southwestern U.S. peanut production area, an estimated 35% of the southern High Plains will not be able to support irrigated agriculture within 30 years at current depletion rates of the Ogallala Aquifer (Scanlon et al., 2012). In addition, global warming is expected to add additional demands on the aquifer (Rosenberg et al., 1999). Recurring and severe droughts have also occurred in the southeastern U.S. (Branch and Kvien, 1992; Strzepek et al., 2010), which produced 65% of U.S. peanuts from 1980-2019 (USDA-ERS, 2021). The importance of drought-tolerant peanuts also extends beyond producing profitable yields under limited water. Higher levels of aflatoxin contamination are observed under drought conditions (Sanders et al., 1985; Holbrook et al., 2000, 2009; Arunyanark et al., 2009, 2010; Girdthai et al., 2010). Despite this, few drought-tolerant peanut cultivars are available.
In peanut, up to 153 marker-trait associations and quantitative trait loci associated with drought-tolerance have been identified, but many of them have minor effects (Ravi et al., 2011; Gautami et al., 2012; Pandey et al., 2014, 2021). Multiple physiological processes mediate drought tolerance, making breeding and phenotyping for this trait especially difficult (Cattivelli et al., 2008; Ravi et al., 2011). Yield and seed quality traits, such as shelling percentage and 100-seed weight, are frequently used to assess drought resistance (Nageswara Rao et al., 1989; Branch and Kvien, 1992; Faye et al., 2015; Pandey et al., 2021). However, yield data are difficult to procure for large mapping populations (Nigam et al., 2005) and may be less reliable as an indicator for multiple locations due to significant genotype x environment interactions (Arunyanark et al., 2008). As a result, surrogate physiological traits associated with drought tolerance are now used to screen larger numbers of genotypes (Pandey et al., 2014, 2021).
Soil plant analysis development measurement (SPAD) or SPAD chlorophyll meter readings (SCMR) are one of the most widely measured physiological traits, perhaps due to the ease of data collection (Upadhyaya, 2005; Rowland and Lamb, 2005; Lal et al., 2006; Arunyanark et al., 2008; Songsri et al., 2008; Balota et al., 2012; Singh et al., 2014; Luis et al., 2016). Several studies with peanut have demonstrated SCMR as a relatively stable trait that correlates well with transpiration efficiency, a trait generally regarded as a reliable indicator of drought tolerance, but one that is difficult to measure on a large scale (Nageswara Rao et al., 2001; Upadhyaya, 2005; Sheshshayee et al., 2006; Arunyanark et al., 2008). Canopy temperature, canopy temperature depression (CTD), and normalized difference vegetation index (NDVI) have also been used for phenotyping drought tolerance (Rucker et al., 1995; Holbrook et al., 2000; Jongrungklang et al., 2008; Nautiyal et al., 2008; Balota et al., 2012; Singh et al., 2014; Luis et al., 2016). Visual indicators of drought stress, such as wilting and leaf color were significantly correlated with aflatoxin contamination (Holbrook et al., 2000; Luis et al., 2016) and are also promising as surrogate traits. Lastly, another visible trait, leaf folding or paraheliotropism, is an adaptation present in some peanut genotypes to reduce solar radiation under drought stress (Matthews et al., 1988; Chapman et al., 1993).
To be useful for peanut breeding, genotypes at the phenotypic extremes of these surrogate traits need to be identified (Shekoofa et al., 2013). Preliminary work identified candidate accessions with divergent phenotypes from the U.S. mini-core. The goal of this project was to evaluate the accessions in three peanut production environments, spanning subhumid southeastern Virginia to semiarid southwestern Oklahoma and the southern Texas High Plains.
Materials and Methods
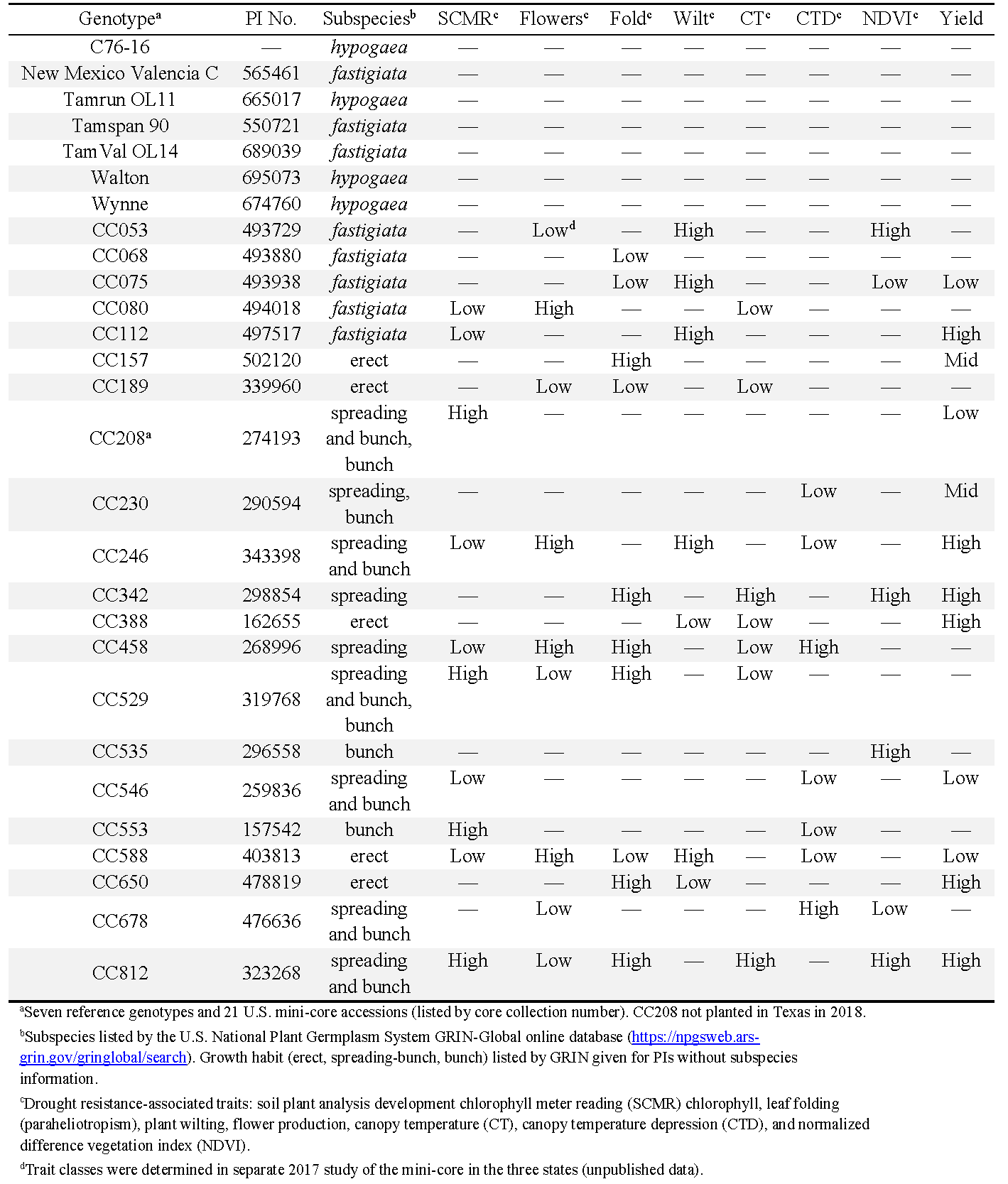
A total of 21 core collection (CC) accessions from the U.S. mini-core collection, in addition to seven reference genotypes, were evaluated in 2018 and 2019 in Oklahoma, Texas, and Virginia (Table 1). The 21 accessions were selected from a 2017 study of the broader mini-core because they exhibited divergent phenotypes for traits associated with drought tolerance (data unpublished). The seven reference genotypes included the following: runner breeding line C76-16 (Holbrook et al., 2013) and cv. Tamrun OL11 (Baring et al., 2013); Spanish cv. Tamspan 90 (Smith et al., 1991); Valencia cvs. New Mexico Valencia C (Hsi, 1980) and TamVal OL14 (Burow et al., 2019); and Virginia cvs. Walton (Balota et al., 2021; tested as 08X09-3-14-1) and Wynne (Plant Variety Protection 201500288). C76-16 (Holbrook et al., 2013; Luis et al., 2016; Bhogireddy et al., 2020; Wang et al., 2021) and Walton (Balota et al., 2021) have demonstrated drought tolerance, and New Mexico Valencia C (Bhogireddy et al., 2020) was recently shown to have poor drought tolerance.
