Introduction
Root-knot nematode (RKN) (Meloidogyne arenaria (Neal) Chitwood) is a soil-borne pest that severely reduces peanut production in infested fields. Only moderate resistance to nematodes has been found in peanut upon screening of more than 1000 plant introductions (Holbrook and Noe 1992), whereas strong resistance has been identified in several wild diploid species such as A. glabrata, A. batizocoi , and A. cardenasii (Baltensperger et al., 1986; Holbrook and Noe, 1990; Nelson et al., 1990). Introgression of high levels of nematode resistance from wild species to cultivated peanut was achieved through two separate pathways. Simpson et al. (1993) produced a highly fertile nematode resistant amphidiploid, TxAG-6, through a three-way interspecific cross ( A. batizocoi Krapov & Gregory) x ( A. cardenasii x A. diogoi Hoehne). TxAG-6 was backcrossed with recurrent cultivated parent Florunner (Norden et al., 1969) to result in the release of nematode resistant cultivars COAN (Simpson and Starr, 2001) and NemaTAM (Simpson et al., 2003). Integration of nematode and TSWV resistance led to the release of Tifguard (Holbrook et al., 2008), a cultivar suitable for the southeastern peanut growing region where TSWV imposes a major threat to peanut production. The second introgression pathway was followed by Stalker et al. (2002) in which A. hypogaea (PI261942) was crossed with A. cardenasii and hexaploid F1 progeny were produced by colchicine treatment. Tetraploid progenies carrying resistance to nematodes were identified upon backcrossing and spontaneous chromosome reduction. Several molecular markers linked to nematode resistance were developed from these breeding materials (Garcia et al., 1996), yet they were not widely used in peanut breeding programs. The main source of nematode resistance introgressed into cultivated peanut was derived from A. cardenasii via TxAG-6 (Burow et al., 1996; Choi et al., 1999) and manifested as complete inhibition of invading juvenile development (Starr et al., 1995).
Thirteen polymorphic markers defined the introgressed region from TxAG-6 when mapped in two populations, Gregory x Tifguard and NemaTAM x GP NC WS14 (Nagy et al., 2010). Recombination was severely repressed with zero and 3.4 cM map distance for Gregory x Tifguard and NemaTAM x GP NC WS14, respectively, although the syntenic region spanned 30.1 cM in an intraspecific A. duranensis genetic map. The lack of recombination prevented further fine mapping of the resistance trait within this region. Nevertheless, molecular markers within the introgressed region were deployed in breeding programs such as a dominant CAPS (cleaved amplified polymorphic sequence) marker S197 (Chu et al., 2007) and a co-dominant SSR (simple sequence repeat) marker GM565 (Chu et al., 2011). Recently, marker and phenotype dissociation was reported in a breeding program using COAN as the donor for nematode resistance (Branch et al., 2014). In order to develop new markers tightly linked to the resistance trait, a recombinant inbred line (RIL) population of Gregory x Tifguard was genotyped with 132 polymorphic SSR (simple sequence repeat) and SNP (single nucleotide polymorphism) markers. A rare recombinant carrying part of the introgressed region while retaining strong nematode resistance was discovered enabling deployment of more tightly linked markers in breeding programs to improve selection for nematode resistance.
Materials and Methods
Plant Materials
Seventy-eight RILs from Gregory x Tifguard F7:9 were used for genotyping with polymorphic SSR or SNP markers. Gregory, the susceptible female parent, is a virginia-type cultivar developed from a cross between two elite parents (Isleib et al., 1999). Tifguard, the resistant male parent, carries a large alien introgressed region on chromosome A09 conferring near-immunity to root-knot nematode (Holbrook et al., 2008; Nagy et al., 2010). The original source of the introgression was from an A-genome wild diploid species A. cardenasii (Simpson et al., 1993; Garcia et al., 1996).
