Introduction
Reliable phenotyping is crucial for a successful crop improvement program. It plays a vital role throughout the plant breeding process, from identification of sources for the trait, to selection and advancement of lines, and evaluation of cultivars prior to release. An ideal phenotyping method is robust, repeatable, resource-efficient, objective, non-destructive, and reliable (Janila et al., 2013). Pod and seed characteristics affect economic returns for peanut producers. Refined genetic mapping of grade components would lead to discovery of molecular markers that could significantly expedite breeding cycles (Chu et al., 2011). To improve measurements of pod and seed characteristics, a group of germplasm accessions with high levels of phenotypic variation was included in this study. In addition to establishing the phenotypic methods, characterization of these genetic materials provides useful information for their selection as crossing parents. The traits evaluated include pod volume, pod density, percent kernel, and testa color.
For pod volume, the liquid or water displacement method has been widely used for measurement in peanut (Agrawal et al., 1973; Aydin, 2007; de Godoy and Norden, 1981; Rucker et al., 1994). Based on the Archimedes’ principle, the volume of water displaced by a completely immersed object equals the volume of the object. However, high surface tension of water and absorption of water by peanut pod hulls may compromise the accuracy of volume measurement using this method. Agrawal et al. (1973) noted that water absorbed by each peanut pod ranged from 0.05 to 0.1 g that amounted to nearly 6 percent error in pod volume measurement of their samples. Aydin (2007) used toluene (C7H8) instead of water, to circumvent these issues.
The pod density of a peanut is defined as the ratio of pod weight to pod volume and also is known as true density (Aydin, 2007) or solid density (Akcali et al., 2006). According to Rucker et al. (1994), pod density of field cured pods increased with an increase in kernel maturity. Therefore, Rucker et.al (1994) also developed a technique to sort shell-stock peanuts into maturity classes based on pod density fractions. Peanut lots with higher percentages of mature kernels are desirable due to their favorable nutritional and flavor qualities and lower tendency for aflatoxin contamination (Ghate et al., 1993; Rucker et al., 1994).
The shelling percentage or percent kernel of a peanut sample is defined as the ratio between seed and pod weights. Knauft and Gorbet (1993) tested various market grade characteristics for consistency on a set of genotypes across several locations and environments during a 5 year period. Percent kernel and 100 seed-weight showed maximum stability across environments, such that even limited data on these characteristics could be used for making selections in a breeding program.
There are five basic solid testa (seed coat) colors for peanut including white, tan, pink, red, and purple (Branch, 2011). Among the US market types, pink is the most acceptable testa color, followed by tan, then red color (valencia type). Even after more than 100 years since Van der Stok (1910) first began testa color genetic studies in cultivated peanut, the inheritance of this trait is not completely understood (Branch, 2011). Therefore, it is important to evaluate testa color to select against the off-types early in the breeding process (Holbrook and Branch, 1989), as well as conduct studies to understand the inheritance of this trait. Testa color phenotyping involves manual sorting of kernels into various color classes. This method is highly subjective, and inconsistent results may arise due to lighting conditions, fatigue of the observer, etc. Hue, as one of the main attributes of color, was defined as an “attribute of a visual perception according to which an area appears to be similar to one of the colours: red, yellow, green, and blue, or to a combination of adjacent pairs of these colours considered in a closed ring” (CIE, 2014). Other properties of color include colorfulness, chroma, saturation, lightness, and brightness. Hue angle can be computed by converting RGB color into HSL (hue-saturation-lightness) or HSV (hue-saturation-value) color space.
Digital imaging and analysis can help to overcome the inherent issues of previously described methods to measure pod and seed characteristics. Image analysis is usually objective, reliable, robust, and resource-efficient. Over the past two decades, machine-vision based methods have been used in several studies in peanut phenotyping, including kernel size and kernel damage (Dowell, 1992), peanut maturity (Ghate et al., 1993; Colvin et al., 2014), pod color (Boldor et al., 2002), and projected pod area (Aydin, 2007), to name a few. Ghate et al. (1993) developed a machine-vision maturity classification method based on detection of surface texture differences among kernels. The gray-level histogram characteristics and gradient transformation were used on kernel images to distinguish among samples (Ghate et al., 1993). Color-based methods to assess pod maturity relied on computer-aided mesocarp color classification and a digital imaging model (DIM) using scanned images of the mesocarp layer (Bolder et al., 2002; Colvin et al., 2014).
The objectives of the present study were to: 1) develop an image acquisition and analysis method that can measure projected pod area and testa color; 2) build a regression model to predict peanut pod volume, by using projected pod area data, and 3) report pod density, percent kernel, and testa color data. The focus of this research was to use digital imaging and analysis for high-resolution phenotyping, and to provide alternative methods for pod volume and testa color measurements.