In Oklahoma, the experiments were conducted at Oklahoma State University’s Caddo Research Station in Fort Cobb. In 2018, entries were planted in a randomized complete block design (RCBD) with four replications/blocks on 18 May. Half of the field was harvested at 134 days after planting (DAP) on 28 September; the remainder was harvested two weeks later on 12 October. In 2019, entries were planted on 17 May using a RCBD, however, three blocks were in a water-restricted field, and two blocks were in a field with normal irrigation. The irrigated field was harvested 141 DAP, but due to logistical and weather delays, the water-restricted field was harvested 172 DAP. In Texas, the experiments were conducted at the USDA-ARS Cropping Systems Research Laboratory in Lubbock. Entries were planted using a RCBD with four blocks in 2018 and 2019 under water-deficit conditions. Planting and harvest dates were 17 May and 8 November (176 DAP) in 2018 and 14 May and 17/18 October (157-158 DAP) in 2019, respectively. In Virginia, the experiment was conducted at Virginia Tech’s Tidewater Agricultural Research and Extension Center (TAREC) in Tidewater. Three blocks arranged in a RCBD were each planted in an irrigated field and a water-restricted field. Rain-out shelters covered the water-restricted fields from 16 July 2018 and 15 July 2019 for 6 weeks and were removed on 30 August 2018 and 27 August 2019. In 2018, the irrigated plots were planted on 17 May and harvested on 9 September (116 DAP), and the water-restricted plots were harvested at 125 DAP. Planting and harvest dates in 2019 were 30 April and 10 September (134 DAP). Plots in all states were managed for weeds and diseases following their respective University Extension recommendations. In all states, water-restricted plots were irrigated to 25% evapotranspiration (ET) replacement from approximately 40 to 105 DAP (weather permitting in Oklahoma and Texas). This growth period includes the pod-filling stage (Boote, 1982), a time that is especially susceptible to water stress (Nageswara Rao et al., 1985, 1989).
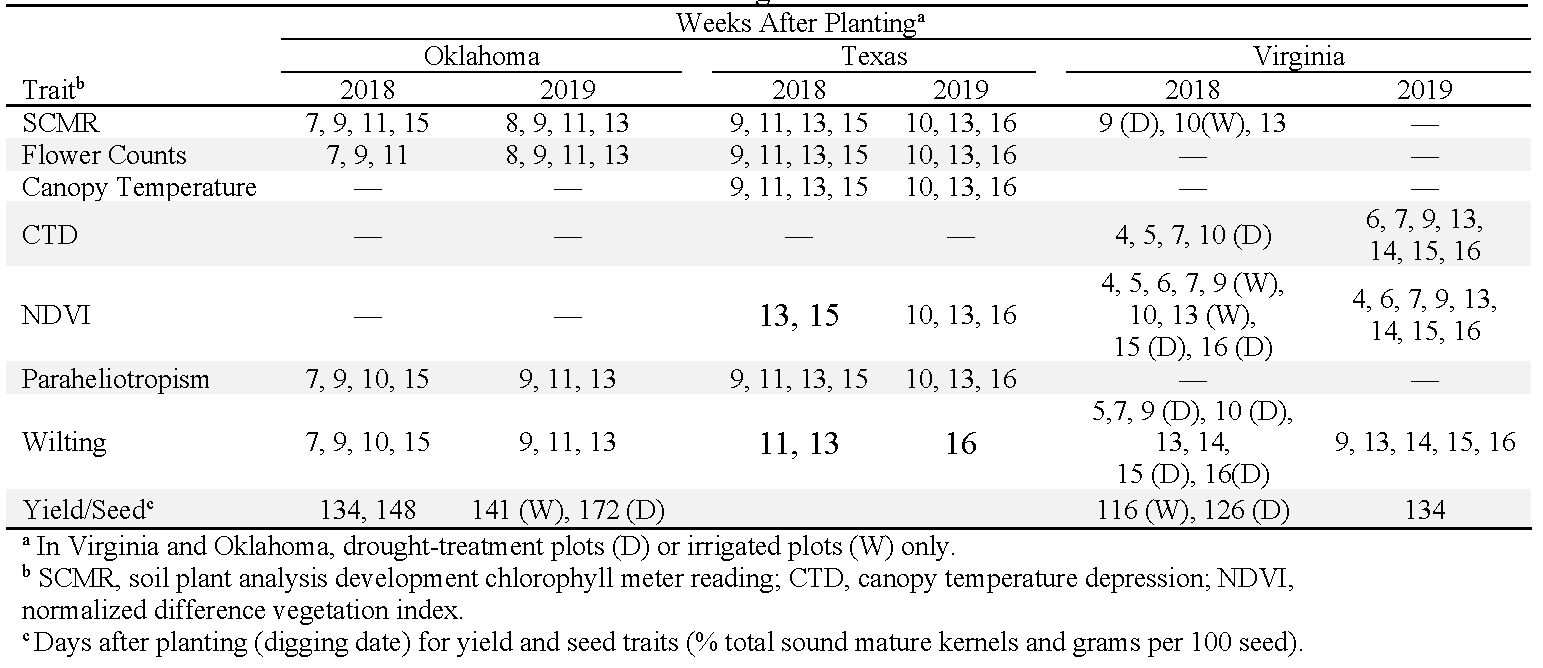
Ten traits associated with drought resistance were measured, but the specific traits measured varied among states depending on equipment availability and labor (Table 2). SCMR readings were taken by recording the mean from four representative plants in each plot. The 4th fully expanded leaf from the top of the mainstem of each of four plants was measured in Oklahoma and Virginia; the 4th fully expanded leaf from lateral branches were measured in Texas. SCMR readings were taken with a SPAD 502 Plus (Spectrum Technologies, Aurora, IL) in Oklahoma, and a SPAD-502 (Minolta, Tokyo, Japan) in Texas and Virginia. In Oklahoma and Texas, fresh, non-wilted flowers were counted in the morning within a representative 1.86-m section inside each plot (Oklahoma) or within the entire plot (Texas). An Agri-Therm II (model 100L, Everest Interscience, Chino Hills, CA) was used to measure afternoon (13:00 to 16:00) canopy temperature and canopy temperature depression (CTD). Normalized difference vegetation index (NDVI) was measured in Texas and Virginia with the GreenSeeker Handheld Crop Sensor (Trimble Agriculture, Sunnyvale, CA). In Oklahoma and Texas, paraheliotropism was rated in the afternoon using the following scale: 0, no folding; 1, 10% leaves folded; 2, 30% leaves folded; 3, 50% leaves folded; 4, 70% folded; 5, 90% folded; and 6, 90% folded but with leaves closed tightly and puckered around leaf margins. Plant wilting was also rated in the afternoon with the following scale in Oklahoma and Texas: 1, no wilting; 2, 25% of plants wilted; 3, 50% wilted; 4, 75% wilted; and 5, 100% leaves wilted and ca. 50% leaves brown for >90% plants within plot. In Virginia, a wilting scale from 0-5 was used (Sarkar et al., 2021). After harvest, pods were dried in a forced-air dryer to approximately 7% moisture by weight and separated from soil and plant debris with a peanut cleaner. One 200-g sample of pods from each plot was used to determine the percentage of total sound mature kernels (TSMK) following USDA-AMS (2019) guidelines. To simplify grade collection from diverse genotypes, TSMK for all entries was determined using the smallest screen for Spanish/Valencia kernels (5.94-mm x 19.05-mm (15/64 x 3/4 inch) slots) in Oklahoma and Virginia.