Genotyping
DNA was extracted using the DNeasy plant mini-kit (Qiagen, Valencia, CA). PCR amplifications for SSR markers were carried out in a GeneAmp® PCR System 9700 (Applied Biosystems, Foster City, CA) using a touchdown program, starting with 95 C for 5 min, followed by 6 cycles of 95 C for 30 s, 64 C (dropping 1 C /cycle) for 30 sec and 72 C for 30 sec, followed by 30 cycles of 95 C for 30 sec, 58 C for 30 sec and 72 C for 30 sec; final extension was performed at 72 C for 7 min. Forward primers were labeled with FAM, HEX, or TAMRA fluorophores (Supplemental table 1). PCR products were run on 1.5% agarose gels to check for amplification and contamination. These PCR products were then diluted 40 times using sterile water. The final mixture for fragment analysis in each well of a 96-well semi-skirted plate was constituted by 1 μl of the diluted PCR product, 9 μl of Hi-Di formamide (Applied Biosystems, Foster City, CA) and ∼ 0.20 μl of the ROX™ dye-labeled size standard. SSR markers were genotyped on an ABI3730XL Capillary DNA Sequencer (Applied Biosystems, Foster City, CA). Genotypes were called using the Gene Mapper 4.0 software (Applied Biosystems, Foster City, CA) and manual correction.
For SNP markers, KASP assays (KBioscience Ltd., Hoddesdon, UK) were developed for a set of 25 markers that were polymorphic between parents (Supplemental table 2). In this study, ‘GKAM' (Groundnut KASP Assay Markers) nomenclature (Khera et al., 2013) was used for the SNPs from the tetraploid array and ‘AdSNP' for the SNPs from the diploid array (Nagy et al., 2012).
Thermocycling and endpoint genotyping for the KASP assays were performed on a Roche LC480 (Roche Applied Science Indianapolis, IN) using two oligos 5′-labeled with FAM and HEX fluorophores. Each 5 μl volume reaction contained 2.5 μl of KASP Genotyping Mix, 0.07 μl of primer assay mix, 1.63 μl of water and 0.8 μl of genomic DNA template (5 ng/μl). The following thermal cycling program was used: hot start or activation at 95 C for 15 min, followed by 9 cycles of 94 C for 20 sec and 61 C for 60 sec, the annealing temperature dropped at the rate of 0.6 C/cycle, followed by 27 cycles at 94 C for 10 sec and 55 C for 60 sec and 2 cycles of 94 C for 20 sec and 57 C for 60 sec. Pre- or post-melt cycles were at 30 C for 1 sec and cooling to 25 C during plate reading. If the signals did not separate sufficiently, three additional cycles of 94 C for 20 sec and 57 C for 60 sec were performed. Roche LC480 allowed for endpoint genotyping based on dual color hydrolysis probes (FAM and HEX) and automated scatterplot analysis.
Phenotyping for Nematode Resistance
Phenotyping for nematode resistance of RIL 46, 48 and parents was performed using a greenhouse screening method as described previously (Holbrook et al., 1983; Timper et al., 2003). Peanut plants were grown in steam-pasteurized loamy sand. Meloidogyne arenaria were propagated on eggplant ( Solanum melongena L.) plants in the greenhouse. Eggs were harvested from roots of eggplant plants and hatched in a mist chamber to obtain second-stage juveniles (J2). Two experiments were carried out one wk apart. Each consisted of 10 randomized blocks with all four genotypes in each block. Prior to germination, all seeds were genotyped with markers AdSNP124 and AdSNP92. High throughput DNA extraction from seeds was performed following a previously published protocol (Chu et al., 2011). To obtain seed tissue for extraction, a small piece of cotyledon tissue (approximately 2 to 3 mm in diameter, 0.5 mm in thickness) was excised from the distal end of the seed. One Tifguard seed appeared to be heterozygous for both markers and was excluded from the experiment; the remaining seeds were homozygous for their expected genotype profiles. Eleven days after germination, 3,600 J2/pot and 5,800 J2/pot were applied to each trial, respectively. Plants were harvested 70 days after inoculation. After roots were washed clean of soil, fresh weights of roots were taken. Nematode eggs were extracted in a 1% NaOCl solution for 4 minutes and collected on nested 150- and 25-μm-pore sieves. Nematode eggs were counted using a dissecting microscope. Gall ratings were performed based on the following scale: 0 = no gall, 1 = 1 to 2, 2 = 3 to 10, 3 = 11 to 30, 4 = 31 to 100, 5 = more than 100 galls per root system (Taylor and Sasser, 1978). Nematode egg numbers per gram of root were transformed by square root for statistical analysis. Gall ratings were analyzed without transformation. General linear model was used to determine whether there were interactions between peanut genotype and experimental trial. Differences (P<0.05) among peanut genotypes were determined by Fisher's LSD test using SAS Enterprise Guide software V6.1 (SAS Institute, Cary, NC).