Materials and Methods
Plant Material
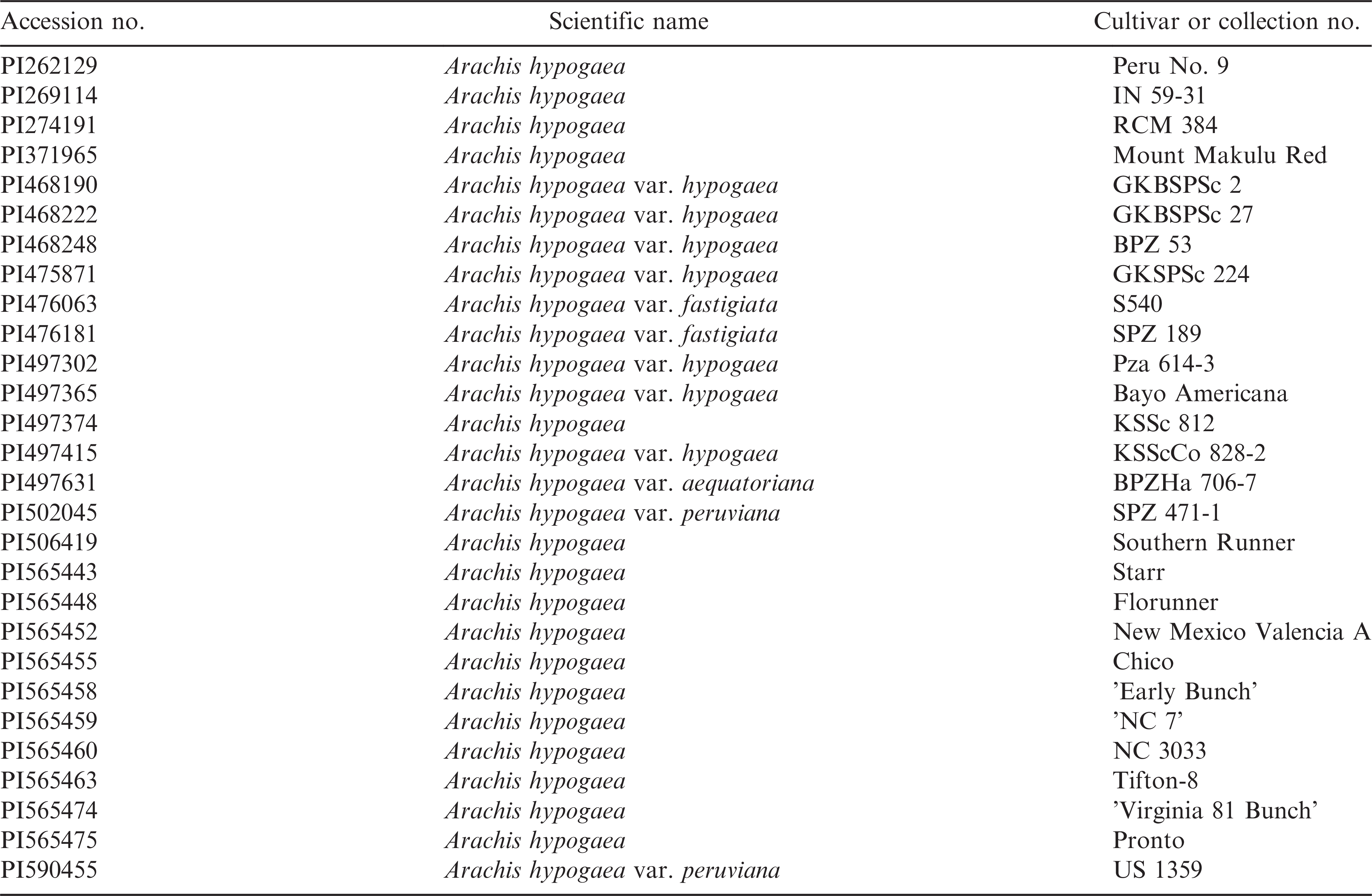
Fruits from 28 Arachis germplasm accessions (Table 1) were used in this high-resolution phenotyping study. These genotypes were grown in 1.8 m x 3.0 m field plots at the Gibbs Farm in Tift County, GA. Genotypes were sorted by maturity and planted on 19 April 2012. Standard cultural practices including irrigation and fungicide sprays were used as needed throughout the growing season. The early maturing genotypes were dug and picked on 16 August 2012, and the later maturing genotypes were dug and harvested on 28 August 2012. Pods were bulk harvested from the plots of each accession, dried to approximately 10% moisture, and stored at room temperature for approximately two months. A representative sample of 10 pods per accession was selected for phenotyping pod and kernel characteristics. Multi-kernel pod samples were selected to represent the maximum kernel production potential of each accession; this consisted of two-kernel pods for most, and three or four-kernel pods for eight of the 28 accessions where these were the predominant pod type. Each pod was carefully shelled by vertically cracking the hull into equal halves and the internal pericarp color (IPC) was used to determine pod maturity (Gilman and Smith, 1977). Immature and/or diseased pods were excluded from the samples to reduce environmental effects. For each accession, individual pods and their kernels were weighed before placing them in labelled coin envelopes for storage prior to image acquisition.
Image Acquisition and Processing
Ten half hulls per genotype collected using the procedure described above were included in one image as 10 replicates for that genotype. A simple image acquisition system consisting of a 12.3 MP digital SLR (single-lens reflex) camera (Nikon D5000) with an 18 to 55 mm lens attachment, a camera mount, and two 15W soft white incandescent bulbs (General Electric) were used to take the images. The camera setup was placed in a laminar flow hood and covered with a black cloth to eliminate background light. A piece of black plush cloth was placed under the samples, to enhance object/background contrast, which enabled easy image segmentation. The camera was vertically mounted at a fixed distance (30 cm) from the samples and the images were captured in manual mode at a fixed focal length of 24 mm, ISO of 400, shutter speed of 1/50 s, and F-stop of F4.5. The images were taken at a resolution of 2144 by1424 pixels and saved in .jpeg file format. The scale of image resolution was determined by a coin of known diameter, a US quarter. For standardization, a US quarter was subjected to imaging and the image was processed to calculate the number of pixels occupied by the quarter (Np ). The diameter (d) of a US quarter is 24.26 mm and its one-face area equals 462.2441 mm2. Area per pixel (Ap ) in the captured image at the given camera settings was calculated (Eq.1).