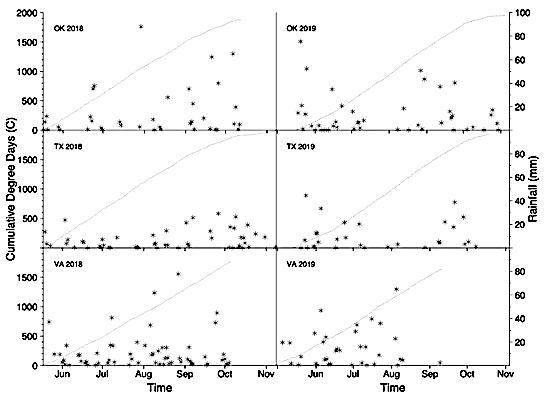
Temperature and rainfall were monitored using the Fort Cobb Oklahoma Mesonet station at the Caddo Research Station (McPherson et al., 2007), the Plant Stress & Water Conservation Meteorological Tower at ARS-Lubbock ( https://www.ars.usda.gov/plains-area/lubbock-tx/cropping-systems-research-laboratory/wind-erosion-and-water-conservation-research/docs/pswc-met-tower-data/), and the WeatherStem system at TAREC ( http://vt-arec.weatherstem.com/arec115). Degree day heat units were calculated using the West Texas method 1 formula for peanut (Rowland et al., 2006)(Figure 1):
Degree Day = (ATmax35 + ATmin12.8)/2 – 12.8
where ATmax35 and ATmin12.8 are the maximum and minimum air temperatures capped at 35C and 12.8C, respectively.
Data were analyzed with SAS version 9.4 (SAS Institute, Cary, NC). To account for environmental differences between years and among states (environments), cumulative degree day was tested as a covariate in all analyses. Differences among genotypes and environments for drought-associated traits were estimated using mixed-model ANOVA with PROC MIXED. Fixed effects included state, genotype, year, and block; year*block was used as the random effect. The SLICE option was used to examine the effects of state and genotype. For flower count data, the sum of flowers counted each season per plot was used for the analyses to minimize the effect of genetic differences in flowering times among genotypes. Similarly, differences among genotypes and water treatments (irrigated and water-deficit) in Oklahoma and Virginia were also examined with PROC MIXED. Pearson’s correlations among traits collected from the water-restricted plots in all states and years were calculated using the raw data with PROC CORR.
Results and Discussion
Since rainout shelters were not available in Oklahoma and Texas, water-restricted plots were exposed to rain. However, rainfall events greater than 20 mm were relatively rare between 40 and 105 DAP in both years, when deficit irrigation was imposed (Figure 1). The following range of cumulative degree days between 40 and 105 DAP were observed: 561-1493 and 429-1374 in Oklahoma, 582-1529 and 420-1384 in Texas, and 426-1269 and 428-1277 in Virginia in 2018 and 2019, respectively (Figure 1). In 2018 in Virginia, irrigated plots were harvested early (116 DAP) due to severe disease outbreaks of late leaf spot, southern blight, and Sclerotinia blight (caused by Nothopassalora personata, Athelia rolfsii, and Sclerotinia minor, respectively). In 2019, plants in the water-restricted plots in Oklahoma were killed by an early freeze (22 October) 13 days before digging.
Comparisons Among Water-restricted Plots in Oklahoma, Texas, and Virginia
For all analyses comparing drought-associated traits within the water-restricted plots, the effect of cumulative degree day was highly significant (P < 0.01), validating its use as a covariate. In the analyses of SCMR readings (Table 3), states and genotypes differed in SCMR (P < 0.01), as well as the genotype*environment interaction (F = 1.77, P < 0.01), which indicated that mean SCMR values for genotype varied with state. For all genotypes, SCMR readings were highest in Texas and lowest in Virginia, perhaps due to differences in SCMR meter models (Huang and Peng, 2004), the time of day the data were collected (Arunyanark et al., 2008), or plant water status (Martínez and Guiamet, 2004). Despite the significant genotype*environment interaction, common genotypes occupied the top five lowest and highest SCMR in all states: CC546, CC080, and New Mexico Valencia C had low SCMR; Tamrun OL11, CC678, and CC529 had high SCMR. In the wilt analyses, genotype (F = 4.04, P < 0.01) had a significant effect on wilting across states, but the effects of state (P = 0.7) and genotype*environment interaction (P = 0.1) were not significant. While genotype had a significant effect across states, no significant differences in wilting among genotypes were observed in Texas, contributing to the near-statistically significant genotype*environment interaction. CC053 had the highest wilt scores in all states, and CC535 and Walton were among the least wilted genotypes in Oklahoma and Virginia.
Paraheliotropism and flower count data were collected from Oklahoma and Texas (Table 4). State (F = 136.13, P < 0.01) and genotype (F = 23.68, P < 0.01) had significant effects on paraheliotropism scores, but the interaction between the two effects was not significant (P = 0.65). Paraheliotropism scores were generally higher in Texas than in Oklahoma. Leaf folding in alfalfa was shown to vary depending on ambient vapor pressure deficit (Reed and Travis, 1987), and the lower humidity in West Texas despite similar daily maximum temperatures to Oklahoma may have resulted in more paraheliotropism. In both states, the least paraheliotropism was observed in CC189, and genotypes exhibiting the most leaf folding included CC678, CC342, and Walton. For flower counts, state (F = 312.7), genotype (F = 10.3), and the genotype*environment interaction (F = 2.45) had highly significant effects (P < 0.01) on the total number of flowers counted. More flowers were counted in Oklahoma than in Texas for 21 of the 28 genotypes (Table 4). In Texas, large numbers of flowers were observed following rainfall or irrigation in 2019 (data not shown), but few were observed on actual data collection dates in Texas. Nonetheless, both states shared genotypes with the highest (CC80) and fewest (CC189) flower counts when grown under drought stress.
