Introduction
Peanut (Arachis hypogaea L.) is an important crop for the United States, especially in Georgia. In 2011, peanut was harvested from more than 192,000 ha in Georgia, and yields averaged 4171 kg/ ha with a total production value that exceeded $586 million (Williams-Woodward, 2013), while in 2014, average peanut yields were 4595 kg/ ha in Georgia, harvested from more than 239,000 ha (USDA - NASS, 2015). Peanut production can be threatened by a variety of diseases that can affect all parts of the peanut plant and reduce the quality and quantity of pods and seeds. Foliar diseases such as tomato spotted wilt, caused by the Tomato spotted wilt tospovirus, early and late leaf spots, caused by Cercospora arachidicola S. Hori and Cercosporidium personatum (Berk & M. A. Curtis Deighton), respectively (Bellgard and Ham, 2004, Damicone and Melouk, 2010), can cause considerable yield losses and consequently substantial economic losses. Yield loss due to damage and increased management costs of these three foliar diseases in 2011 in Georgia was estimated at $32.2 million (Williams-Woodward, 2013).
Peanut rust, caused by the fungus Puccinia arachidis Speg., is another damaging foliar disease in peanut cultivation that is common in countries with warm, tropical climates, but is relatively rare in the United States (Bromfield, 1971, Subrahmanyam et al., 1985). Peanut rust was first reported in Georgia in 1953, but it was indicated in that report that rust was not expected to become a serious problem in Georgia (Rankin, 1953) since the pathogen cannot survive the cold winter temperatures. However, peanut rust did become an important problem in southern Texas in 1965 (Van Arsdel and Harrison, 1972). Under normal cultivation conditions, yield losses to peanut rust can be considerable, with losses as high as 50% reported in India (Subrahmanyam, et al., 1983abc). Currently in the U.S., damage due to peanut rust is typically localized, so little information on yield losses in the U.S. is available.
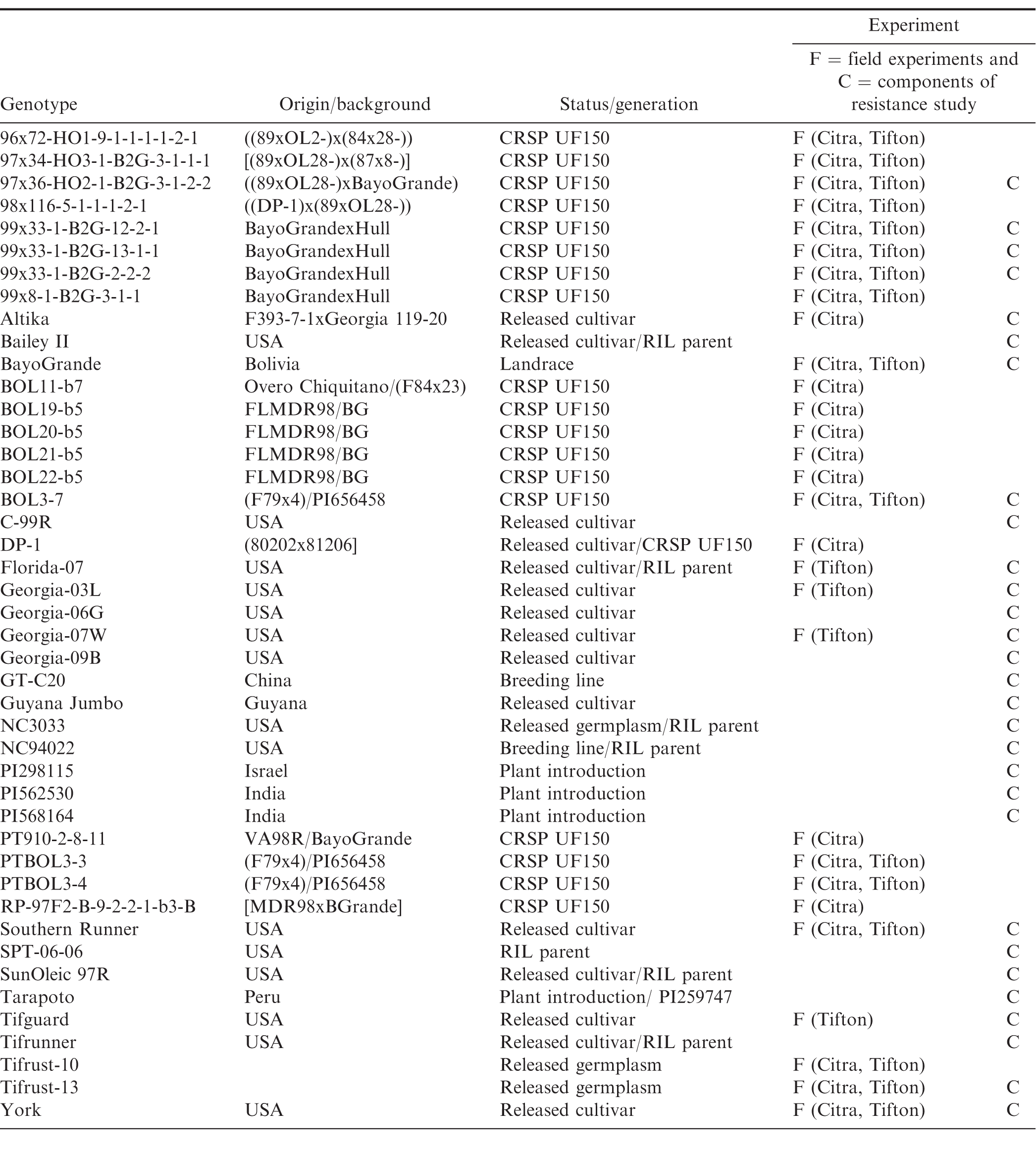
Management methods for peanut rust in regions where this disease is prevalent (developing countries such as India, Haiti, and Guyana) include cultural practices such as eradicating volunteer plants to reduce the inoculum source, and allowing fallow periods of at least one month between crops (Melouk and Shokes, 1995, Subrahmanyam, 1997). Multiple fungicide applications throughout the season (Bromfield, 1971, Subrahmanyam, 1997, Subrahmanyam et al., 1985) with chlorothalonil, strobilurins, and triazoles are effective; however, chemical control is often not an option in developing countries due to increased production costs or lack of availability. The pathogen may also develop resistance with frequent fungicide applications (Smith and Littrell, 1980) although the risk is not the same for the different chemical groups. The use of resistant peanut cultivars is thus a very desirable management approach. Many germplasm accessions were screened for resistance during the late 1970s through the early 1990s, resulting in the identification of several peanut accessions with resistance to peanut rust (Bromfield, 1971, Subrahmanyam et al, 1989, 1997, Wynne et al., 1991), with sources of resistance mainly originated from Peru, Bolivia, and India (Bromfield, 1971, Melouk and Shokes, 1995, Subrahmanyam et al, 1989, 1997). Little new information on rust resistance has become available in the last two decades. Recently, several breeding lines have been developed in the UF150 project of the Peanut Collaborative Research and Support Program (Peanut CRSP) as part of the United States Agency for International Development (USAID). These CRSP breeding lines were developed from crosses with parents including a Bolivian landrace cultivar, 'BayoGrande'; U.S. cultivars 'Hull' and 'Florida MDR98'; plant introduction (PI) PI656458, and the breeding lines VA98R and F79x4. The CRSP breeding lines are currently being screened for multiple disease resistance in the United States and multiple peanut producing countries in the western hemisphere (Table 1).
Resistance to P. arachidis typically is quantitative, where multiple components of resistance provide varying levels of partial resistance, leading to a reduced rate of the rust epidemic development. Components of peanut rust resistance that have been described include increased incubation and latent periods, reduced infection frequency, pustule size, percent diseased area, spore production, and spore germination (Bromfield, 1971, Cook, 1980, Subrahmanyam et al., 1983c). These components were characterized in the 1980s and early 1990s (Cook, 1980, Subrahmanyam, et al., 1983c), but little work has been reported on the more recently developed breeding lines.
The objectives of this study were to determine the level of field resistance to peanut rust and components of resistance of (a) newly developed peanut CRSP breeding lines with Bolivian genetic background, (b) commonly grown peanut cultivars in Georgia, and (c) parents of existing breeding populations. Part of this research has been reported previously (Power et al., 2013), as part of a symposium proceedings paper, in which field studies conducted in 2010 and 2011 were included, as well as growth chamber studies that lasted 31 d. The field experiments in this current study were carried out from 2010 to 2013, and the methods for the growth chamber studies were adjusted so that data from 16 d were included.