Ap : Area per pixel
Np : Number of pixels in one-face quarter area
ImageJ, a public domain program (NIH, 2015), was used for image processing. A set of 10 color images was opened each time for processing. Under “File” menu, “Import” submenu was selected and “Image sequence” command was used to import images for batch processing. Interactive image thresholding was performed to achieve image segmentation in HSB (hue, saturation, and brightness) color space. Under “Image” menu, “Adjust” submenu was selected and “Threshold” command was used to segment images into regions of interest (ROI) and background. Under “Analyze” menu, “Analyze Particles” command was used to individually label and measure the number of pixels occupied by the projected area of each sample.
Pod Volume Estimation Model
Two pods per accession, a total of 56 pods representing 28 germplasm accessions, were selected to build a model for estimated pod volume (Ve). The model was based upon the statistically dependent relationship between half-hull projected area (Pa ) (cm2), measured by image processing, and actual pod volume (Va ) (cm3), calculated by water displacement.
To estimate pod volume based on water displacement, a 30 ml beaker filled with a solution of 0.1% Triton X-100 (Sigma-Aldrich, St. Louis, Mo) was placed in a 100 mm by 15 mm polystyrene petri dish (Fisher Scientific, Pittsburgh, PA) lined with one layer of qualitative filter paper (Fisher Scientific). Triton X-100 was used to reduce surface tension in order to maintain a constant level of water when the beaker was filled to capacity. The setup was placed on a scale to ensure accurate replacement of water for each measurement. To measure pod volume, a pod was submerged in the beaker full of water and the overflow was collected in the petri dish. The preweighed, highly absorbent paper lining the petri dish, soaked up the excess water. The wet paper pieces were reweighed and the gain in weight (g) was recorded as Va (cm3), assuming that the density of water was 1.0 g/cm3. After measuring each sample, the beaker was refilled carefully to the top using a Rainin P1000 pipette (Mettler Toledo, Oakland, CA) until the initial weight of the setup was achieved.
Projected Pod Area
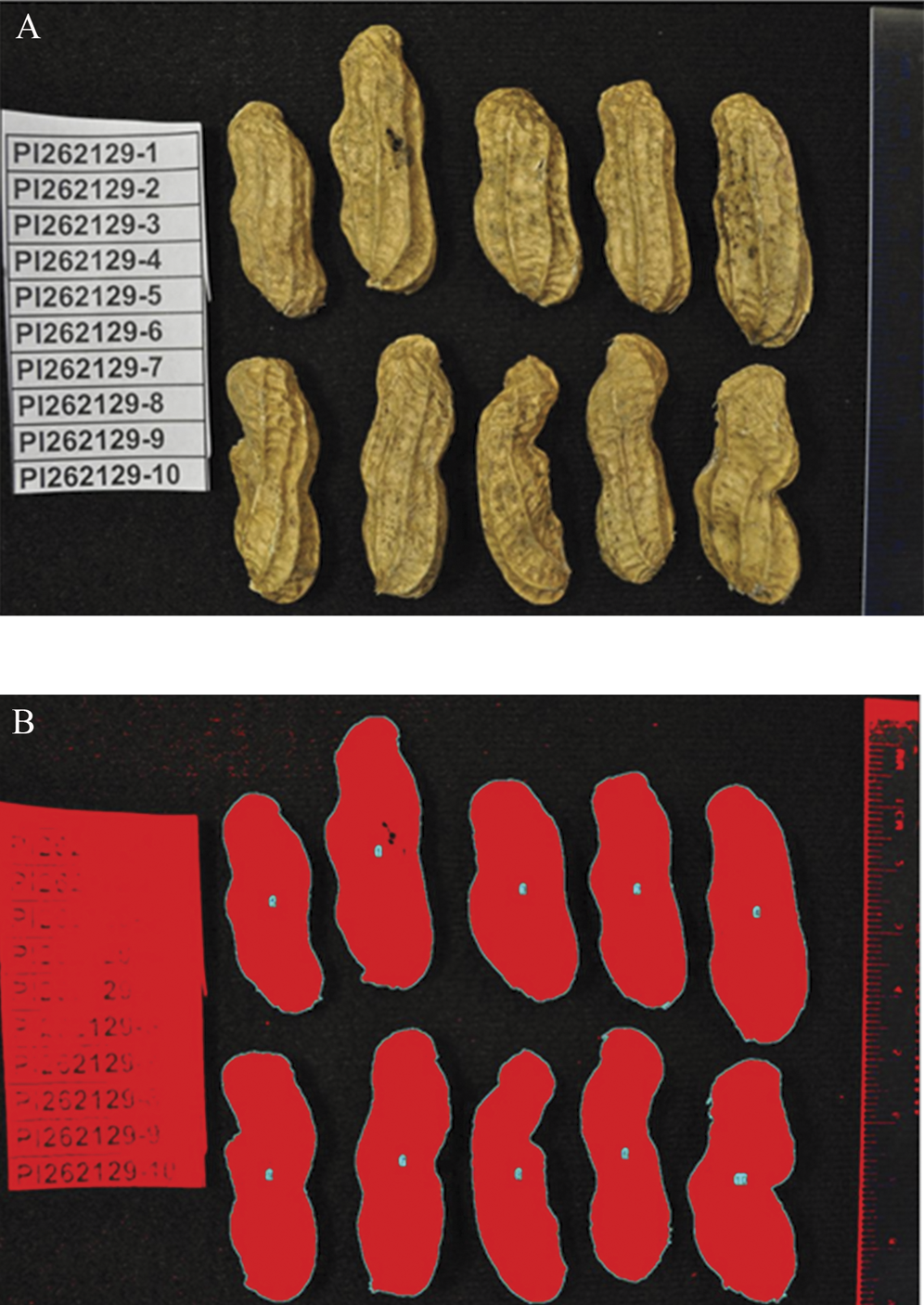
For each accession, one-half hull of every pod for 10 samples was included in a single image. The 10 half-hull samples were adjacently placed, in two rows of 5, on a plush black cloth (Figure 1). Pa for each half hull, in cm2, was calculated using the previously determined formula (Eq.1).