Results and Discussion
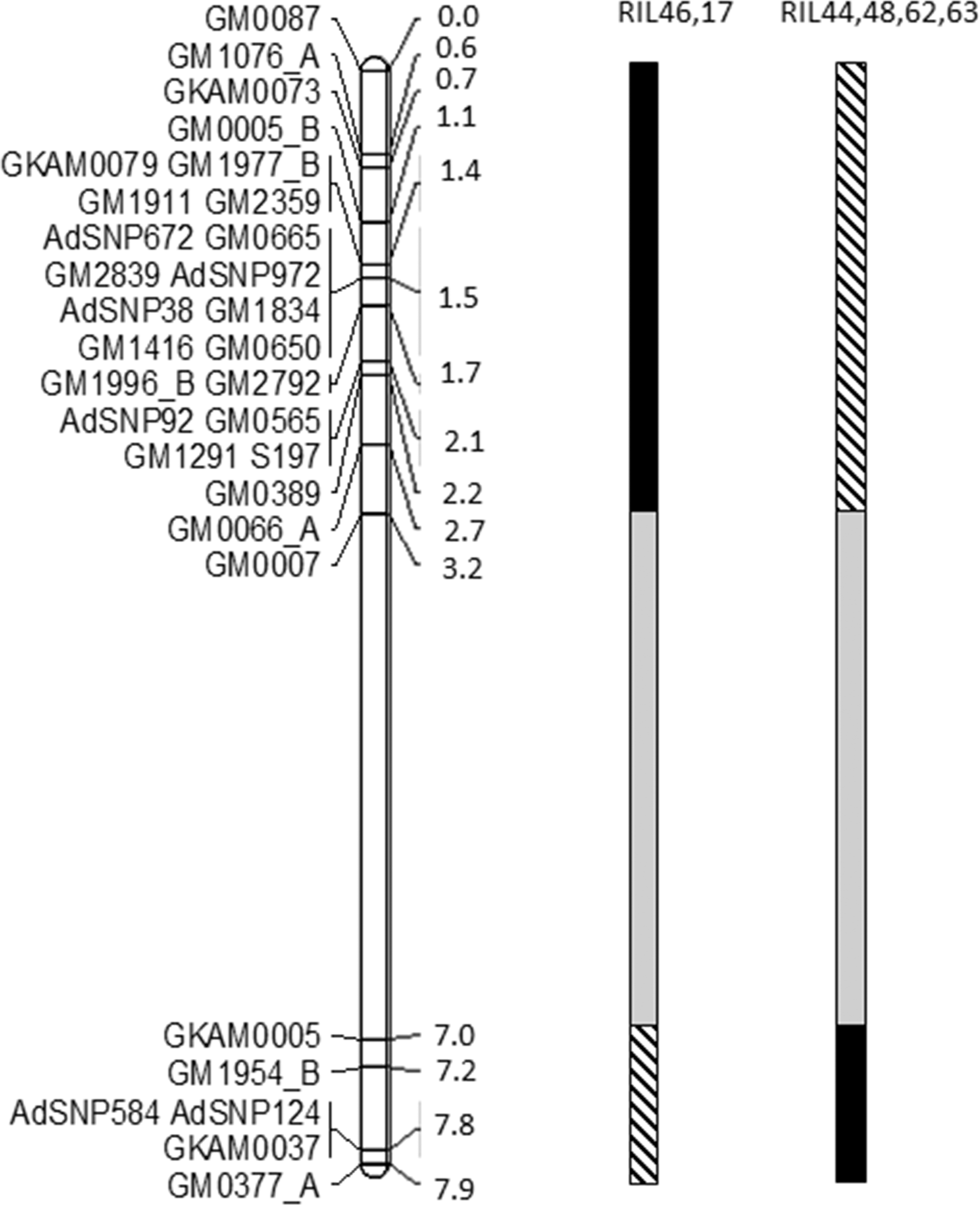
A total of 109 SSRs (24% of screened primer pairs) and 24 SNP markers were polymorphic between Gregory and Tifguard and mapped to 17 linkage groups, with 31 markers (23%) distributed on linkage group A09 at a map distance of 7.9 cM (Fig. 1). Marker coverage was biased for the introgressed region and too sparse across other linkage groups to generate a useful whole genome map. Previously, no recombination among markers in the introgressed region was observed by genotyping F4 RILs of this population with 13 markers (Nagy et al., 2010). Markers on linkage group A09 were clustered in two regions designated as the first (0 to 3.2 cM) and second introgressed regions (7 to 7.9 cM). The first introgressed region was heavily populated with markers and all six markers (GM665, GM650, GM565, S197, GM389 and GM66) that were common to both current and published maps (Nagy et al., 2010) were clustered within this region, although the order of one marker, GM66, was discrepant with the published NemaTAM x GP NC WS 14 map. Significant segregation distortion caused by excessive heterozygotes in the NemaTAM x GP NC WS 14 population has been reported previously (Nagy, et al., 2010). In the present study, the amount of heterozygosity was reduced by removing five individuals that had greater than 14% heterozygous loci. Four loci with greater than 40% heterozygous alleles and 7 loci with significant (P<0.001) linkage distortion were also excluded. The total percentage of heterozygotes in the data analyzed was reduced from 3.2% to 1%. The second introgressed region had five markers positioned between 7 and 8 cM and these markers were not mapped in previous publications. Increased marker density in this study allowed for detection of recombination events in the population and increased genetic map distance.