NDVI and canopy temperature data were collected in Texas and Virginia (Table 5). State (F = 20.0, P < 0.01) had a significant effect on NDVI, but no differences among genotypes or a genotype*environment interaction were observed (P ≥ 0.93). In addition, no differences among genotypes were observed for canopy temperatures collected in Texas (F = 0.81, P = 0.74) or CTD collected in Virginia (F = 1.38, P = 0.10). Canopy temperature meters focused on relatively narrow sections of plots, and considerable variability was observed within plots in Texas (data not shown). Thus, imaging systems capable of obtaining mean temperatures from entire plots might be more useful.
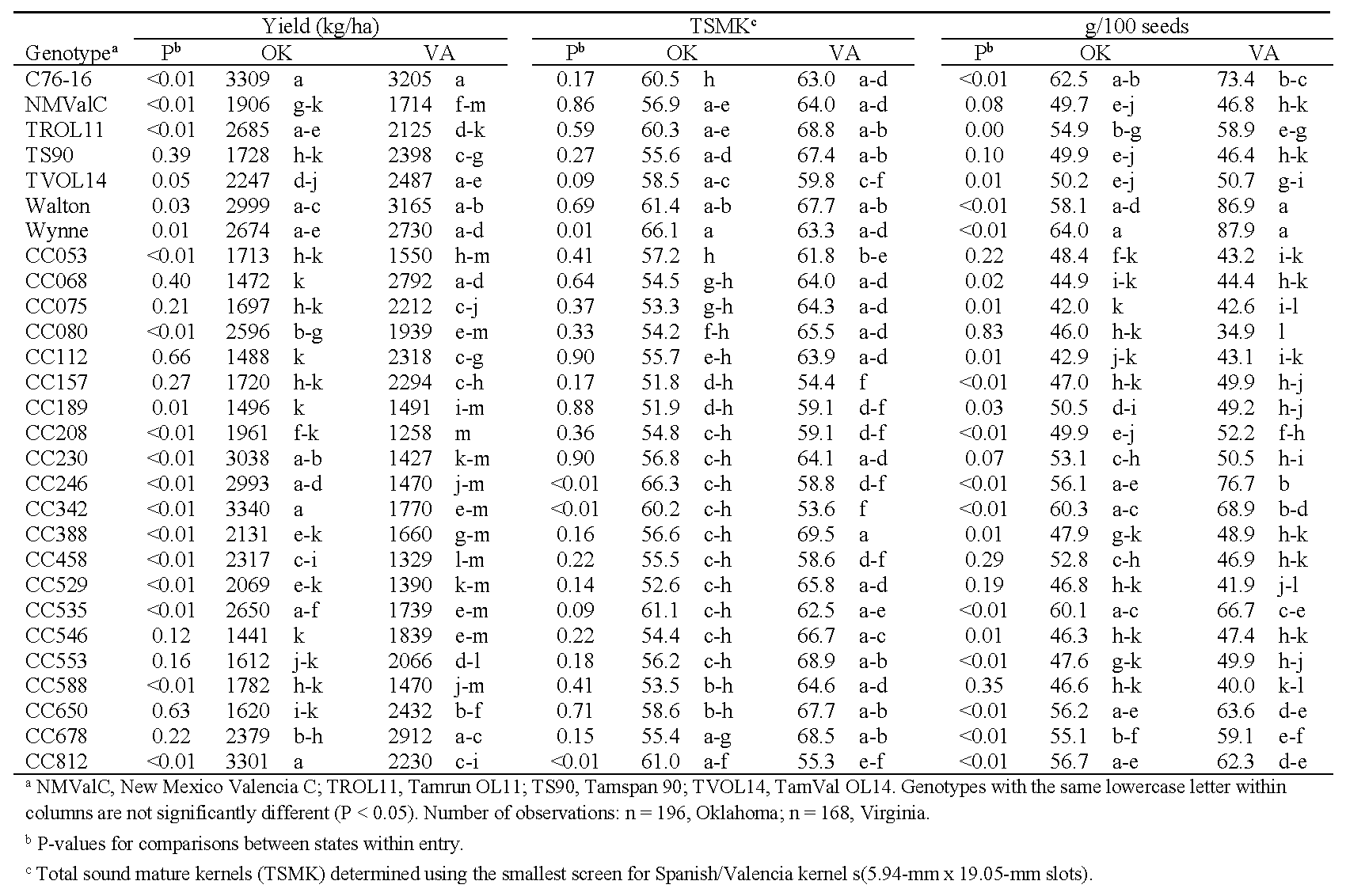
Yield, grade, and seed weights were collected from plots in Oklahoma and Virginia (Table 6). Under water-limited conditions, the effects of state and genotype on yield and seed weights were highly significant (P < 0.01). For TSMK, the effect of state was not significant (P = 0.18) though genotype was significant (P < 0.01). However, the genotype*environment interactions were also significant for all traits (P < 0.01). Maximum yields under drought conditions were similar in both Oklahoma and Virginia, ca. 3200-3300 kg/ha. Yields from C76-16 and Walton were among the top five numerically, but the lowest yields were produced by different genotypes in each state (Table 6). The states also had few genotypes in common for the highest and lowest total sound mature kernels. Seed weight was more consistent between the two locations: Wynne, Walton, C76-16, and CC342 were among the heaviest seed, and CC075, CC080, and CC112 were among the lightest.
Correlation analyses revealed few strong associations among drought-associated traits (Table 7). While genotypes did not differ in canopy temperature, the strongest correlation observed was between canopy temperature and total flower counts in Texas (r = -0.8). NDVI (r = 0.6) and paraheliotropism (r = -0.4) were also moderately correlated with flower counts. Paraheliotropism and SCMR were also moderately associated (r = 0.5). Several traits were significantly, albeit moderately, correlated with yield and seed-quality traits, the strongest of which was between yield and TSMK with SCMR (r = 0.48).