Pod Density
Pod density for each sample was calculated by dividing pod weight (Wp ) by Ve .
Percent Kernel
The percent kernel value for each pod was calculated by dividing the weight of kernels (Wk ) by Wp , and multiplying the result by 100 to express the observation as a percentage.
Testa Color
Testa color was quantified on accessions with solid testa colors by measuring hue angle values and visual grouping using the RHS color chart (The Royal Horticultural Society, London, SW1P2PE). To avoid complexity, five accessions with variegated testa were not included in this test. For each accession, a set of randomly chosen kernels was selected for imaging. All images were transferred from RGB (red, green and blue) color space to HSL (hue, saturation and lightness) color space. Five kernels were randomly selected in each image and the central areas of these kernels were defined as regions of interest (ROI) for mean hue angle extraction. Hue, one of the properties of color, was measured, to differentiate among various accessions. Hue angle (in degrees) of peanut testa was calculated to represent their respective color groups.
Statistical Analyses
Correlation and regression analyses were used to estimate the statistical relationship between Pa and Va . In order to predict Ve , a regression equation (Eq. 2) was fitted to the Pa and Va data.
Pod density, percent kernel, and hue angle (testa color) data were statistically analysed (PROC ANOVA) using SAS 9.2 software (SAS Institute, Cary, NC) and where significant, mean separations were obtained using Tukey’s HSD multiple comparison test at P ≤ 0.05. The relationship between pod density and percent kernel was determined statistically by Pearson’s correlation analysis.
Results and Discussion