To test nematode resistance levels conferred by the two introgressed regions on linkage group A09, recombinant lines between these two regions were identified by revisiting the genotyping data. Two RIL lines (17 and 46) retained alien alleles in the second introgressed region and exhibited a haplotype from susceptible parent Gregory in the first introgressed region (Fig. 1). Four RIL lines (44, 48, 62 and 63) possessed alien alleles in the first introgressed region and a Gregory haplotype in the second introgressed region. Due to the lack of markers between 3.2 cM and 7.0 cM, the source of allele transmission within this region is unknown. Only 6 out of 78 RILs were recombinants within the introgression, and recombination during marker-assisted selection has only recently been reported (Chu et al., 2011; Branch et al., 2014).
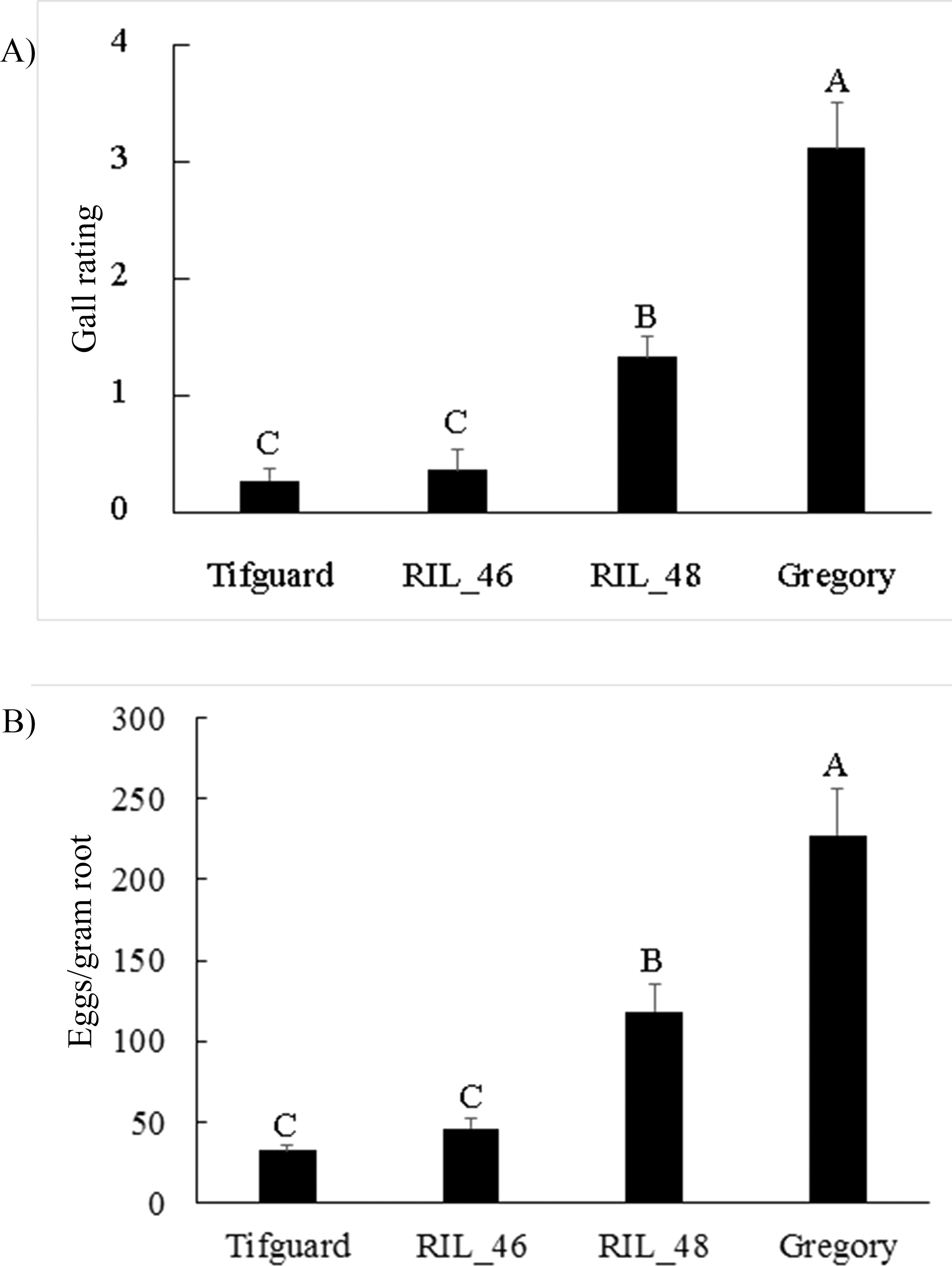
To further analyze the effect of the recombination, two inoculation trials, with 10 replicates per trial, were performed. Although a standard greenhouse inoculation test usually consists of 3 to 5 replicates (Holbrook and Noe, 1992), high variation among nematode phenotyping data has been reported in previous studies (Starr, et al., 1995, Timper, et al., 2000). Increasing the number of replications stabilizes variation. Quantitative data on egg number was reported to better indicate nematode resistance than egg mass index (Hussey and Janssen, 2002); therefore, egg counts were taken in the current study in addition to gall rating. Due to seed availability, phenotyping was performed only with RILs 46, 48, and parental