Comparisons Among Water-restricted and Irrigated Plots in Oklahoma and Virginia
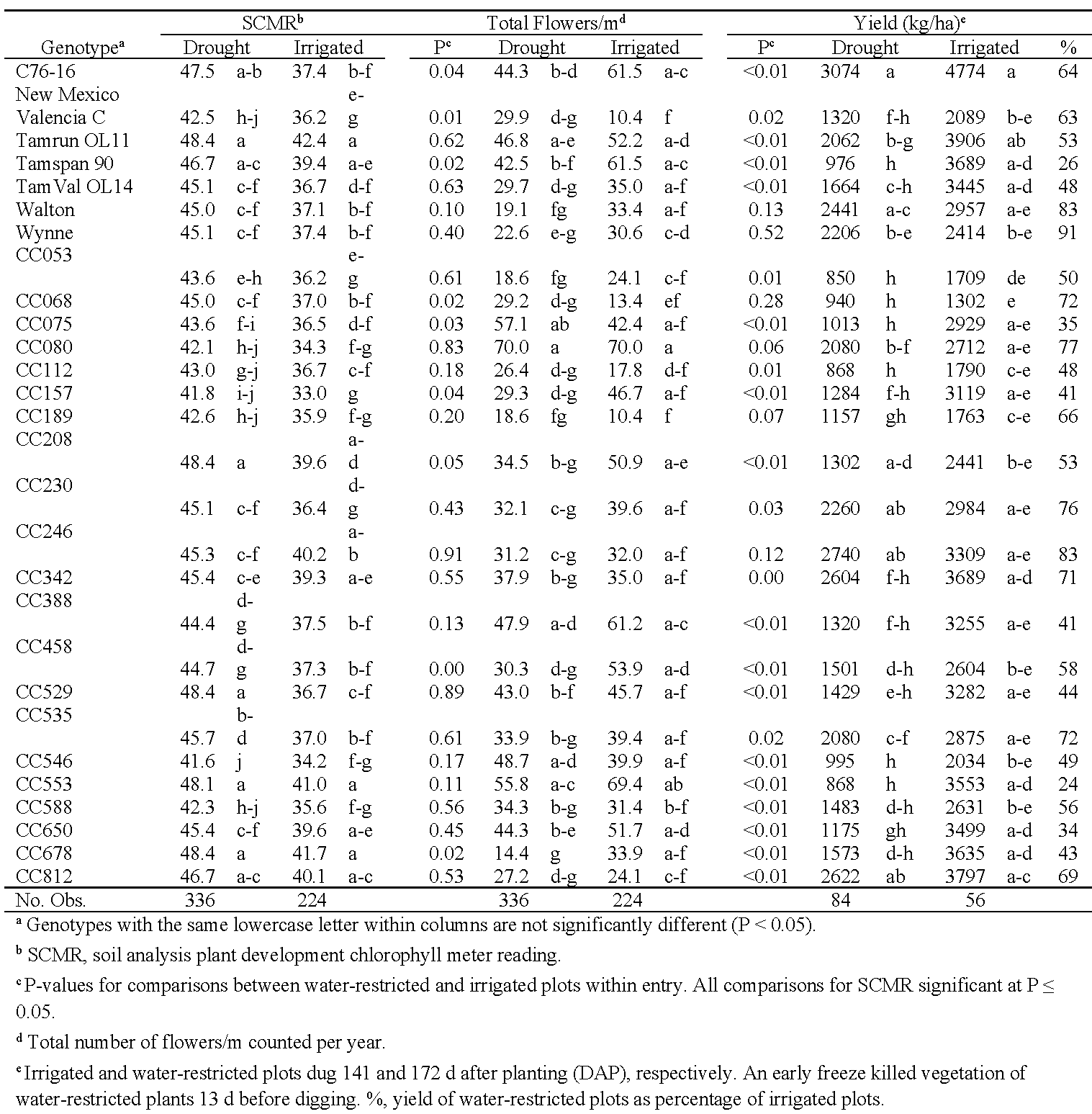
To assess genotype responses to limited water in comparison to normal irrigation, analyses were conducted between water-restricted and irrigated plots in 2019 in Oklahoma and in both years in Virginia (Tables 8 and 9). In Oklahoma, no paraheliotropism and wilting data were collected from the irrigated plots because little leaf folding or wilting were observed under irrigation. The effect of cumulative degree day was significant only in the SCMR analysis (P < 0.01), so analysis of covariance was used only for this trait in Oklahoma. For SCMR, irrigation (F = 253.3) and genotype (F = 9.3) had significant effects (P < 0.01), and the irrigation*genotype interaction was not significant (P = 0.48). All genotypes had significantly higher SCMR values when grown under drought conditions than with normal irrigation. For flower counts, the water treatments did not have a significant effect (F = 2.1, P = 0.19), but both genotype and the genotype*water treatment interaction were significant (P < 0.01; Table 7). Seven genotypes (C76-16, CC157, CC208, CC458, CC553, CC650) produced significantly more flowers under irrigation than when drought stressed. Conversely, New Mexico Valencia C, CC068, and CC112 produced more flowers under drought stress, and CC080 produced an equally high number of flowers under both water treatments. Meisner and Karnok (1992), taking daily flower counts, observed that flowering in Florunner could recover after being suppressed by drought. In our experiment, flower counts were taken on 3 d in 2018 and 4 d in 2019, so it is unclear if the three genotypes consistently produced many flowers over the duration of their flowering period. Unfortunately, the Oklahoma yield, TSMK, and seed weight data are confounded by a 31-d difference between harvest dates for the irrigated (141 DAP) and water-restricted plots (172 DAP), due in part to inclement weather. The early hard freeze in combination with the late harvest of the water-restricted plots likely had detrimental effects on yield and seed quality, particularly in early-maturing genotypes. Yield in Oklahoma was influenced by genotype (F = 11.4, P < 0.01) and genotype*water treatment (F = 3.3, P < 0.01), but the effect of the water treatment bordered on being significant (F = 4.8, P = 0.08). Yields from water-restricted plots ranged from 24% (CC553) to 91% (Wynne) of the irrigated plots (Table 8). For TSMK and seed weight, water treatment and genotype had significant effects (P < 0.05), but their interaction was not significant (P = 0.27 and 0.34, respectively; data not shown).