A set of 28 diverse peanut germplasm accessions (Table 1) was phenotyped for pod and seed traits using image processing techniques. In the process of data acquisition from image J software, it was noted that the sequence of data output follows the relative position of an object along the Y-axis, i.e., an object placed higher produces data prior to any object placed in a lower position (Figure 1). Data output in this manner posed a challenge for the subsequent step of data sorting. We therefore recommend to arrange peanut pods such that their apical ends are aligned in descending order of their top co-ordinates from left to right. This alignment enables easy feature extraction during image processing.
Pod Volume Estimation Model
The relationship between Va and Pa data, calculated from 56 pods representing 28 peanut accessions, was statistically analysed to generate a simple linear regression model:
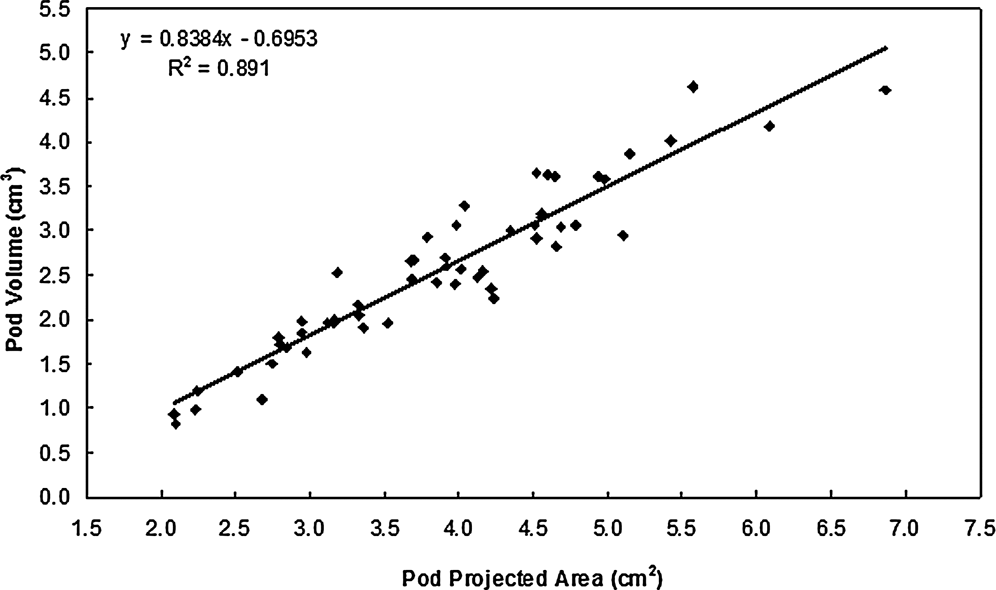
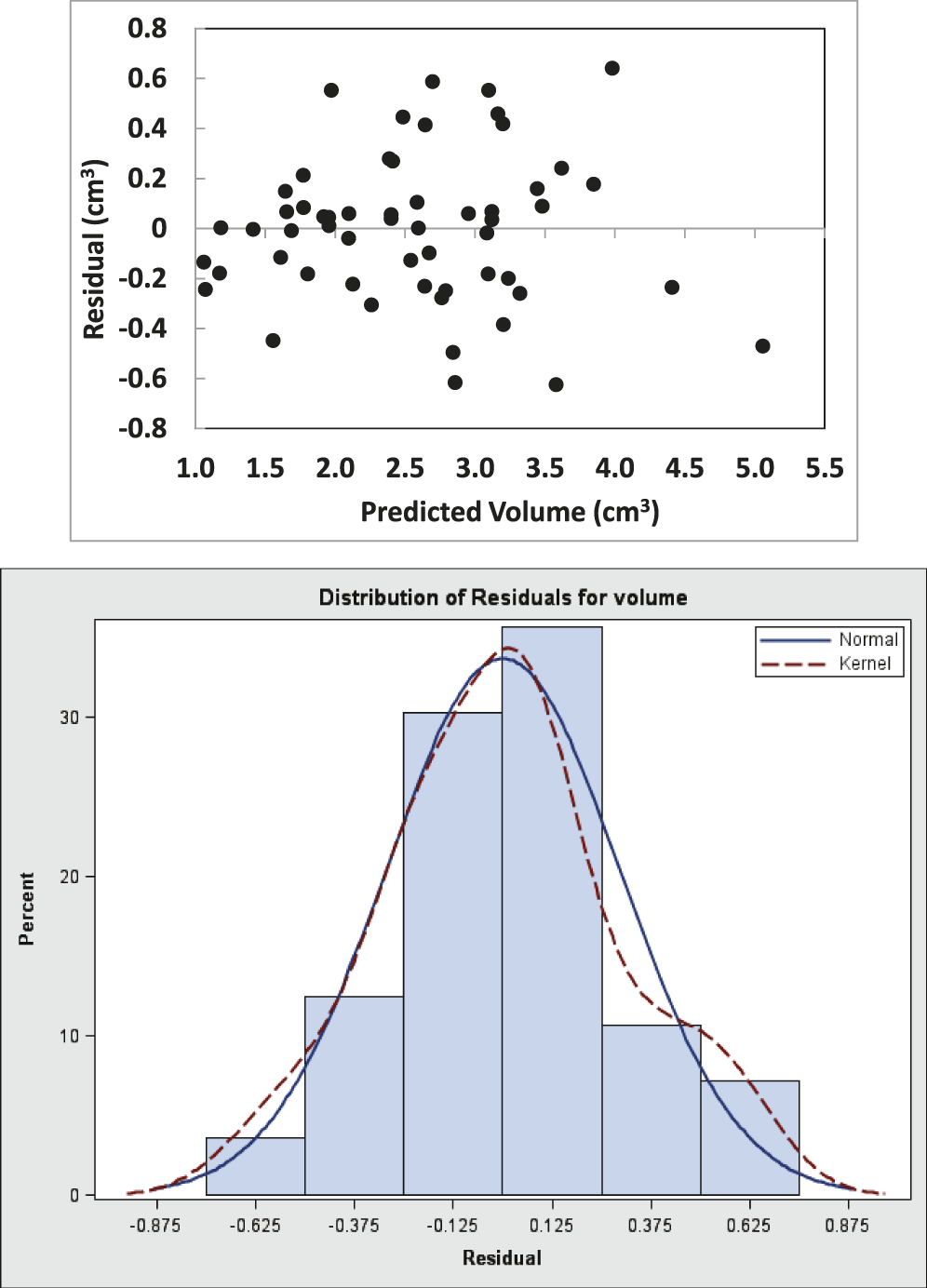
The parameters, Va and Pa , also indicated significant coefficient of correlation (r = 0.944, P = 0.05) between them. The linear regression model fit the data well (P<0.0001; R2 = 0.891) and provided significant evidence that Pa is a predictor of pod volume in the peanut germplasm set evaluated in this study. The fit plot (Figure 2) shows the positive slope of the fitted line. The random pattern of residuals in the scatter plot and normality distribution histogram (Figure 3) shows homogeneous error variances across the values of Pa , thus indicating that the simple linear regression model adequately describes the relationship between these data. From the model, it can be concluded that the estimated increase in average pod volume for a unit increase in projected pod area is 0.8384 cm3.