lines, Tifguard and Gregory. Since there was no significant interaction between trial and plant genotype, data from the two trials were pooled for analysis. RIL 46 demonstrated resistance to nematodes as strong as the resistant parent, Tifguard, and was significantly more resistant than RIL 48 based on both egg count and gall rating (Fig. 2). RIL 48 showed an intermediate level of resistance to nematodes and its egg counts and gall rating were significantly lower than the susceptible parent, Gregory. Therefore, the first introgressed region has genes conferring moderate resistance to nematodes as detected in RIL 48. The second introgressed region harbors major nematode resistance gene(s) as demonstrated by the high level of resistance in RIL 46. Previously, two dominant resistance genes conditioning galling and egg production were reported in a F2 population segregating for nematode resistance introgressed from A. cardenasii (Garcia, et al., 1996). Phenotyping data from our recombinant inbred lines does not support the effect of two dominant genes controlling separate gall and egg phenotypes. Instead, since both galling and egg production were suppressed at a similar level within either the highly resistant RIL 46 line or the moderately resistant RIL 48 line, it appears that resistance gene(s) residing in either introgressed region show pleiotropic effects for galling and egg production. Identification of these RILs with shorter introgressed chromosomal segments will facilitate finer scale mapping and expression analysis of nematode resistance genes. Integrating RIL 46 into peanut breeding programs can reduce any linkage drag across the larger alien introgression while transferring high levels of nematode resistance.