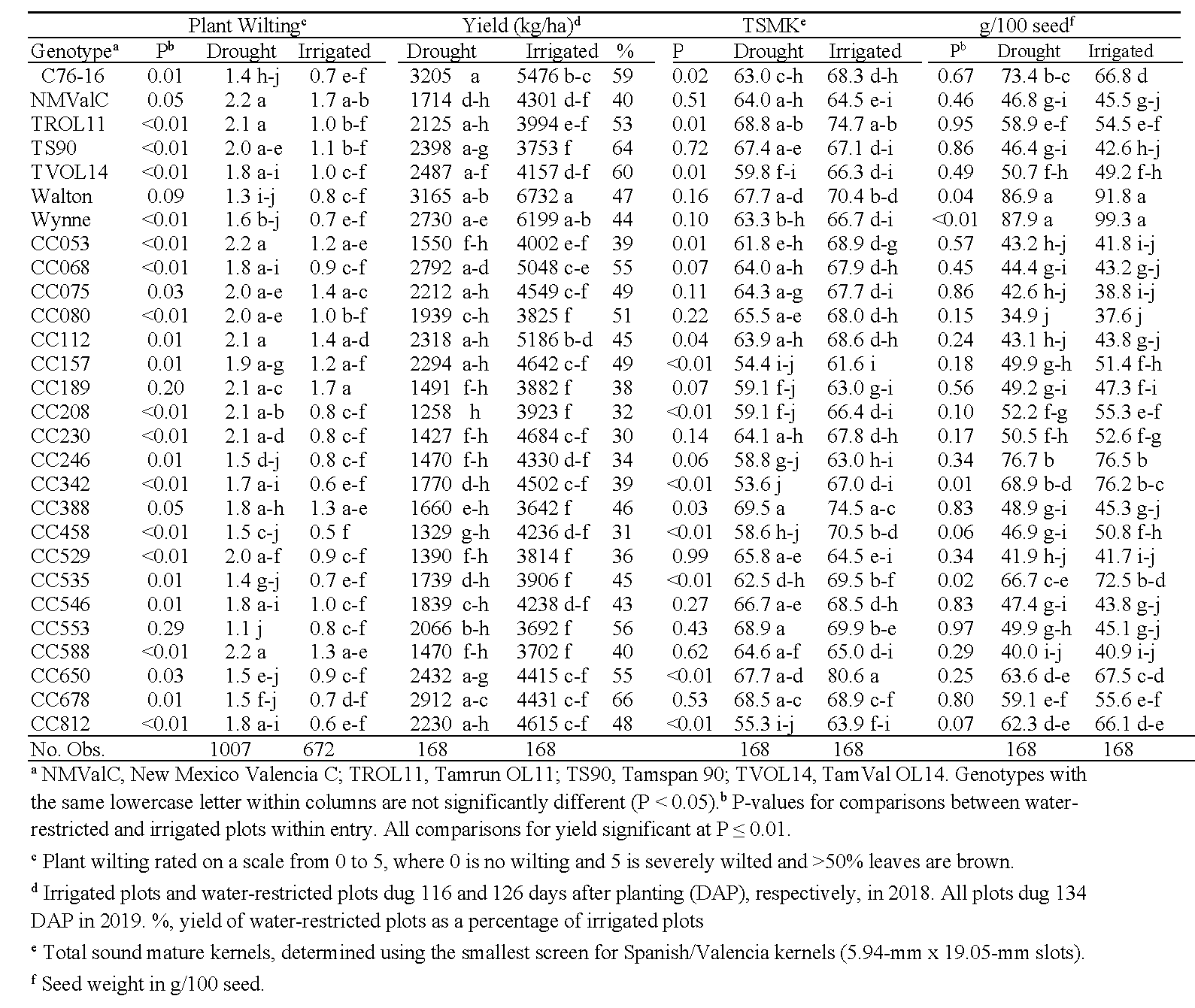
In Virginia, cumulative degree day had significant effects on each trait (P < 0.01) and was therefore used as a covariate for all analyses between irrigated and water-restricted plots. For CTD, the effects of water treatment, genotype, and the interaction between the two were not significant (P>0.75). Water treatment significantly affected NDVI ratings (F = 55.9, P < 0.01), but genotype and the genotype*water treatment interaction were also not significant (P > 0.32). Overall, NDVI readings taken from irrigated plots were numerically greater than those from drought-stressed plots (data not shown), showing that NDVI is affected by water stress on peanuts (Rouse et al., 1974; Vergara-Diaz et al., 2015). Analyses of plant wilting showed significant effects of water treatment and genotype (P < 0.01), and a nonsignificant interaction (P = 0.84). As expected, plant wilting was greater in the water-restricted than irrigated treatments (Table 9). Wilt scores for CC553, CC189, and Walton did not differ statistically between the two water treatments. For yield and seed weights, both water treatment and genotype were significant (P < 0.01), and the interaction between the two were nonsignificant (P > 0.56). Yields from the water-restricted plots ranged from 30% (CC230) to 66% (CC678) of the irrigated treatments (Table 9). Numerically, the five highest-yielding genotypes under drought conditions in Virginia were C76-16, Walton, CC678, CC068, and Wynne. For most genotypes, 100-seed weights between the two water treatments did not differ, except for Wynne, CC342, CC535, and Walton, where seed weights were significantly larger from the irrigated plots. For TSMK, all effects of water treatment, genotype, and their interaction were significant (P < 0.04). When TSMK significantly differed between water treatments within genotypes, shell-out in irrigated plots was greater than under water restriction. Genotypes with similar TSMK (P ≥ 0.43) regardless of water treatment included CC529, Tamspan 90, CC588, CC678, New Mexico Valencia C, and CC553.
Outside the U.S., germplasm has been screened for multiple drought tolerance-associated traits in geographically and climatically diverse environments (Faye et al., 2015; Pandey et al., 2021). However, to date, just one study in the U.S. has evaluated drought tolerance in considerably different production environments—Georgia and Arizona (Holbrook et al., 2000). Here, we identified mini-core accessions with divergent phenotypes for drought-associated traits that were consistent across three distinct environments. A significant genotype by environment interaction was observed for SCMR, but common genotypes occupied the extremes in all states. Similar to observations by Holbrook et al. (2009), a significant correlation was not observed between SCMR and visual drought stress/wilting, but moderate, significant correlations were observed with paraheliotropism, yield, and TSMK. Others have also observed correlations between SCMR and pod yield (Upadhyaya, 2005; Songsri et al., 2008; Singh et al., 2014). Genotypes were significantly different for plant wilting and paraheliotropism, and these traits appeared to be less affected by genotype by environment interactions. Thus, wilting and paraheliotropism may be particularly useful for breeding as surrogate traits for drought tolerance as previously suggested (Holbrook et al., 2000; Luis et al., 2016). While these visually-assessed traits are non-destructive and less time-consuming to collect than other physiological traits such as leaf water potential (O’Toole et al., 1984), rating for wilt and paraheliotropism is still labor intensive. Remote sensing technologies may offer high-throughput solutions for phenotyping these traits (Sullivan and Holbrook, 2007; Sarkar et al., 2021).
Acknowledgements
The authors thank the following people for technical assistance over multiple years: A. Harting and L. Myers, USDA-ARS-Stillwater, OK; R. Weidenmaier, H. Houston, and B. Leighton, OAES Caddo Research Station, Fort Cobb, OK; F. Bryant and D. Redd, Tidewater AREC, Suffolk, VA; undergraduate students K. Carter, N. Cash, I. Hover, G. Lane, and C. Skinner. This project was supported by: AFRI award #2017-67013-26193 from the USDA-NIFA; CRIS Projects 3072-21220-008-00D and 3096-21000-022-000-D; and USDA Hatch Project 880350. Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture. USDA is an equal opportunity provider and employer.
Literature Cited
Arunyanark A., Jogloy S., Akkasaeng C., Vorasoot N., Kesmala T., and Patanothai A.. 2008. Chlorophyll stability is an indicator of drought tolerance in peanut. J. Agron. Crop Sci. 194:113–125.
Arunyanark A., Jogloy S., Wongkaew S., Akkasaeng C., Vorasoot N., Kesmala T., and Patanothai A.. 2009. Association between aflatoxin contamination and drought tolerance traits in peanut. Field Crops Res. 114:14–22.
Arunyanark A., Jogloy S., Wongkaew S., Akkasaeng C., Vorasoot N., Wright G.C., Rachaputi R. C., and Patanothai A.. 2010. Heritability of aflatoxin resistance traits and correlation with drought tolerance traits in peanut. Field Crops Res. 117:258–264.
Balota M., Isleib T.G., and Tallury S.. 2012. Variability for drought related traits of virginia-type peanut cultivars and advanced breeding lines. Crop Science 52:2702–2713.