Materials and Methods
Field studies
Field studies were conducted at the University of Florida Plant Science and Education Unit, in Citra, FL (29.41°, -82.11°) in 2010, 2011 and 2012, and at the University of Georgia Coastal Plain Experiment Station, in Tifton, GA (31.48°, -83.51°) in 2011, 2012 and 2013, to evaluate field resistance of the breeding lines. Soil type in Citra, FL was fine sand (loamy, siliceous, semiactive, hyperthermic grossarenic paleudult), and in Tifton, GA was loamy sand (fine-loamy, kaolinitic, thermic Plinthic Kandiudults). A randomized complete block design was used for the experiments with three replications from 2010 through 2012, and four replications in 2013. Three and four replications were used because of the seed availability of the breeding lines. In Citra, 25 genotypes (Table 1) were planted on 25 May 2010, 1 June 2011 and 31 May in 2012. In Tifton, 19 genotypes (Table 1) were planted on 27 June 2011, and 20 genotypes were planted on 22 June 2012 and 11 June 2013.
Peanut seeds were planted at a rate of 20 seed/m in two-row plots bordered by cultivars Florida-07, Tufrunner-727, or Georgia-09B. The plots were 6 m long and 1.8 m wide with 0.91 m between rows. In both locations, leaf spot epidemics were suppressed by alternating sprays with flusilazole (Punch, DuPont de Nemours, Wilmington, DE) and thiophanate methyl (Topsin 4.5FL, United Phosphous, Inc., King of Prussia, PA). Applications were on a 14-d schedule starting approximately 40 d after planting (DAP), with 6 to 7 sprays per season. These fungicides have no activity against P. arachidis (personal communication, A. K. Culbreath). Peanuts in all experimental plots at both locations and in all three years were inoculated by brushing plants in the experimental plots with heavily rust-infected peanut plants collected from a nearby earlier planted field. In Citra, plants were inoculated on 25 August 2010, 20 September 2011, and 29 August 2012, and in Tifton on 22 and 23 September and 18 October 2011, 6 and 20 September 2012, and on 23 October 2013. Plants in all plots were also inoculated at night time in Tifton, when the leaves were closed, with a urediniospore suspension of approximately 10,000 spores/ml on 25 August 2012 and 7 September 2013, and in Citra on 12 September 2012 during the day with a urediniospore suspension of approximately 1,000 spores/ml. The suspension used for the inoculation was prepared by adding vacuumed urediniospores from locally collected, heavily rust-infected peanut plants, to 0.005% Tween 20 solution. The suspension was sprayed using a pump-up air hand sprayer.
Rust severity was determined using a modified nine-point ICRISAT scale based on lesion density and leaf necrosis (Subrahmanyam et al, 1995): 1 = no disease (0% severity); 2 = sparsely distributed lesions, primarily on lower leaves (1-5% severity); 3 = many lesions on the lower leaves with evident necrosis and very few lesions on middle and upper leaves (6-10% severity); 4 = numerous lesions on lower and middle, with severe necrosis on lower leaves (11-20% severity); 5 = severe necrosis of middle and lower leaves and fewer lesions on top leaves (21-30% severity); 6 = extensive damage to lower leaves, lesions densely present on middle leaves with necrosis and lesions may be on top leaves as well (31-40% severity); 7 = severe damage to lower and middle leaves and lesions are densely distributed on top leaves (41-60% severity); 8 = 100% damage to lower and middle leaves and lesions on top leaves with severe necrosis (61-80% severity) and 9 = almost all leaves are withering and bare stems are present (81-100% severity). In Citra, peanut rust severity was evaluated 108, 122, 136 DAP in 2010; 112, 118, 126 DAP in 2011; and 96, 110, 117, 124, 131, 138, DAP, in 2012. In Tifton, disease severity was assessed 123, 130, 137 DAP in 2011; 101, 122, 129, 136, 140 DAP in 2012; and 134, 147, 154 DAP in 2013. Despite the fungicide applications for leaf spot suppression, late leaf spot pressure was high in Citra, so leaf spot severity was assessed on 136 DAP in 2010, 126 DAP in 2011 and 124, 131 and 138 DAP in 2012, using the 1-10 ICRISAT scale (2010) and the 1-10 Florida scale (2011 and 2012) (Chiteka et al., 1988).
Rust severity data were used to calculate area under the disease progress curve (AUDPC) for each plot (Shaner and Finney, 1977). The number of d between disease assessments differed per year and per location, so AUDPC values were standardized (stAUDPC) by dividing the AUDPC by the number of d between the first and last assessment date. The effects of genotype on stAUDPC and final disease severity were analyzed using the Proc MIXED with ddfm = satterth option on the model statement (SAS v 9.3, SAS Institute Inc., Cary, NC). Year, rep, and the interaction with these factors were treated as random effects, and genotype as fixed effect. Fisher's protected LSD (P < 0.05) was used to determine significant differences in stAUDPC and final disease severity among genotypes.
Components of resistance
To assess the components of resistance, a detached leaf experiment was carried out as described by Cook (1980). A single-pustule isolate was developed by harvesting urediniospores with a vacuum pump from peanut leaves collected from fields in Georgia, and inoculating healthy leaves with urediniospores from a single pustule. This cycle was repeated several times to ensure purified single-pustule isolates. Urediniospores were maintained on leaves of the susceptible cultivar 'Altika' that were placed on 15-cm Petri dishes with 10% water agar, and incubated at 25 C. Freshly produced urediniospores were collected from these leaves to prepare a spore suspension of 40,000 spores/mL of 0.005% Tween 20, which was quantified using a hemacytometer. Percentage germination of the urediniospores was assessed 1 day before the start of the experiments, and on the day of the experiments. A spore suspension was sprayed on 10% water agar plates, and incubated at 25 C in darkness overnight. The next day, the number of germinated urediniospores out of 50 randomly chosen urediniospores was determined using a compound microscope at 100x magnification. Urediniospores were considered germinated if the germination tube was longer than the greatest diameter of the urediniospore.
The peanut genotypes used in this study included currently grown cultivars, CRSP breeding lines, ICRISAT plant introductions, and parents of existing recombinant inbred line (RIL) populations that are currently being screened for multiple disease resistance (Table 1). Peanut plants were grown from seed in the greenhouse at 25 C, in 15-cm pots filled with commercial potting soil (Sunshine Professional Growing Mix, Sun Gro Horticulture Distribution Inc, Bellevue, WA), and were watered as needed. Four or five seeds treated with a commercial seed treatment consisting of 45% captan, 15% pentachloronitrobenzene and 10% carboxin (Vitavax PC, Bayer CropScience LP, Research Triangle Park, NC) were planted in each pot. The youngest fully expanded leaves of 5 to 6 wk-old plants were collected, the leaflets detached, and placed on sterile moistened filter paper in a Petri dish (9-cm diameter) with the abaxial side up. The leaflets were inoculated by spraying them for 1 sec using a compressed air sprayer containing the uredinial spore suspension. The experimental design was a randomized complete block with three replicates per genotype, and for each genotype a control was included by spraying leaves with a sterile 0.005% Tween 20 solution. The Petri dishes containing inoculated leaflets were incubated in darkness for 16 hr at 25 C. After the 16-hr dark period, the closed Petri dishes were incubated at 25 C, with a 12-hr photoperiod for 16 d. The filter paper was kept moist with sterile distilled water. The leaflets were examined on 7, 11, and 16 d after inoculation (DAI) for the development of pustules.
The components of resistance measured included: latent period, infection frequency, lesion diameter, and percent diseased area. The latent period was determined by counting the number of d between inoculation and spore production of at least one pustule. The infection frequency was determined as the number of pustules/cm2 of leaf area, and lesion diameter was determined by measuring the lesion area of 10 arbitrarily selected lesions and calculating the mean lesion diameter from:
Leaf area, lesion area, and percent diseased area were measured from digital images of leaves that were taken 16 DAI, using the ASSESS 2.0 Image Analysis Software for plant disease quantification (APS Press, St. Paul, MN). For ASSESS analyses, the leaflets were glued to blue paper background, covered with a sheer plastic sheet, scanned at 300 dpi, and stored as TIFF files. The detached leaf experiment was carried out three times.
The effects of genotype on the components of resistance were analyzed using the Proc MIXED with ddfm = satterth option on the model statement (SAS v 9.3, SAS Institute Inc., Cary, NC). Trial, rep, and the interaction with these factors were treated as random effects, and genotype as fixed effect. Fisher's protected LSD (P < 0.05) was used to determine significant differences in the components of resistance among genotypes. The correlation of these components from the growth chamber assays with stAUDPC and the final disease severity from the field experiments was determined by calculating the correlation coefficient at (P ≤ 0.05) with the Proc CORR procedure.