Pod Density, Percent Kernel and Pod Volume.—
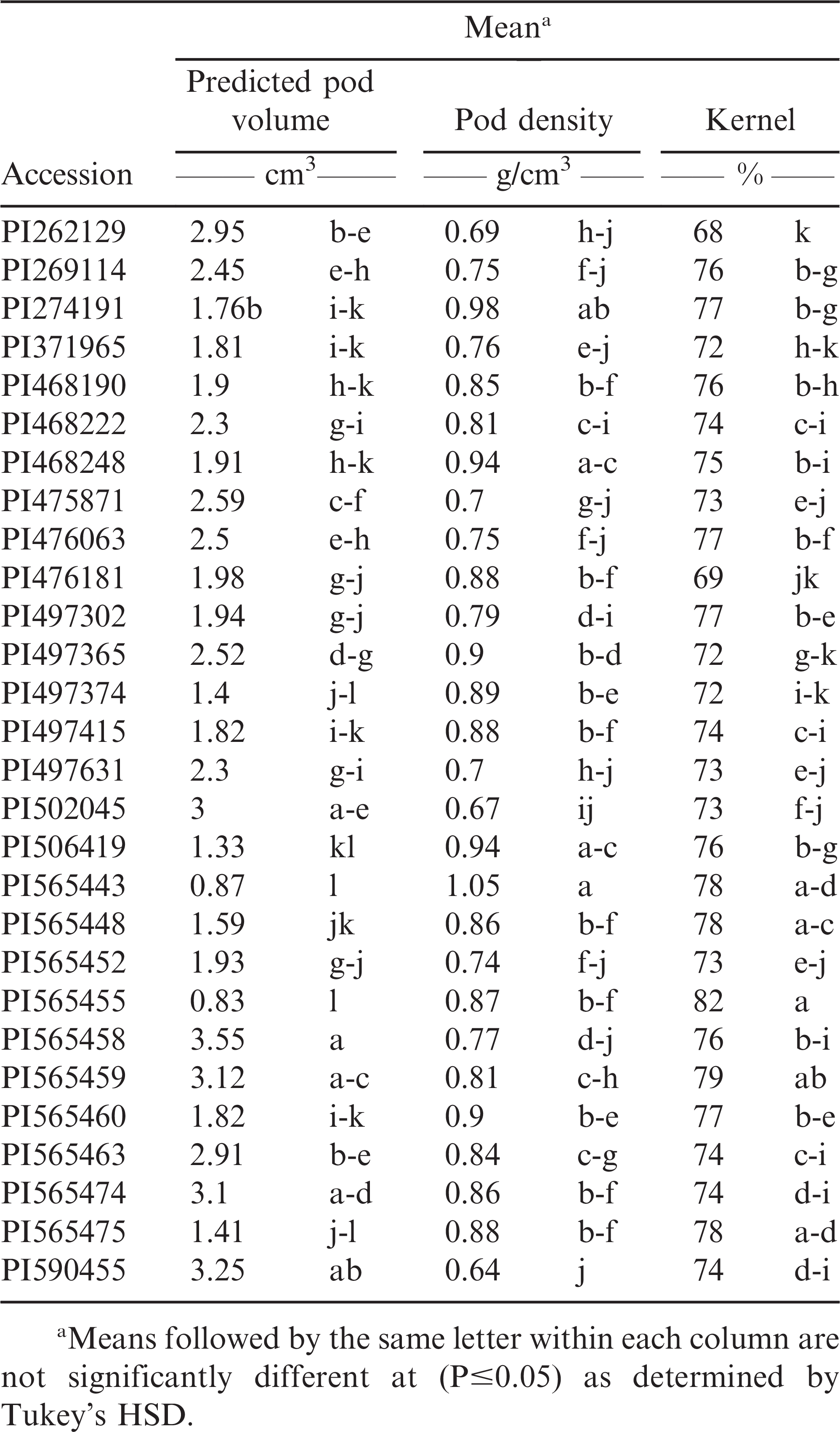
The genotypes differed (P<0.0001) for pod volume, pod density and percent kernel indicating that substantial variability existed for these traits in the test set. Tukey’s mean separation was calculated for each trait to further distinguish among accessions (Table 2). Mean predicted pod volume ranged from 0.83 cm3 (PI565455) to 3.55 cm3 (PI565458) suggesting large pod size variation among the accessions. The mean pod densities among 28 accessions were 0.64 (±0.0087) g/cm3 (PI590455) to 1.05 (±0.0163) g/cm3 (PI565443) (Table 2). Interestingly, both the Arachis hypogaea var. peruviana accessions (PI590455 and PI505045) had four-kernel pods, and the lowest pod density among the 28 entries. Whereas the samples with the highest density belonged to an A. hypogaea accession (PI565443) with two-kernel pods. However, there was insufficient data to suggest a correlation between pod density and number of kernels/pod. The mean portion of kernel among 28 accessions was 68% (±0.75) (PI262129) to 82% (±0.15) (PI565455) (Table 2).
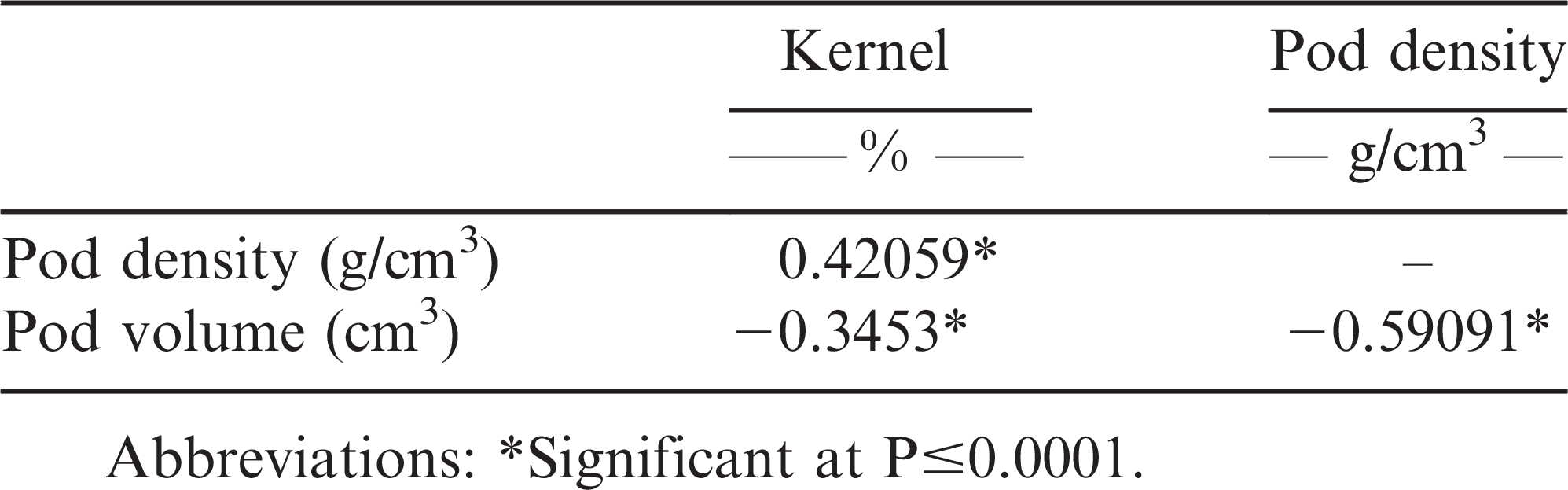
Data indicated that pod volume had statistically negative (P<0.0001) associations with both pod density (r = −0.59091) and percent kernel (r = −0.3453) (Table 3). Both negative and positive correlations between pod size and shelling percentage have been previously reported (de Godoy and Norden, 1981; Venuprasad et al., 2011). It is suspected that the large pod size and thicker pericarp traits in peanut may be linked, thus making it difficult for breeders to select for larger pods with higher shelling outturns (de Godoy and Norden, 1981; Hammons, 1973; Venuprasad et al., 2011).