Balota M., Tillman B.L., Paula-Moraes S.V., and Anco D.. 2021. ‘Walton’, a new virginia-type peanut suitable for Virginia and northern U.S. growing regions. J. Plant Regist. 15:422-434.
Baring M.R., Simpson C.E., Burow M.D., Cason J.M., and Ayers J.L.. 2013. Registration of ‘Tamrun OL11’ peanut. J. Plant Regist. 7:154–158.
Bhogireddy S., Xavier A., Garg V., Layland N., Arias R., Payton P., Nayak S.N., Pandey M.K., Puppala N., and Varshney R.K.. 2020. Genome-wide transcriptome and physiological analyses provide new insights into peanut drought response mechanisms. Sci. Rep. 10:4071.
Boote K.J. 1982. Growth stages of peanut (Arachis hypogaea L.). Peanut Sci. 9:35–40.
Branch W.D., and Kvien C.K.. 1992. Peanut breeding for drought resistance. Peanut Sci. 19:44–46.
Burow M.D., Baring M.R., Chagoya J., Trostle C., Puppala N., Simpson C.E., Ayers J.L., Cason J.M., Schubert A.M., Muitia A., and López Y.. 2019. Registration of ‘TAMVal OL14’ Peanut. J. Plant Regist. 13: 134–138.
Cattivelli L., Rizza F., Badeck F.-W., Mazzucotelli E., Mastrangelo A.M., Francia E., Marè C., Tondelli A., and Stanca A.M.. 2008. Drought tolerance improvement in crop plants: An integrated view from breeding to genomics. Field Crops Res. 105:1–14.
Chapman S.C., Ludlow M.M., Blamey F.P.C., and Fischer K.S.. 1993. Effect of drought during early reproductive development on growth of cultivars of groundnut (Arachis hypogaea L.). I. Utilization of radiation and water during drought. Field Crops Res. 32:193–210.
Faye I., Pandey M.K., Hamidou F., Rathore A., Ndoye O., Vadez V., and Varshney R.K.. 2015. Identification of quantitative trait loci for yield and yield related traits in groundnut (Arachis hypogaea L.) under different water regimes in Niger and Senegal. Euphytica 206:631–647.
Gautami B., Pandey M.K., Vadez V., Nigam S.N., Ratnakumar P., Krishnamurthy L., Radhakrishnan T., Gowda M.V.C., Narasu M.L., and Hoisington D.A.. 2012. Quantitative trait locus analysis and construction of consensus genetic map for drought tolerance traits based on three recombinant inbred line populations in cultivated groundnut (Arachis hypogaea L.). Mol. Breed. 30:757–772.
Girdthai T., Jogloy S., Vorasoot N., Akkasaeng C., Wongkaew S., Holbrook C.C., and Patanothai A.. 2010. Associations between physiological traits for drought tolerance and aflatoxin contamination in peanut genotypes under terminal drought. Plant Breed. 129:693–699.
Holbrook C.C., Guo B.Z., Wilson D.M., and Timper P.. 2009. The U.S. breeding program to develop peanut with drought tolerance and reduced aflatoxin contamination. Peanut Sci. 36:50–53.
Holbrook C.C., Isleib T.G., Ozias-Akins P., Chu Y., Knapp S.J., Tillman B., Guo B., Gill R., and Burow M.D.. 2013. Development and phenotyping of recombinant inbred line (RIL) populations for peanut (Arachis hypogaea). Peanut Sci. 40:89–94.
Holbrook C.C., Kvien C.K., Rucker K.S., Wilson D.M., Hook J.E., and Matheron M.E.. 2000. Preharvest aflatoxin contamination in drought-tolerant and drought-intolerant peanut genotypes. Peanut Sci. 27:45–48.
Hsi D.C. 1980. Registration of New Mexico Valencia C peanut (Reg. No. 24). Crop Sci. 20:113–114.
Huang J., and Peng S.. 2004. Comparison and standardization among chlorophyll meters in their readings on rice leaves. Plant Prod. Sci. 7:97–100.
Jongrungklang N., Toomsan B., Vorasoot N., Jogloy S., Kesmala T., and Patanothai A.. 2008. Identification of peanut genotypes with high water use efficiency under drought stress conditions from peanut germplasm of diverse origins. Asian J. Plant Sci. 7:628.
Lal C., Hariprasanna K., Rathnakumar A.L., Gor H.K., and Chikani B.M.. 2006. Gene action for surrogate traits of water-use efficiency and harvest index in peanut (Arachis hypogaea). Ann. Appl. Biol. 148:165–172.
Luis J.M., Ozias-Akins P., Holbrook C.C., Kemerait Jr R.C., Snider J.L., and Liakos V.. 2016. Phenotyping peanut genotypes for drought tolerance. Peanut Sci. 43:36–48.
Martínez D., and Guiamet J.. 2004. Distortion of the SPAD 502 chlorophyll meter readings by changes in irradiance and leaf water status. Agronomie 24:41–46.
Matthews R.B., Harris D., Williams J.H., and Rao R.N.. 1988. The physiological basis for yield differences between four genotypes of groundnut (Arachis hypogaea) in response to drought. II. Solar radiation interception and leaf movement. Exp. Agric. 24:203–213.
McPherson R.A., Fiebrich C.A., Crawford K.C., Kilby J.R., Grimsley D.L., Martinez J.E., Basara J.B., Illston B.G., Morris D.A., Kloesel K.A., Melvin A.D., Shrivastava H., Wolfinbarger J.M., Bostic J.P., Demko D.B., Elliott R.L., Stadler S.J., Carlson J.D., and Sutherland A.J.. 2007. Statewide monitoring of the mesoscale environment: a technical update on the Oklahoma Mesonet. J. Atmos. Oceanic Technol. 24:301–321.
Meisner C.A., and Karnok K.J.. 1992. Peanut root response to drought stress. Agronomy Journal 84:159–165.
Nageswara Rao R.C., Singh S., Sivakumar M.V.K., Srivastava K.L., and Williams J.H.. 1985. Effect of water deficit at different growth phases of peanut. I. Yield responses. Agron. J. 77:782–786.
Nageswara Rao R.C., Talwar H.S., and Wright G.C.. 2001. Rapid assessment of specific leaf area and leaf nitrogen in peanut (Arachis hypogaea L.) using a chlorophyll meter. Agron. J. 186:175–182.
Nageswara Rao R.C., Williams J.H., and Singh M.. 1989. Genotypic sensitivity to drought and yield potential of peanut. Agron. J. 81:887–893.
Nautiyal P.C., Rajgopal K., Zala P.V., Pujari D.S., Basu M., Dhadhal B.A., and Nandre B.M.. 2008. Evaluation of wild Arachis species for abiotic stress tolerance: I. Thermal stress and leaf water relations. Euphytica 159:43–57.
Nigam S.N., Chandra S., Sridevi K.R., Bhukta M., Reddy A.G.S., Rachaputi N.R., Wright G.C., Reddy P.V., Deshmukh M.P., and Mathur R.K.. 2005. Efficiency of physiological trait-based and empirical selection approaches for drought tolerance in groundnut. Ann. Appl. Biol. 146:433–439.