Greenhouse evaluations
The same genotypes used in the components of resistance study were used for this study. A randomized complete block design was used with four replicates per genotype, and the experiment was repeated twice. Three to 4 wk after planting as described previously, individual plants were transferred to cones. The plants were inoculated 5 to 6 wk after planting by spraying leaves (with previously described inoculum) until covered completely with the spore suspension. Effort was made to direct sprays on the underside of leaves. The inoculated plants were placed in a moist chamber constructed of PVC pipe and covered with black plastic (4 m x 1.5 m x 1.5 m, L, W, H), and incubated in darkness for 16 h at 25 C. Humidifiers were used to keep the leaves wet and the humidity high (> 90%). The youngest fully developed leaf was labeled, and at 21 DAI the labeled leaves were processed for ASSESS 2.0 analysis as described above. The components of resistance measured for the labeled leaves were infection frequency, lesion diameter, and percent diseased area, and they were determined as described above.
The effects of genotype on the components of resistance were analyzed using the Proc MIXED with ddfm = satterth option on the model statement (SAS v 9.3, SAS Institute Inc., Cary, NC). Trial, rep, and the interaction with these factors were treated as random effects, and genotype as fixed effect. Fisher's protected LSD (P ≤ 0.05) was used to determine significant differences in the components of resistance among genotypes.
Results and Discussion
Field evaluations of rust resistance
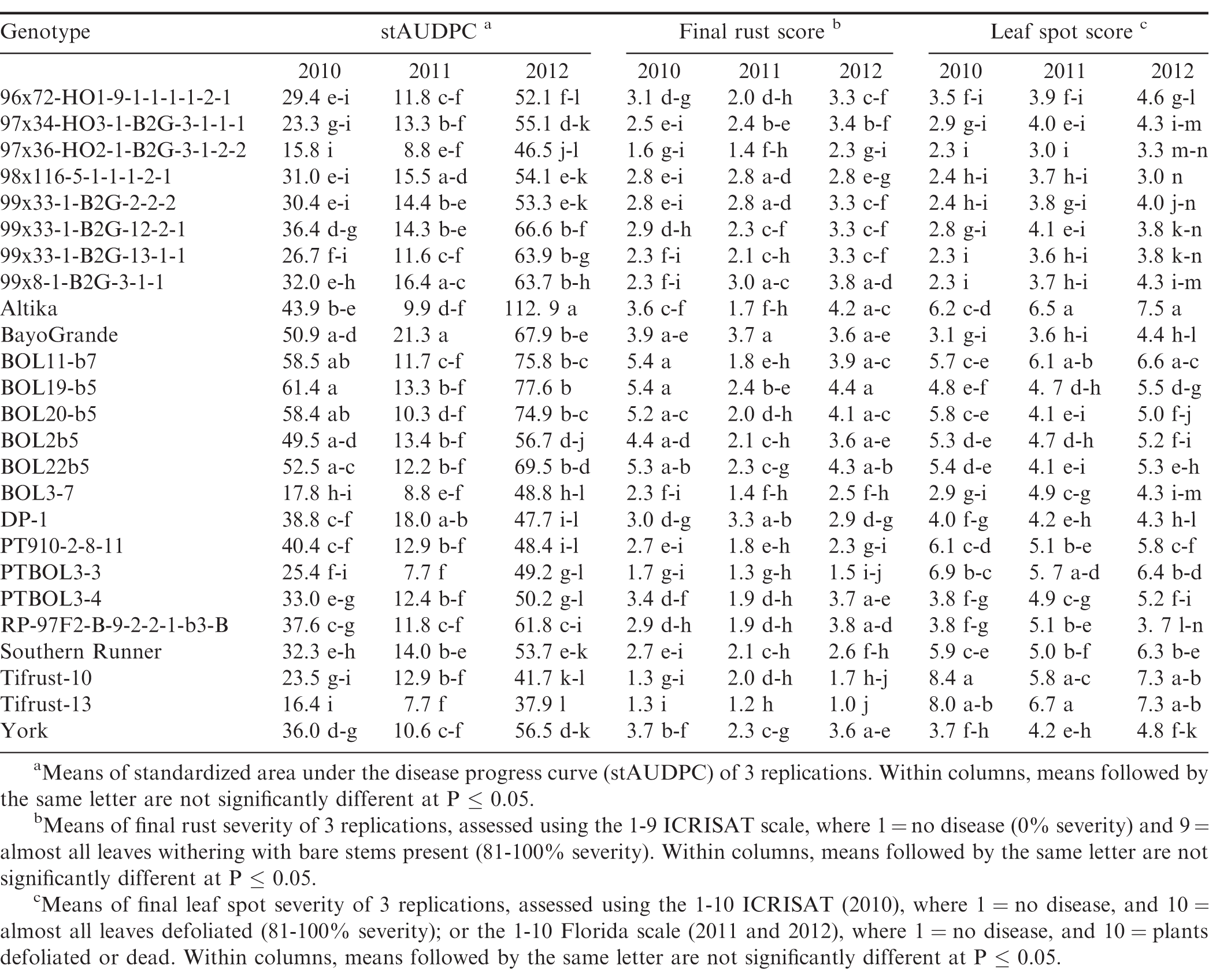
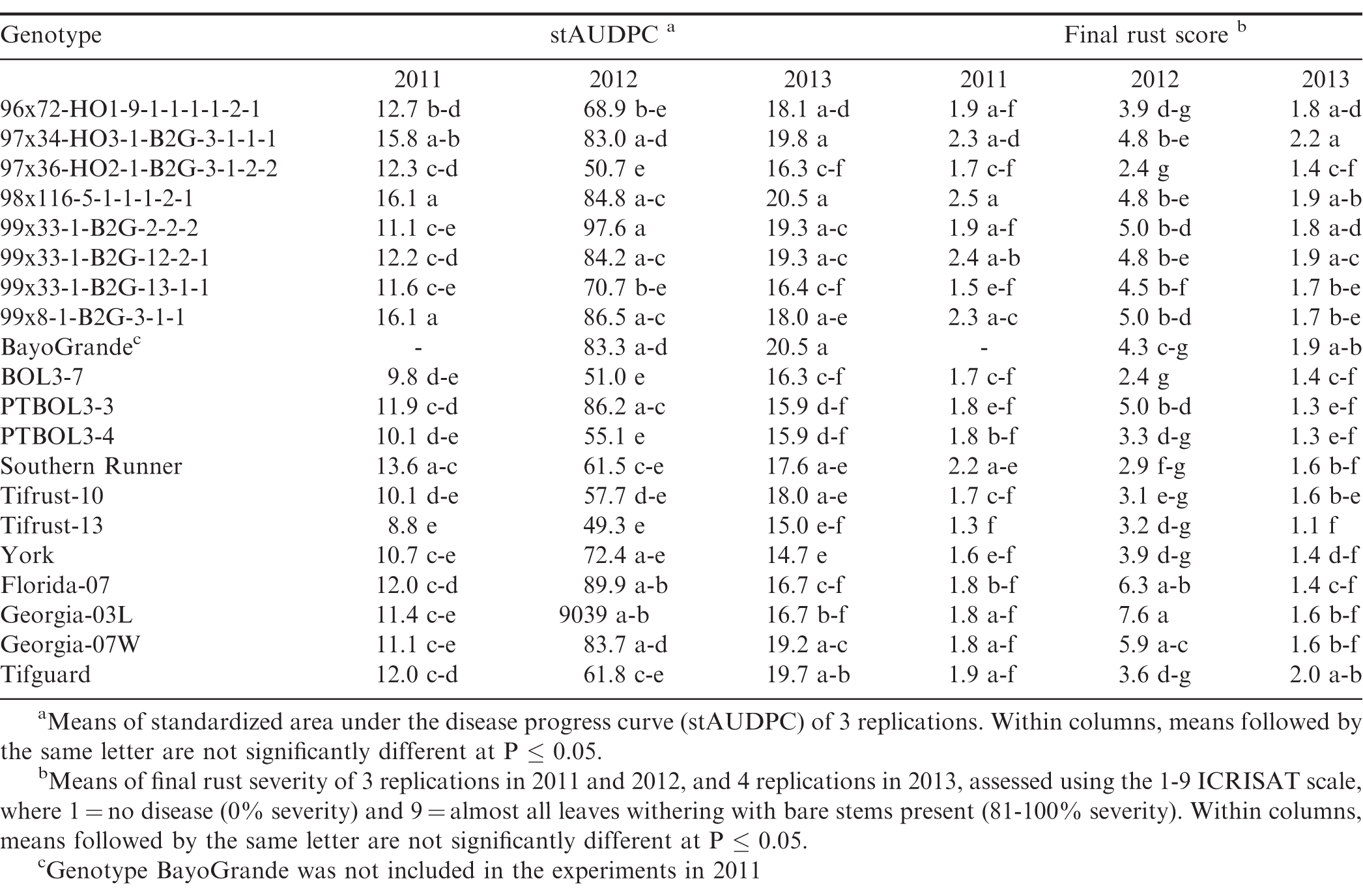
There were significant year × genotype interactions (P ≤ 0.001) for rust severity and stAUDPC among years, and the experiments in the different locations were not identical with respect to genotypes included, so each experiment was analyzed separately. Rust was present in both locations and in each experimental year, but severity of the epidemics varied among years and between locations. Development of epidemics depends on the introduction of inoculum, usually by tropical storms, and subsequent environmental conditions that are conducive for disease development, such as warm temperatures, rainfall, and high humidity. This was the case in 2010 and 2012, but not in 2011 and 2013, in both Citra and Tifton. In Citra, rust was first observed approximately 90 to 95 DAP in 2010 through 2012, while in Tifton rust epidemics started 60 to 70 DAP in 2011 through 2013. There were significant differences (P ≤ 0.05) in stAUDPC and final rust severity among the genotypes (Tables 2, 3). In Citra, the disease pressure was high in 2010 and 2012 due to favorable weather conditions. In 2011, disease pressure was low in Citra and Tifton, due to very dry and hot weather conditions. This resulted in few differences among genotypes for either final rust severity or stAUDPC. Moreover, the epidemic started too late in 2011 in Tifton and as a result the final disease severity was too low to distinguish among genotypes. Due to wet and cold weather in Tifton in 2013, final disease severity was too low to distinguish resistant from susceptible genotypes. No immunity to rust was noted, as all genotypes were infected in all experimental years and in both locations.