Gall rating and number of eggs/gram root from nematode inoculation study. A) Gall rating; B) Eggs/gram root. Gall ratings are based on the following scale: 0 = no gall, 1 = 1 to 2, 2 = 3 to 10, 3 = 11 to 30, 4 = 31 to 100, 5 = more than 100 galls per root system. Eggs/gram root was transformed by square root. Different letters indicate significant differences at P<0.05 for mean separation by Fisher's LSD test.
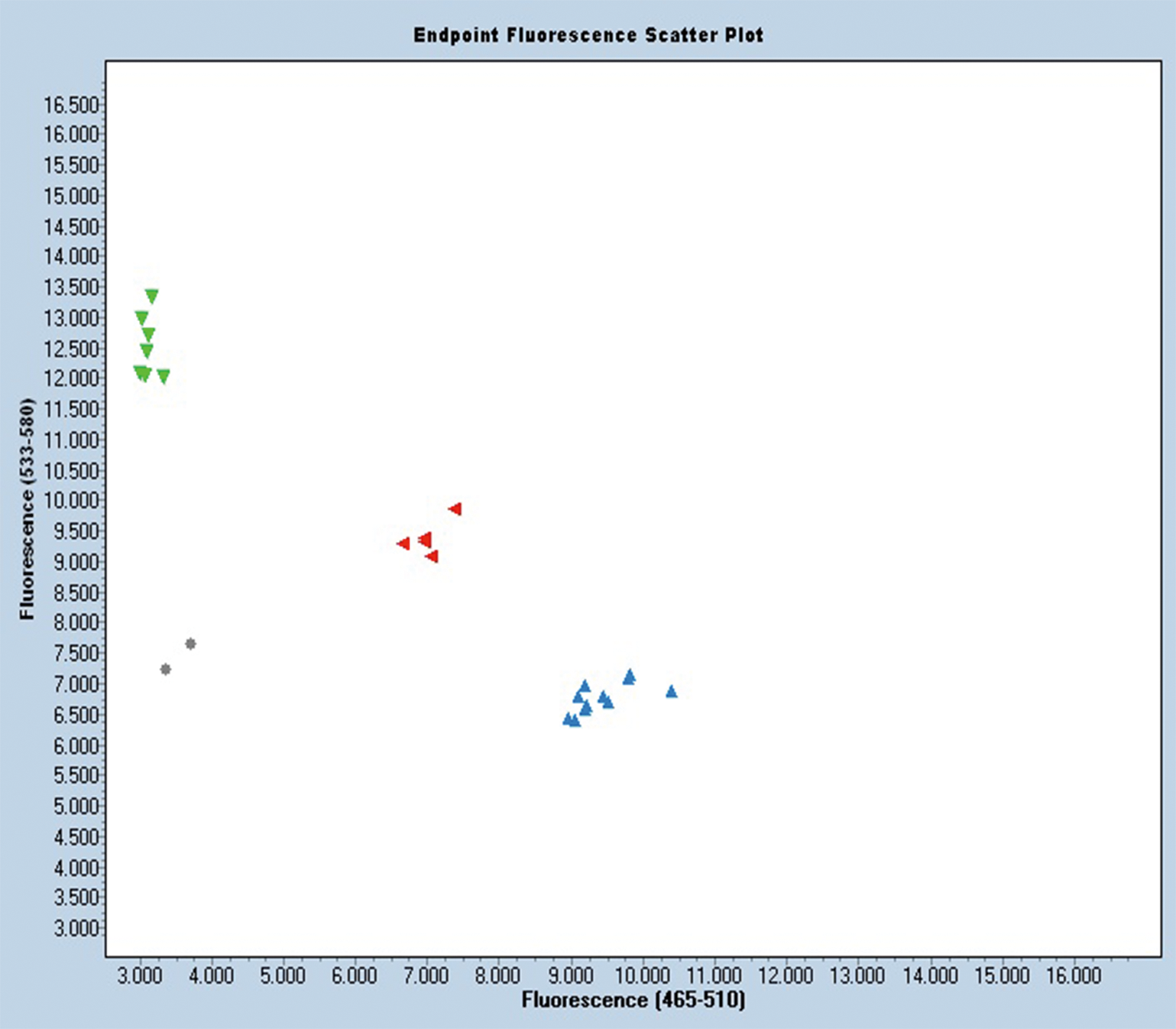
SNP marker AdSNP 584 detects a simple SNP, i.e., two alleles at a single locus in one of the two sub-genomes. It is located at the second introgressed region that has an “A” in Gregory and a “G” in Tifguard (Supplemental table 2). KASP analysis shows that signals from homozygotes and heterozygotes separate perfectly in the scatter plot with this marker (Figure 3). This new marker is linked tightly to the major resistance gene(s) and can be deployed in breeding programs to improve selection for nematode resistance. This finding also offers a potential explanation for the reported marker phenotype dissociation (Branch et al., 2014). Since the four resistant lines dissociated with markers are derivatives of COAN, they could be recombinant lines similar to RIL 46. Testing these lines with markers in the second introgressed region, including AdSNP 584, would clarify their genetic inheritance.
Acknowledgements
This work was supported by The Peanut Foundation, The National Peanut Board, Georgia Peanut Commission, and The Agriculture and Food Research Initiative competitive grant # 2010-85117-20550 of the USDA National Institute of Food and Agriculture.
Literature Cited
D. D., Baltensperger, R. A Dunn and G. M Prine (1986). Root-knot nematode resistance in Arachis glabrata . Peanut Sci 13 : 78 – 80 .