O’Toole J.C., Turner N.C., Namuco O.P., Dingkuhn M., and Gomez K.A.. 1984. Comparison of some crop water stress measurement methods. Crop Sci. 24:1121–1128.
Pandey M.K., Gangurde S.S., Sharma V., Pattanashetti S.K., Naidu G.K., Faye I., Hamidou F., Desmae H., Kane N.A., and Yuan M.. 2021. Improved genetic map identified major QTLs for drought tolerance-and iron deficiency tolerance-related traits in groundnut. Genes 12:37.
Pandey M.K., Upadhyaya H.D., Rathore A., Vadez V., Sheshshayee M.S., Sriswathi M., Govil M., Kumar A., Gowda M.V.C., Sharma S., Hamidou F., Kumar V.A., Kera P., Bhat R.S., Khan A.W., Singh S., Li H., Monyo E., Nadaf H.L., Mukri G., Jackson S.A , Guo B., Liang X., and Varshney R.K.. 2014. Genomewide association studies for 50 agronomic traits in peanut using the ‘reference set’ comprising 300 genotypes from 48 countries of semi-arid tropics of the world. PLoS One 9:e105228.
Ravi K., Vadez V., Isobe S., Mir R.R., Guo Y., Nigam S.N., Gowda M.V.C., Radhakrishnan T., Bertioli D.J., Knapp S.J., and Varshney R.K.. 2011. Identification of several small main-effect QTLs and a large number of epistatic QTLs for drought tolerance related traits in groundnut (Arachis hypogaea L.). Theor. Appl. Genet. 122:1119–1132.
Reed R., and Travis R.L.. 1987. Paraheliotropic leaf movements in mature alfalfa canopies. Crop Sci. 27:301-304.
Rosenberg N.J., Epstein D.J., Wang D., Vail L., Srinivasan R., and Arnold J. G.. 1999. Possible impacts of global warming on the hydrology of the Ogallala aquifer region. Clim. Change 42:677–692.
Rouse J.W., Haas R.H., Schell J.A., and Deering D.W.. 1974. Monitoring vegetation systems in the Great Plains with ERTS. Third ERTS Symposium. NASA SP-351, Washington DC. p. 309–317.
Rowland D.L., and Lamb M.C.. 2005. The effect of irrigation and genotype on carbon and nitrogen isotope composition in peanut (Arachis hypogaea L.) leaf tissue. Peanut Sci. 32:48–56.
Rowland D.L., Sorensen R.B., Butts C.L., and Faircloth W.H.. 2006. Determination of maturity and degree day indices and their success in predicting peanut maturity. Peanut Sci. 33:125–136.
Rucker K.S., Kvien C.K., Holbrook C.C., and Hook J.E.. 1995. Identification of peanut genotypes with improved drought avoidance traits. Peanut Sci. 22:14–18.
Sanders T.H., Blankenship P.D., Cole R.J., and Hill R.A.. 1985. Temperature relationships of peanut leaf canopy, stem, and fruit in soil of varying temperature and moisture. Peanut Sci. 12:86–89.
Sarkar S., Ramsey A.F., Cazenave A.-B., and Balota M.. 2021. Peanut leaf wilting estimation from RGB color indices and logistic models. Front. Plant Sci. 12:713.
Scanlon B.R., Faunt C.C., Longuevergne L., Reedy R.C., Alley W.M., McGuire V.L., and McMahon P.B.. 2012. Groundwater depletion and sustainability of irrigation in the US High Plains and Central Valley. Proc. Natl. Acad. Sci. U.S.A. 109:9320–9325.
Shekoofa A., Devi J.M., Sinclair T.R., Holbrook C.C., and Isleib T.G.. 2013. Divergence in drought-resistance traits among parents of recombinant peanut inbred lines. Crop Sci. 53:2569–2576.
Sheshshayee M.S., Bindumadhava H., Rachaputi N.R., Prasad T.G., Udayakumar M., Wright G.C., and Nigam S.N. 2006. Leaf chlorophyll concentration relates to transpiration efficiency in peanut. Ann. Appl. Biol. 148:7–15.
Singh D., Balota M., Isleib T.G., Collakova E., and Welbaum G.E.. 2014. Suitability of canopy temperature depression, specific leaf area, and SPAD chlorophyll reading for genotypic comparison of peanut grown in a sub-humid environment. Peanut Sci. 41:100–110.
Smith O.D., Simpson C.E., Grichar W.J., and Melouk H.A.. 1991. Registration of ‘Tamspan 90’ peanut. Crop Sci. 31:1711.
Songsri P., Jogloy S., Kesmala T., Vorasoot N., Akkasaeng C., Patanothai A., and Holbrook C.C.. 2008. Heritability of drought resistance traits and correlation of drought resistance and agronomic traits in peanut. Crop Sci. 48:2245–2253.
Strzepek K., Yohe G., Neumann J., and Boehlert B.. 2010. Characterizing changes in drought risk for the United States from climate change. Environ. Res. Lett. 5:044012.
Sullivan D.G., and Holbrook C.C.. 2007. Using ground-based reflectance measurements as selection criterion for drought and aflatoxin resistant peanut genotypes. Crop Sci. 47:423–432.
Upadhyaya H.D. 2005. Variability for drought resistance related traits in the mini core collection of peanut. Crop Sci. 45:1432–1440.
USDA ERS - Oil Crops Yearbook - Peanuts. 2021.
USDA-AMS. 2019. Farmers’ Stock Peanut Inspection Instructions. USDA-AMS.
Vergara-Diaz O., Kefauver S.C., Elazab A., Nieto-Taladriz M.T., and Araus J.L.. 2015. Grain yield losses in yellow-rusted durum wheat estimated using digital and conventional parameters under field conditions. Crop J. 3:200–210.
Wang X., Yang X., Feng Y., Dang P., Wang W., Graze R., Clevenger J.P., Chu Y., Ozias-Akins P., Holbrook C., and Chen C.. 2021. Transcriptome profile reveals drought-induced genes preferentially expressed in response to water deficit in cultivated peanut (Arachis hypogaea L.). Front. Plant Sci. 12:555.
Notes
- First, tenth, and eleventh authors: Research Plant Pathologist, Research Biologist, and Research Geneticist, USDA-ARS, Stillwater, OK 74075; Second and fourth authors: Professor and Technician II, Texas A&M AgriLife Research, Lubbock, TX 79403; Second, and sixth authors, Professor and Graduate Student, Dept. Plant and Soil Science., Texas Tech University, Lubbock, TX 79409; Third and fifth authors: Professor and Former Graduate Student, Tidewater Agricultural Research and Extension Center, Virginia Tech, Suffolk, VA 23437; Seventh author, Chair and Professor, Dept. Biomedical Sciences, Rocky Vista University, Parker, CO, 80134; Eighth author, Professor, Dept. Biosystems and Agricultural Engineering, Oklahoma State University, OK74078; Ninth author, Research Plant Physiologist, USDA-ARS, Lubbock, TX 79415. [^] Corresponding author’s E-mail: rebecca.bennett@usda.gov