The present study was undertaken in order to devise an objective, reliable, and a relatively quick method for measuring pod volume in peanut. While the goal of increased objectivity was accomplished by incorporating an imaging system, reliability largely depends on how well the various potential confounding factors are controlled during the experiment.
Aydin et al. (2007) reported that an increase in moisture content may increase the projected pod area and pod density, among other physical properties. Therefore, in order to accurately measure pod volume and density using the proposed method, it is imperative that all entries be dried to the same moisture content before starting the experiment. In addition, the effectiveness of the proposed regression model in predicting pod volume can be validated by carrying out another experiment where the pod volume is measured by using the liquid displacement, as well as from the projected pod area using image acquisition. Since kernel maturity may influence pod density, representative samples from each accession should belong to the same maturity class.
While the proposed pod volume measurement method has most of the characteristics of an ideal phenotyping method as enumerated in Janila et al. (2013), it still falls short due to its destructive nature of sample processing. However, it provides a more accurate method than the liquid displacement method, which is time consuming and prone to erroneous pod volume measurement owing to the liquid absorption during the process.
Testa Color
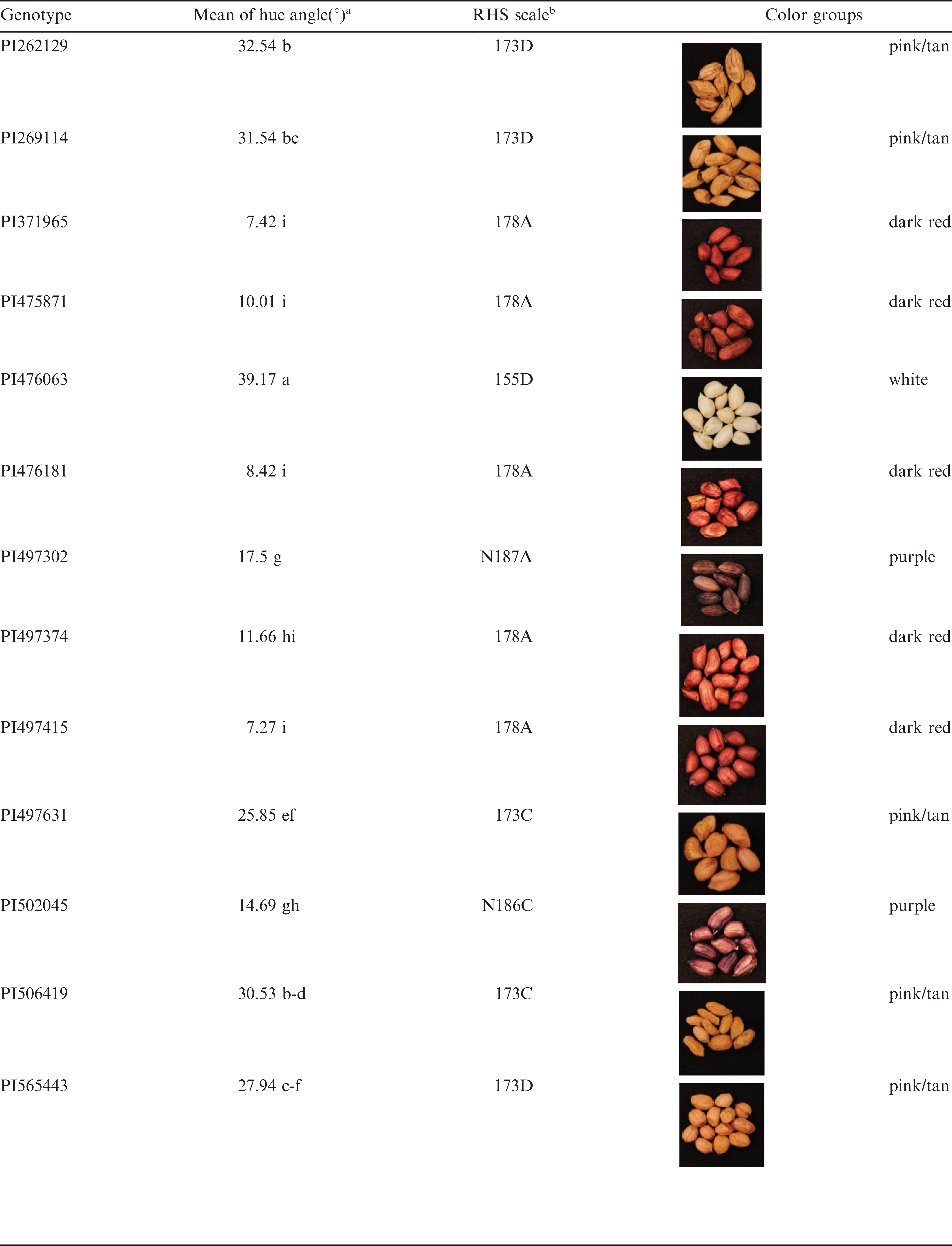
The five basic solid testa colors for peanut, including white, tan, pink, red, and purple (Branch, 2011), were represented in the diverse set of 28 accessions used in the present study (Table 4). To avoid ambiguity, five accessions, that had variegated testa color, were excluded from evaluation.