Genotypes Tifrust-10 (Hammons et al., 1982b), Tifrust-13 (Hammons et al., 1982a), two rust-resistant standards, and PTBOL3-3 were among the genotypes with the lowest stAUDPC and final disease severity in all 3 years in Citra, and in Tifton in 2012, however, they had high late leaf spot (caused by Cercosporidium personatum) severity ratings, with more than 90% defoliation at harvest, in Citra (Tables 2, 3). Although these genotypes show little promise for use for peanut production, they may be of use in breeding for rust resistance. CRSP breeding lines 97x36-HO2-1-B2G-3-1-2-2 and BOL3-7 appeared to be among the most resistant to both rust and late leaf spot, as these breeding lines had low stAUDPC and low final disease severity values for both diseases in all 3 years in Citra, and in 2012 in Tifton (Tables 2, 3). These genotypes show potential to be developed into cultivars, since they appear to have multiple disease resistance. The cultivars Altika and BayoGrande, and CRSP breeding line BOL19-b5 had the highest stAUDPC for rust and high final leaf spot severity in 2010, 2011, and 2012 (Table 2). The cultivars Southern Runner (Black and Smith, 1987) and York (personal communication B. L. Tillman) are partially resistant to rust, and Tifguard is moderately resistant to rust as well (personal communication A. K. Culbreath). Georgia-03L, a cultivar observed to have some resistance to rust in Nicaragua and Haiti (field experiments in 2010, personal communication T. B. Brenneman), appeared to be very susceptible in Tifton in 2012. Georgia-07W, a cultivar currently grown in Georgia with resistance to stem rot (caused by Sclerotium rolfsii), did not appear to be resistant to rust, as it had high stAUDPC and final disease severity scores.
The environmental conditions (frequent rains and hot summer temperatures) led to very conducive conditions for peanut rust development in both Citra and Tifton in 2012, resulting in relatively high disease pressure by the end of the season. The resistance in some of the breeding lines was not as effective under this high disease pressure; however, breeding lines 97x36-HO2-1-B2G3-1-2-2 and BOL3-7 maintained resistance in both Citra and Tifton in 2012. Breeding line PTBOL3-3 had a low final rust severity rate in Citra, but had one of the higher scores at Tifton in that same year.
Components of rust resistance
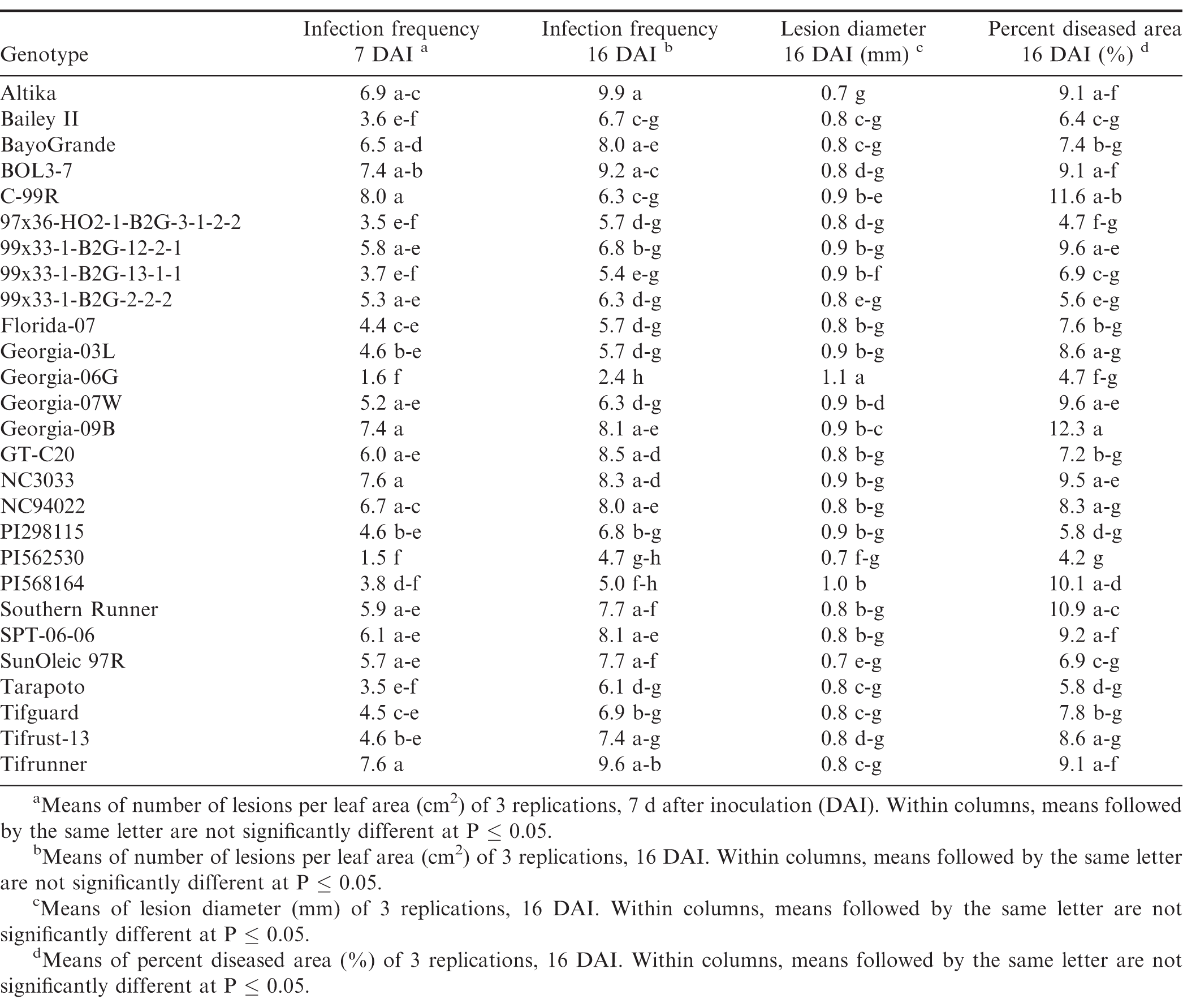
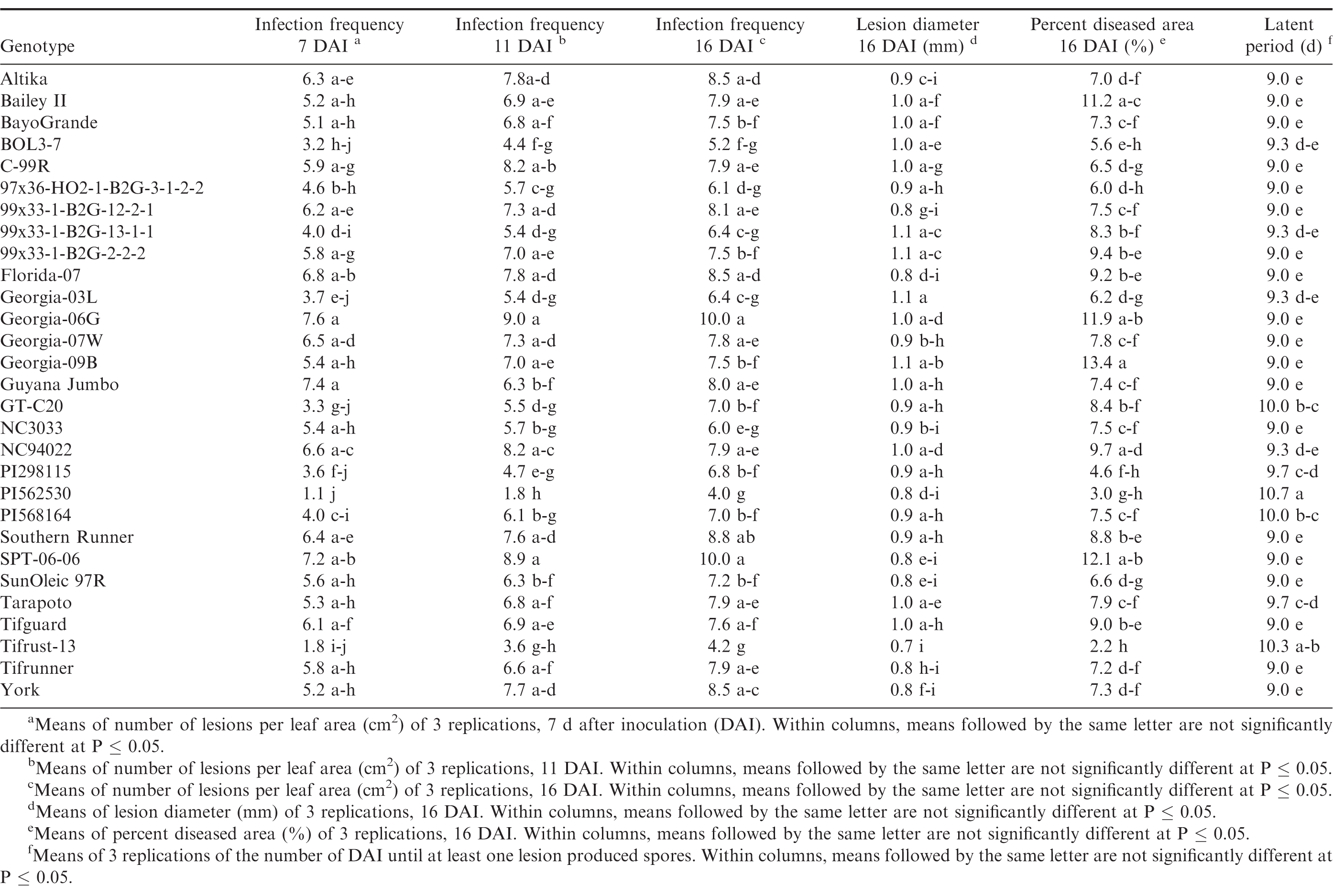
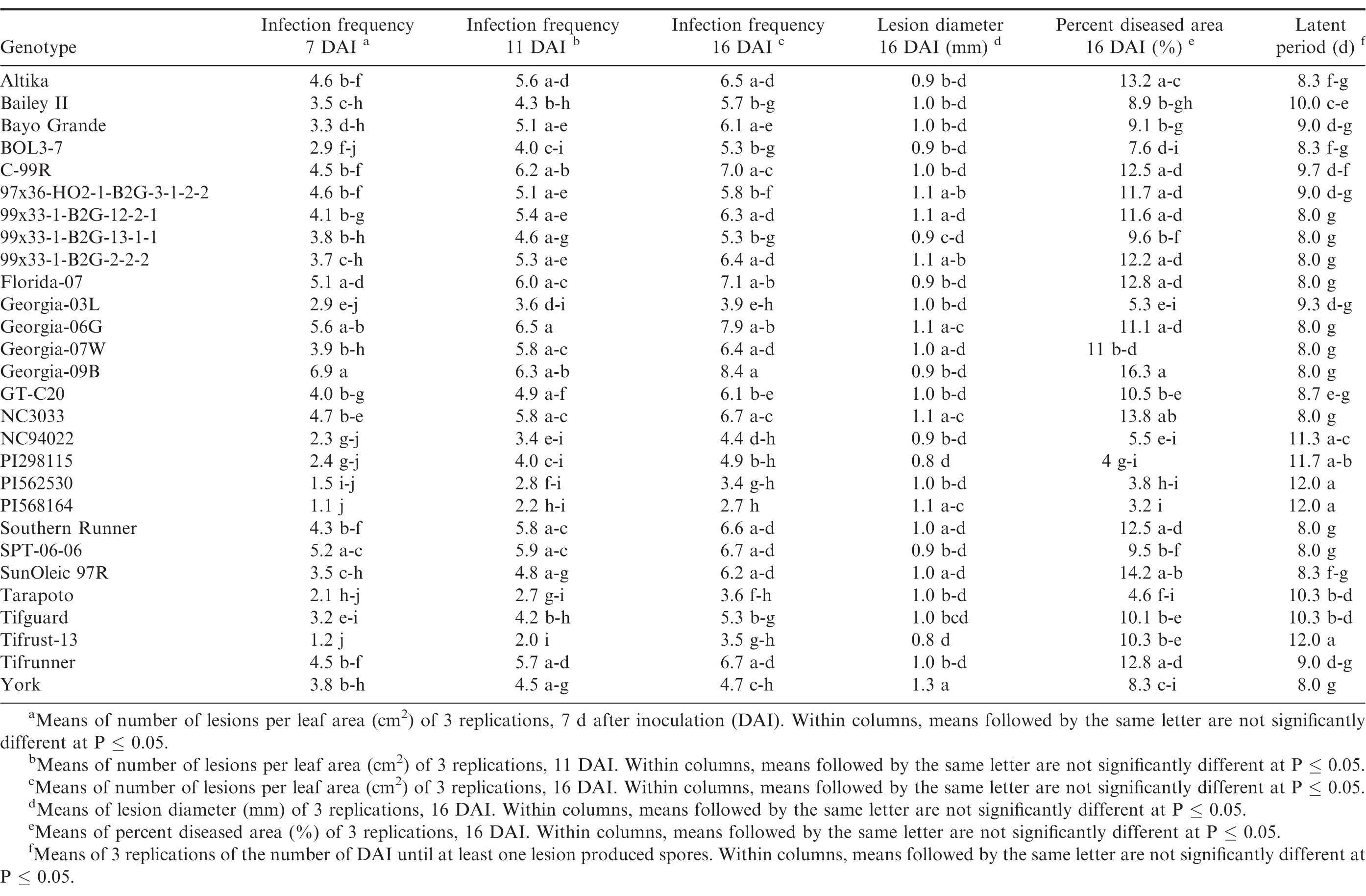