W. D., Branch, T. B Brenneman and G Hookstra (2014). Field test results versus marker assisted selection for root-knot nematode resistance in peanut. Peanut Sci 41 : 85 – 89 .
M. D., Burow, J. L Starr, A. H Paterson and C. E Simpson (1996). Identification of peanut ( Arachis hypogaea L.) RAPD markers diagnostic of root-knot nematode (Meloidogyne arenaria (Neal) Chitwood) resistance. Mol. Breed 2 : 369 – 379 .
K., Choi, A. H Paterson, C. E Simpson, J. L Starr, M. D Burow, G Church and G Burow (1999). Genetics and mechanism of resistance to Meloidogyne arenaria in peanut germplasm. J. Nematol 31 : 283 – 290 .
Y., Chu, C. C Holbrook, P Timper and P Ozias-Akins (2007). Development of a PCR-based molecular marker to select for nematode resistance in peanut. Crop Sci 47 : 841 – 847 .
Y., Chu, C. L Wu, C. C Holbrook, B. L Tillman, G Person and P Ozias-Akins (2011). Marker-assisted selection to pyramid nematode resistance and the high oleic trait in peanut. Plant Genome 4 : 110 – 117 .
G. M., Garcia, H. T Stalker, E Shroeder and G Kochert (1996). Identification of RAPD, SCAR, and RFLP markers tightly linked to nematode resistance genes introgressed from Arachis cardenasii into Arachis hypogaea . Genome 39 : 836 – 845 .
C. C., Holbrook, D. A Knauft and D. W Dickson (1983). A technique for screening peanut for resistance to Meloidogyne arenaria . Plant Dis 67 : 957 – 958 .
C. C Holbrook, and J. P Noe (1990). Resistance to Meloidogyne arenaria in Arachis spp. and the implications on development of resistant peanut cultivars. Peanut Sci 17 : 35 – 38 .
C. C Holbrook, and J. P Noe (1992). Resistance to the peanut root-knot nematode (Meloidogyne arenaria) in Arachis hypogaea . Peanut Sci 19 : 35 – 37 .