As previously stated, hue angle was measured to reflect the testa color differences among various entries. Visually, sample testa color ranged from dark red, purple, pink, tan and white as the hue angle value increased, and statistically significant differences were obtained among these color groups by hue angle measurement (Table 4). Hue angle analysis matched well with RHS scale except for the pink and tan group. The RHS visual scale does not match 100% with hue angle measurement due to the subtle color differences within this group. The smallest (7.27±0.1836°) and largest (39.17±0.2374°) mean hue angle values among 23 entries were observed for PI497415 (dark red) and PI476063 (white), respectively. Previously, testa color was categorized into arbitrary groups by visual selection (Holbrook et al. 1993). Current digitized image analysis of testa color provides an objective alternative method for testa color classification. It is important to note that hue is one of the main attributes of color. Therefore, hue angle represents one dimension or property of the color, and not the testa color itself. Nevertheless, hue angle measurement was shown to be suitable to distinguish among peanut germplasm accessions according to testa color variation, although intra-sample variation confounded the distinction between pink and tan. Chukwumah et al. (2009) observed a strong association between hue angle of the peanut testa and total polyphenol content of the kernel, and hypothesized that hue angle may act as a biomarker of total polyphenol content. In the process of shelling, dark testa color can negatively affect the color sorting efficiency since damaged seeds from mold, heat or moisture are discolored (Davidson et al., 1982).
In addition, since measurements from the current study are performed on acquired images of the peanut pod hulls and kernels, an image repository will be developed over time. These images can be used for comparison and/or reference in additional pod and seed phenotyping studies. Furthermore, future developments in image analysis technology may allow for extracting more phenotypic information from such image archives.
Acknowledgements
This work was supported by the National Peanut Foundation and Georgia Peanut Commission.
Literature Cited
Agrawal K Clary B. and Schroeder E. 1973 Mathematical models of peanut pod geometry Trans. ASAE 16 : 315 – 319 .
Akcali I Ince A. and Guzel E. 2006 Selected physical properties of peanuts Intl. J. Food Prop. 9 : 25 – 37 .
Aydin C. 2007 Some engineering properties of peanut and kernel J. Food Engineer. 79 : 810 – 816 .
Boldor D Sanders T. Swartzel K. and Simunovic J. 2002 Computer-assisted color classification of peanut pods Peanut Sci. 29 : 41 – 46 .
Branch W.D. 2011 First 100 years - Inheritance of testa color in peanut (Arachis hypogaea L.) Crop Sci. 51 : 1 – 4 .
Chu Y Wu C.L. Holbrook C.C. Tillman B.L. Person G. and Ozias-Akins P. 2011 Marker-assisted selection to pyramid nematode resistance and the high oleic trait in peanut Plant Genome 4 : 110 – 117 .
Chukwumah Y Walker L.T. and Verghese M. 2009 Peanut skin color: A biomarker for total polyphenolic content and antioxidative capacities of peanut cultivars Intl. J. Mol. Sci. 10 : 4941 – 4952 .
CIE Hue. International Commission on Illumination, "http://eilv.cie.co.at/term/542" (accessed 13 October 2014) .
Colvin B.C Rowland D.L Ferrell J.A. and Faircloth W.H. 2014 Development of a digital analysis system to evalute peanut maturity Peanut Sci. 41 : 8 – 16 .
Davidson J.I Whitaker T.B. and Dickens J.W. 1982 Grading, cleaning, storage, shelling, and marketing of peanuts in the United States Yoakum, TX: Am. Peanut Res. Edu. Soc.: 571 – 623 .
de Godoy I.J. and Norden A. 1981 Shell and seed size relationships in peanuts Peanut Sci. 8 : 21 – 24 .
Dowell F.E. 1992 Automatic control of a peanut grade sample inspection system Food Contr. 3 : 105 – 108 .
Ghate S Evans M. Kvien C. and Rucker K. 1993 Maturity detection in peanuts (Arachis hypogaea L.) using machine vision Trans. ASAE 36 : 1941 – 1947 .
Gilman D. and Smith O. 1977 Internal pericarp color as a subjective maturity index for peanut breeding Peanut Sci. 4 : 67 – 70 .
Hammons R.O. 1973 Genetics of Arachis hypogaea Peanuts: Culture and uses. Amer. Peanut Res. and Educ. Assoc.: 135 – 173 .
Holbrook C.C. and Branch W.D. 1989 Additional locus with a recessive allele for red testa color in peanut Crop Sci. 29 : 312 – 314 .
Holbrook C.C Anderson W.F. and Pittman R.N. 1993 Selection of a core collection from the U.S. germplasm collection of peanut Crop Sci. 33 : 859 – 861 .
Janila P Nigam S. Pandey M.K. Nagesh P. and Varshney R.K. 2013 Groundnut improvement: Use of genetic and genomic tools Front. Plant Sci. 4 : 23 .
Knauft D. and Gorbet D. 1993 Consistency of rank correlations of peanut breeding lines for market grade characteristics and yield Crop Sci. 33 : 697 – 699 .
National Institutes of Health ImageJ , http://imagej.nih.gov/ij (accessed 11 Jun 2015) .
Rucker K Kvien C. Calhoun K. Henning R. Koehler P. Ghate S. and Holbrook C. 1994 Sorting peanuts by pod density to improve quality and kernel maturity distribution and to reduce aflatoxin Peanut Sci. 21 : 147 – 152 .
Van der Stok J. 1910 Resultaten Van Kruisingsproeven Bij Arachis hypogaea L.(Katjang-Tanah) Onderzoekingen omtrent rijst en tweede ge wassen.(In Dutch.) Med. Uitg. van het Dep. v. Landbouw 12 : 218 – 221 .
Venuprasad R Aruna R. and Nigam S. 2011 Inheritance of traits associated with seed size in groundnut (Arachis hypogaea L.) Euphytica 181 : 169 – 177 .
Notes
- First, second, third, and fifth authors: Research Technician, Graduate Student, Research Professional, and Professor, Institute of Plant Breeding, Genetics & Genomics, University of Georgia Tifton Campus, Tifton, GA 31793; Fourth author: Research Geneticist, USDA-ARS, Coastal Plain Experiment Station, Tifton, GA 31793 *Corresponding author’s E-mail: pozias@uga.edu
Author Affiliations