Germination of the urediniospores was higher than 90% for all three trials. Pustules developed on all inoculated leaves, and no pustules were present on the control leaves in any of the repeated studies. There were significant trial × genotype interactions for infection frequency at 7 and 16 DAI, percent diseased area, lesion diameter and latent period, so each experiment was analyzed separately. There were significant differences (P ≤ 0.05) among the genotypes for all of the components measured in trial 1 (Table 4) and trial 2 (Table 5). Differences among the genotypes were significant (P ≤ 0.05) for infection frequency at 7, 11 and 16 DAI, percent diseased area, and latent period in the third trial, but not for lesion diameter (P = 0.25) (Table 6). In general, genotypes that had high infection frequencies at 7 DAI also had high infection frequencies at the end of the experiment, higher percent diseased area, and shorter latent periods. On the other hand, genotypes with longer latent periods generally had lower infection frequencies at 7, 11 and 16 DAI, and smaller percent diseased areas. Georgia-09B, NC3033, BayoGrande, and C99R were among the genotypes with higher infection frequencies, larger percent diseased areas, and shorter latent periods in most of the experimental trials (Tables 4, 5, and 6). In trials 2 and 3 (Table 5 and 6, respectively), SPT-06-06 and Florida-07 also were among genotypes with higher infection frequencies, larger percent diseased areas, and shorter latent periods. Tifrust-13, PI568164, PI562530, typically were among those with lower infection frequencies, smaller percent diseased areas, and longer latent periods.
High values were obtained for infection frequency and percent disease area in the growth chamber experiments; infection frequency in the majority of the genotypes reached about 50% by 7 DAI, even though these pustules were very young at that time point. In contrast to these findings, Mehan et al. (1994) and Subramanyam et al. (1983ab) reported incubation periods (time from inoculation to 50% of the pustules developed) ranging from 8.5 or 9 d for the highly susceptible genotypes, to 18 or 19 d for the resistant ones. The differences between results herein and previous studies may be due to inconsistencies in the Tween concentration used to prepare the inoculum. Mehan et al. (1994) used a 0.1% Tween solution and Subramanyam et al. (1983ab) added "a few drops of Tween 80" to sterile water. In earlier experiments of this study (Power et al., 2013) a 0.1% Tween 20 solution was used for the inoculum suspension and few lesions developed in the growth chamber studies at 20 and 31 DAI. Although optimum conditions for peanut rust development were maintained in the experiments, differences between results herein and previous studies may also be due to conditions such as photoperiod and plant age. These studies corroborate previous reports of rust resistance in genotypes such as Tarapoto, PI298115, PI568164, PI562530, Tifrust-10 and Tifrust-13 (Hammons et al., 1982ab, Subrahmanyam et al., 1983abc, 1995). These genotypes were among those with the lower infection frequencies, percent disease areas, and smaller lesion diameters.