C. C., Holbrook, P Timper, A. K Culbreath and C. K Kvien (2008). Registration of ‘Tifguard’ peanut. J. Plant Reg 2 : 92 – 94 .
Hussey, R.S. and G.J. Janssen 2002 Root-knot nematodes: Meloidogyne species Engham , UK, CABI.
T. G., Isleib, P. W Rice, R. W Mozingo and H. E Pattee (1999). Registration of ‘Gregory’ peanut. Crop Sci 39 : 1526 .
Khera, P., H.D Upadhyaya, M. K Pandey, M Roorkiwal, M Sriswathi, P Janila, Y Guo, M.R McKain, E. D Nagy, S.J Knapp, J Leebens-Mack, J.A Conner, P Ozias-Akins, and R.K Varshney 2013 SNP-based genetic diversity in the reference set of peanut ( Arachis spp.) by developing and applying cost-effective KASPar genotyping assays Plant Genome 6: doi: 10.3835/plantgenome2013.06.0019 .
E. D., Nagy, Y Chu, Y. F Guo, S Khanal, S. X Tang, Y Li, W. B Dong, P Timper, C Taylor, P Ozias-Akins, C. C Holbrook, V Beilinson, N. C Nielsen, H. T Stalker and S. J Knapp (2010). Recombination is suppressed in an alien introgression in peanut harboring Rma, a dominant root-knot nematode resistance gene. Mol. Breed 26 : 357 – 370 .
E.D., Nagy, Y Guo, S Tang, J. E Bowers, R. A Okashah, C. A Taylor, D Zhang, S Khanal, A. F Heesacker and N Khalilian (2012). A high-density genetic map of Arachis duranensis , a diploid ancestor of cultivated peanut. BMC Genomics 13 : 469 .
S. C., Nelson, C. E Simpson and J. L Starr (1990). Expression of resistance to Meloidogyne arenaria in Arachis batizocoi and A. cardenasii . J. Nematol 22 : 242 – 244 .
A.J., Norden, R.W Lipscomb, and W.A Carver (1969). Registration of ‘Florunner' peanuts. Crop Sci 9 : 850 .
C. E., Simpson, M. D Burow, A. H Paterson, J. L Starr and G. T Church (2003). Registration of ‘NemaTAM’ peanut. Crop Sci 43 : 1561 – 1561 .
C. E., Simpson, S. C Nelson, J Starr, K. E Woodward and O. D Simth (1993). Registration of TxAG-6 and TxAG-7 peanut germplasm lines. Crop Sci 33 : 1418 .
C. E Simpson, and J. L Starr (2001). Registration of ‘COAN’ peanut. Crop Sci 41 : 918 – 918 .
H. T., Stalker, K. R Barker, B. B Shew and M. K Beute (2002). Registration of two root-knot nematode-resistant peanut germplasm lines. Crop Sci 42 : 312 – 313 .
J. L., Starr, T. A Lee, and C. E Simpson (1995). Resistance to Meloidogyne arenaria in advanced generation breeding lines of peanut. Peanut Sci 22 : 59 – 61 .
A. L Taylor, and J. N Sasser (1978). Biology, identification, and control of root-knot nematodes (Meloidogyne species). Raleigh: North Carolina State Univ. Graphics. , .
P., Timper, W. F Anderson and C. C Holbrook (2003). Reproduction of Meloidogyne spp. on resistant peanut genotypes from three breeding programs. J. Nematol 35 : 417 – 421 .
P., Timper, H. Q Xue and C. C Holbrook (2000). Expression of nematode resistance in plant introductions of Arachis hypogaea . Peanut Sci 27 : 78 – 82 .
Notes
- First, second, third and last authors: Research Professional, MS student, PhD student, and Professor, Department of Horticulture and Institute of Plant Breeding, Genetics & Genomics, University of Georgia Tifton Campus, Tifton, GA 31793; Fourth and fifth authors: Research Plant Pathologist and Research Geneticist, USDA-ARS, P.O. Box 748, Tifton, GA 31793. [^] *Corresponding author: P. Ozias-Akins
Author Affiliations
Email: pozias@uga.edu