Greenhouse evaluations of rust resistance
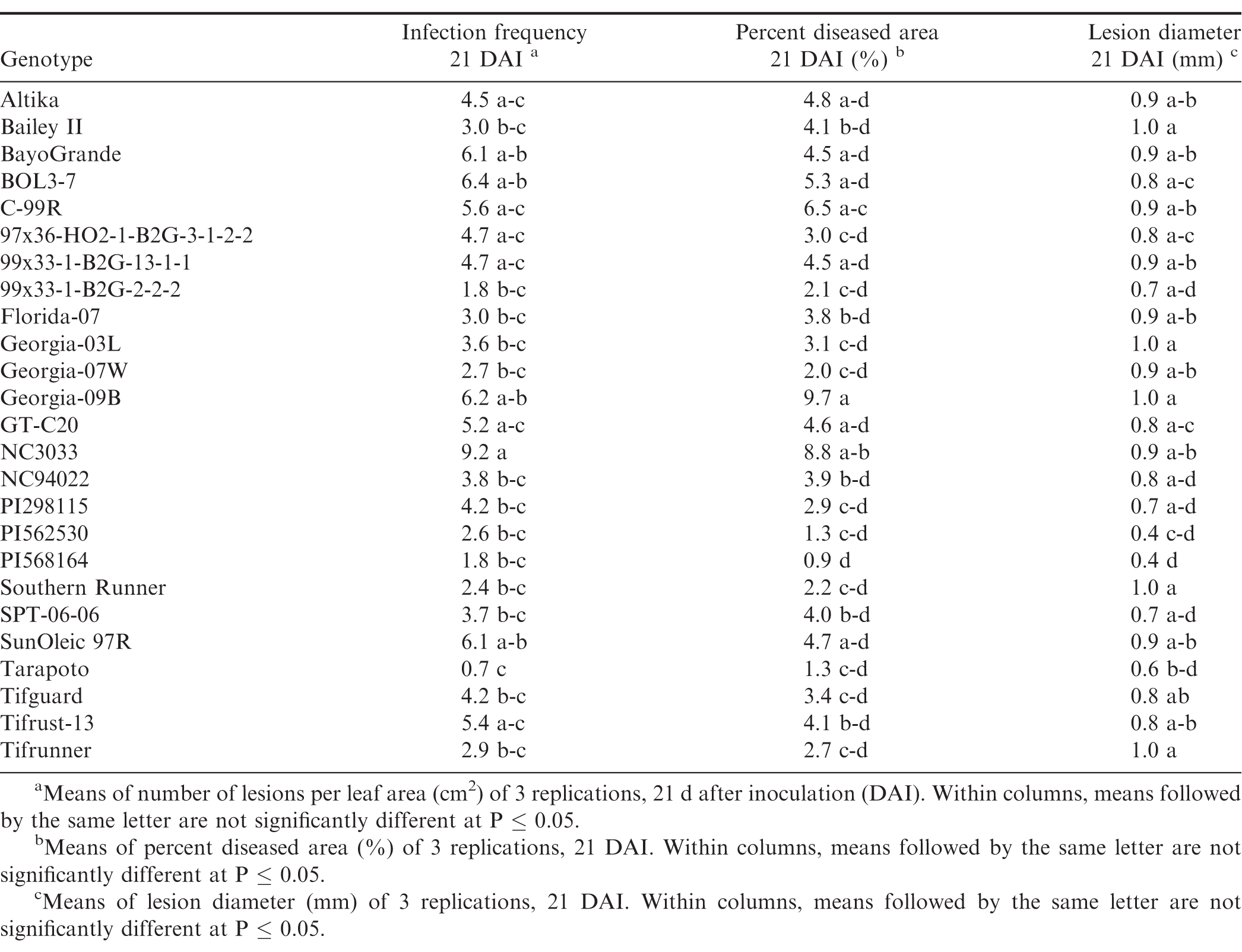
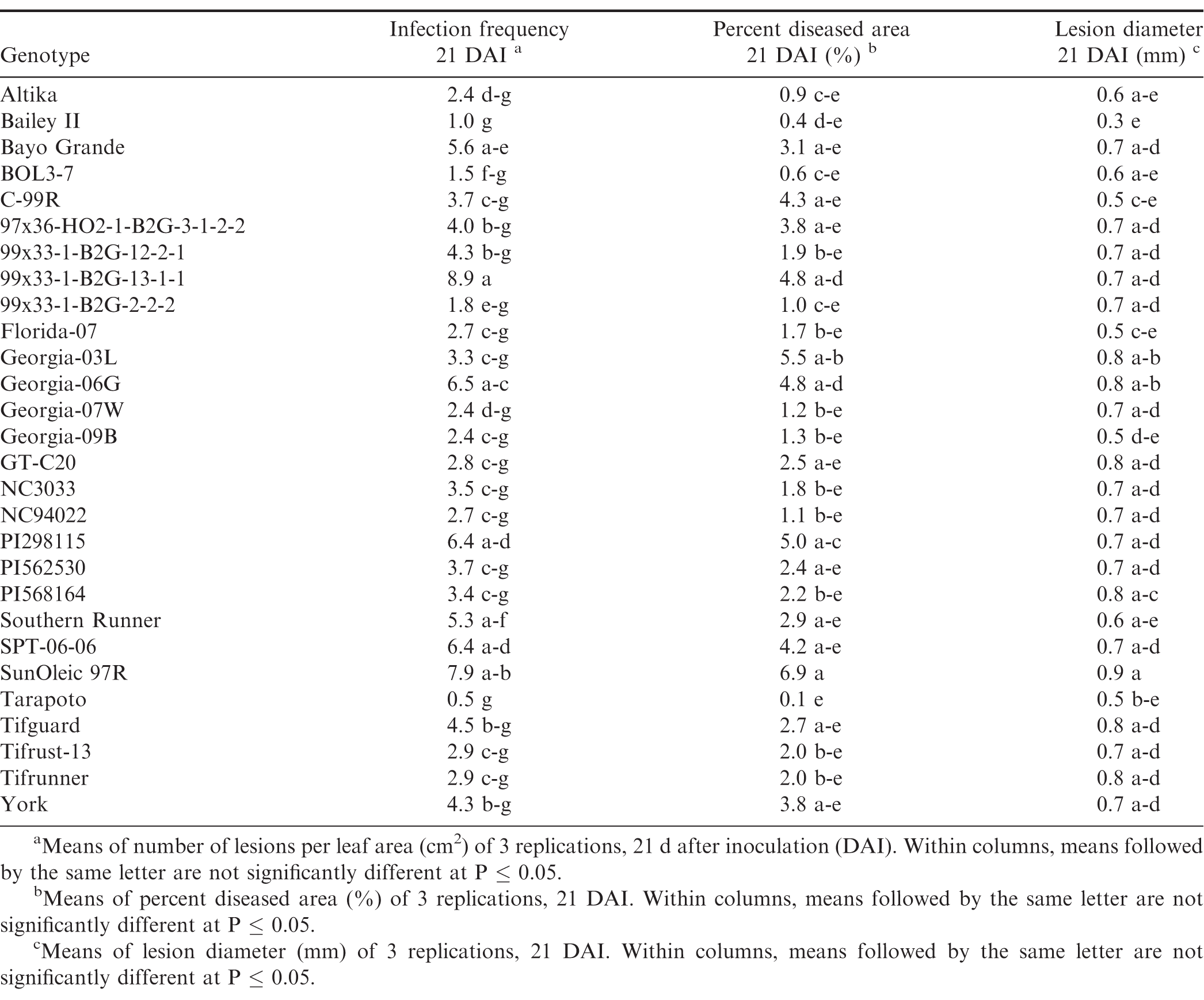
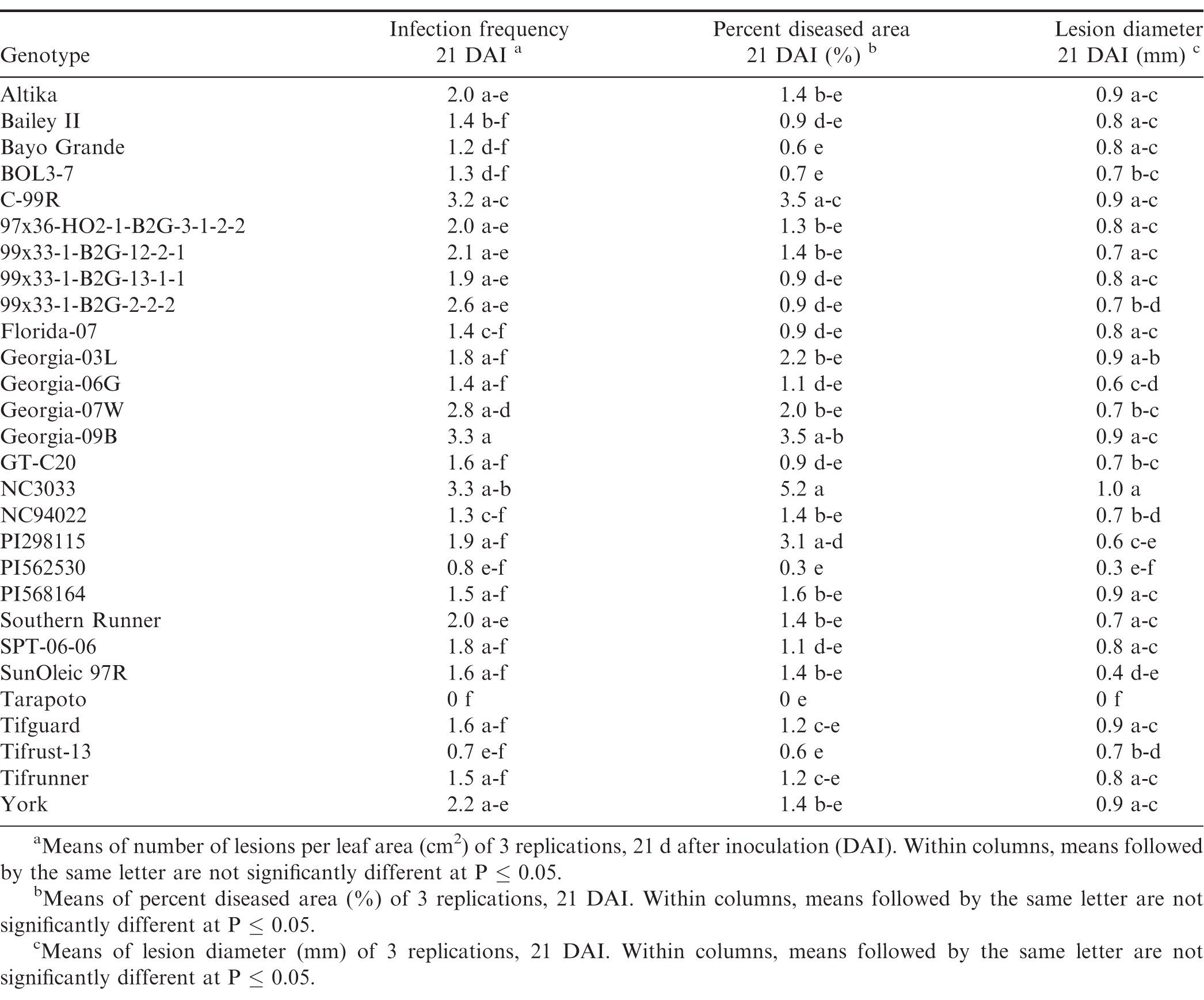
There were significant trial × genotype interactions for the components between repeated trials, so each experiment was analyzed separately. In all three trials (Tables 7, 8, and 9), pustules developed on all inoculated leaves, except for Tarapoto in the third trial. There were no significant differences among genotypes for infection frequency (P = 0.21), lesion diameter (P = 0.06) or percent diseased area (P = 0.08) in the first trial (Table 7), however, based on Fisher's protected LSD, genotypes NC3033, and Georgia-09B had significant highest infection frequency scores, while these scores were significantly lower for PI568164 and Tarapoto. PI568164 had also the smallest percent diseased area and lesion diameter. Differences among genotypes were significant for infection frequency (P = 0.05), but not for lesion diameter (P = 0.31), or percent diseased area (P = 0.34) in the second trail (Table 8), but based on Fisher's protected LSD, genotype SunOleic 97R had the largest percent diseased area and lesion diameter, while Bailey II had the smallest lesion diameter. In the third trial (Table 9) differences among genotypes were significant for lesion diameter (P = 0.005) and percent diseased area (P = 0.009), but not for infection frequency (P = 0.34), but based on Fisher's protected LSD, genotype Bailey II had the lowest infection frequency. Although not all genotype differences were significant, the trend was similar to that observed in the detached leaf studies, in which genotypes NC3033, Georgia-09B, and C-99R were among those with the highest infection frequency 21 DAI in all three trials (Tables 7, 8, and 9) and percent diseased area in the first (Table 7) and third trial (Table 9). Furthermore, Tarapoto, PI562530, PI568164, and 99x33-1-B2G-2-2-2 were among genotypes with the lowest infection frequency and percent diseased area 21 DAI. Resistance ratings were inconsistent for field-resistant CRSP breeding lines BOL3-7, 97x36-HO2-1-B2G3-1-2-2 and 99x33-1-B2G-13-1-1, and rust-resistant standards Tifrust-13 and PI298115; infection frequencies and percent diseased areas were highly variable for these genotypes in all the greenhouse experiment trials.
Correlations between field resistance and components of resistance
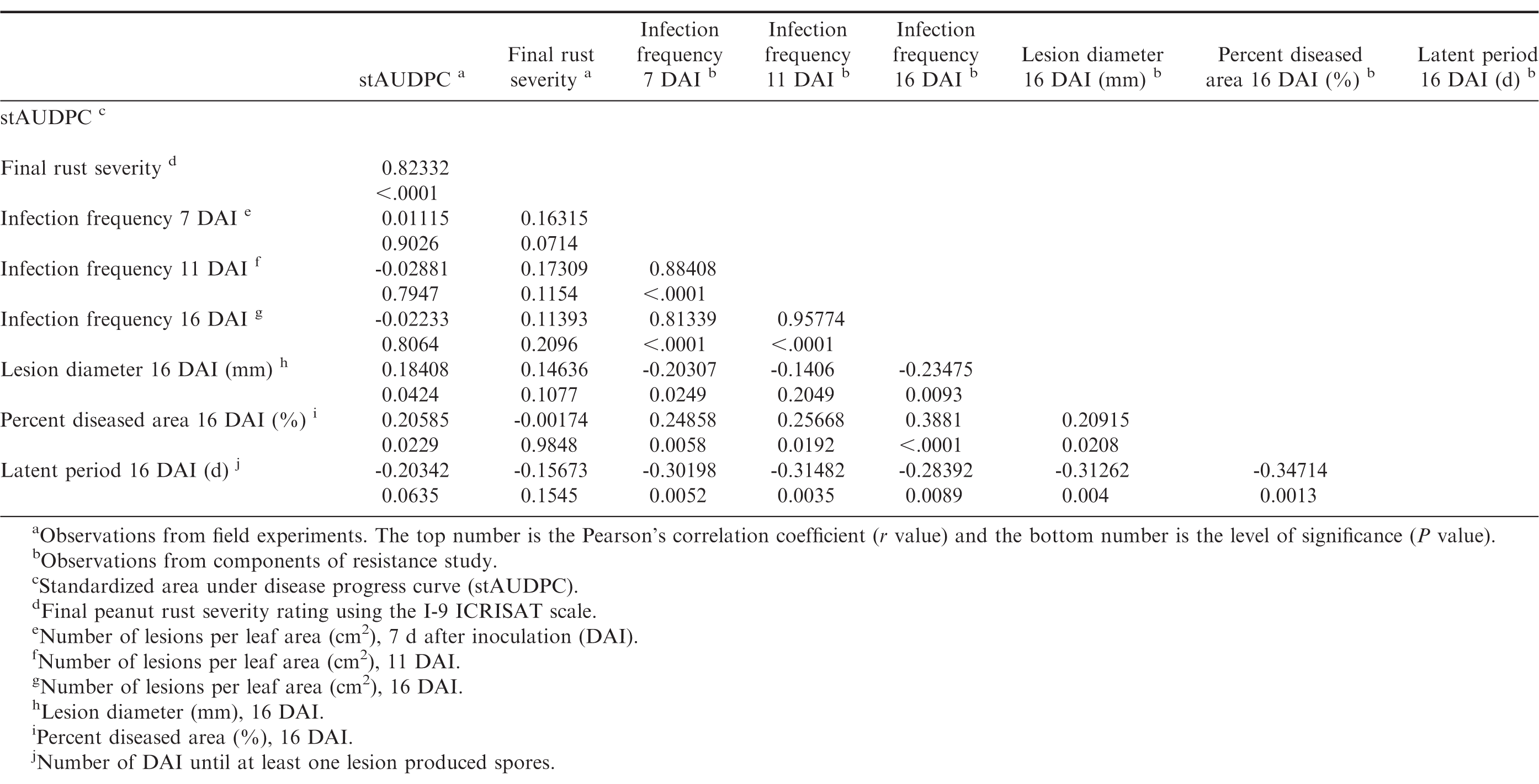
Infection frequencies at 7, 11, and 16 DAI were highly positively correlated to each other (P ≤ 0.05, r > 0.8), and infection frequencies at 7 and 16 DAI were also positively correlated with percent diseased area (P ≤ 0.05), and negatively correlated with lesion diameter and latent period (P ≤ 0.05) (Table 10). The general trend observed was that genotypes with short latent periods had high infection frequencies and large percent diseased area, whereas genotypes with long latent periods had low infection frequencies and small percent diseased areas. However, infection frequencies were not correlated (P > 0.05) with field observations. Standardized AUDPC was positively correlated with lesion diameter (P = 0.04) and percent diseased area (P = 0.02), however the correlation coefficient was low (r < 0.21). There were no correlations between final disease severity and any of the components (P > 0.05). Correlations between stAUDPC and final severity were high (r > 0.8, P ≤ 0.05). Although there were correlations between field resistance and the components of resistance as measured in controlled environments, the correlation coefficients were low. There are several factors that may explain the low or lack of correlation. One explanation may be that there are environmental factors in the field situations related to the growth habit of a genotype that may affect susceptibility of the genotypes to rust, and enhance or reduce disease development. For example, Tifrust-13 has a vine-like growth habit, which may enable it to escape spore deposition, and/or reduce the humidity necessary for successful peanut rust inoculations.
In conclusion, newly developed CRSP breeding lines with Bolivian genetic background, commonly grown peanut cultivars in Georgia, and parents of existing breeding populations were compared in this study for their response to peanut rust by determining the level of field resistance and components of resistance in growth chamber and greenhouse experiments. These studies resulted in the identification of several genotypes with multiple disease resistance in different environments and under high disease pressure. Furthermore, these results indicate sources of rust resistance in the CRSP breeding lines, including several genotypes that could be used as parents in peanut germplasm enhancement programs. Since several peanut mapping populations are being evaluated for multiple traits, including disease resistance (personal communication A. K. Culbreath), information on the response of the parents to peanut rust would be beneficial, because these populations may then be screened for peanut rust resistance as well to enable the identification of QTLs for rust resistance. Together with the availability of microsatellite markers, this could enable marker-assisted breeding for peanut rust resistance. All eight parental lines that were evaluated in this study were among the genotypes with high infection frequencies at 16 DAI in the growth chamber studies. These studies also indicate that latent period, percent diseased area, and lesion diameter may be used as indicators for rust resistance in growth chamber studies.
Acknowledgements
This research was supported, in part, by the National Peanut Board, USAID Peanut Collaborative Research and Support Program (Project UF150). We thank Justin McKinney, Mark Gomillion, Michael Heath, Ronald Hooks, Matthew Wiggins, Samuel Holbrook, Patricia Hilton, Miranda Goodman, Cassidy Reeh, Rachel Woodard, Sarah Rooks, Stephen Mullis, Dr. Sergio Morichetti, Dr. Graeme Wright, Jeff Tatnell and Alyssa Cho for assistance in field and growth chamber studies. We also thank Dr. Peggy Ozias-Akins and Dr. Corley Holbrook, for providing peanut germplasm.
Literature Cited
Bellgard, S., and K.C Ham 2004 Common diseases of peanuts in the top end of the NT In: Agnote Agdex No: 141/633 A. A. N. 141/633.
Black, M.C., and D.H Smith 1987 Spotted wilt and rust reactions in south Texas among selected peanut genotypes Proc. Amer. Peanut Res. Educ. Soc 19: 31 (abstr.).
K.R Bromfield, (1971). Peanut rust: A review of literature. Proc Amer. Peanut Res. Ed. Soc 3: (1): 111- 121.
Z.A., Chiteka, T.A Kucharek, D.A., Knauft, D.W Gorbet, and F.M Shokes (1988). Components of resistance to late leafspot in peanut. I. Levels and variability-implications for selection. Peanut Sci 15: (1): 25- 30.
Cook, M 1980 Host-parasite relations in uredial infections of peanut by Puccinia arachidis Phytopathology 70 (8): 822- 826 doi:10.1094/Phyto-70-822.
J., Damicone, and H.A Melouk (2010). Foliar diseases of peanut. Oklahoma State University, Division of Agricultural Sciences and Natural Resources, Oklahoma Cooperative Extension Service EPP-7655. .
Hammons, R.O., W.D Branch, K.R Bromfield, P Subrahmanyam, V.R Rao, S.N Nigam, R.W Gibbons, and E Goldin 1982 a Registration of Tifrust-13 peanut germplasm Crop Sci 22: 697.
Hammons, R.O., S.N Nigam, R.W Gibbons, P Subrahmanyam, and V.R Rao 1982 b Registration of eight peanut germplasm lines resistant to rust Crop Sci 22 (2): 452- 453.
Mehan, V.K., P.M Reddy, K.V Rao, and D McDonald 1994 Components of rust resistance in peanut genotypes Phytopath 84 (12): 1421- 1426 doi:10.1094/Phyto-84-1421.
Melouk, H.A., and F.M Shokes 1995 Peanut rust pp 72 In: Peanut Health Management H.A Meloukand F.M Shokes, eds Amer. Phytopathological Soc. Press, St. Paul, MN.
Power, I.L., A.K Culbreath, and B.L Tillman 2013 Characterization of resistance of peanut to Puccinia arachidis Plant Health Progress doi:10.1094/PHP-2013-1125-02-RS
H.W Rankin, (1953). Peanut rust in Georgia. Plant Disease Reporter 37: 528.
G., Shaner, and R.E Finney (1977). The effect of nitrogen fertilization on the expression of slow-mildewing resistance in Knox wheat. Phytopathology 67: 1051- 1056.
N.B., Smith, and R.H Littrell (1980). Management of peanut foliar diseases with fungicides. Plant Dis 64: 356- 360.
Subrahmanyam, P., R.O Hammons, S.N Nigam, D McDonald, R.W Gibbons, M.Y Fan, and W.L Yeh 1983 c International cooperative screening for resistance of peanut to rust and late leaf spot Plant Dis 67: 10: 1108- 1111 doi:10.1094/pd-67-1108.
P., Subrahmanyam, D McDonald, R.W Gibbons, and L.J Reddy (1985). Peanut rust: A major threat to peanut production in the semiarid tropics. Plant Dis 69 ((9)): 813- 819.
Subrahmanyam, P., D McDonald, and P.V Subba Rao 1983 b Influence of host genotype on uredospore production and germinability in Puccinia arachidis Phytopathology (73): 726- 729.
Subrahmanyam, P., D McDonald, F Waliyar, L.J Reddy, S.N Nigam, R.W Gibbons, V.R Rao, A.K Singh, S Pande, P.M Reddy, and P.V Subba Rao 1995 Screening methods and sources of resistance to rust and late leaf spot of groundnut pp 1- 14 In: Information Bulletin no. 47. Intl. Crops Res Institute for the Semi-Arid Tropics, Andhra Pradesh, India.
Subrahmanyam, P., V.R Rao, D McDonald, J.P Moss, and R.W Gibbons 1989 Origins of resistances to rust and late leaf spot in peanut (Arachis hypogaea, Fabaceae) Econ. Bot 43 (4): 444- 455 doi:10.1007/bf02935917.
Subrahmanyam, P., P.V Subba Rao, R.W Gibbons, and D McDonald 1983 c Components of resistance to Puccinia arachidis in peanuts Phytopath 73 (2): 253- 256.
Subrahmanyam, P 1997 Rust pp 31- 33 In Compendium of Peanut Diseases, 2nd ed N Kokalis-Burelle, D.M Porter, R Rodriguez-Kabana, D.H Smith, and P Subrahmanyam (eds.) Amer. Phytopath. Soc., St. Paul, MN.
USDA- NASS 2015 Georgia County Estimates Peanuts 2013- 2014 https://www.nass.usda.gov/Statistics_by_State/Georgia/Publications/County_Estimates/2015/GAPeanuts15.pdf.
E. P., Van Arsdel, and A.L Harrison (1972). Possible origin of peanut rust epidemics in Texas. Phytopathology 62 ((7)): 794.
Williams-Woodward, J 2013 Georgia Plant Disease Loss Estimates 2011 The University of Georgia Cooperative Extension Service, Ann Publ. 102-4 Athens, GA.
Wynne, J. C., M.K Beute, and S.N Nigam 1991 Breeding for disease resistance in peanut (Arachis hypogaea L) Annu. Rev. of Phytopath 29: 279- 303 doi:10.1146/annurev.py.29.090191.001431.
Notes
- Department of Plant Pathology, University of Georgia, Tifton 31793-5766 [^]
- Current address of first author: Center for Agricultural Research in Suriname (CELOS), Prof. Dr. Ir. J. Ruinardlaan, Postbus 1914, Paramaribo, Suriname [^]
- Department of Agronomy, University of Florida, Marianna 32446-7906. [^] *Corresponding author e-mail: imanapower@yahoo.com
Author Affiliations