Introduction
Real-time quantitative PCR (qPCR) is currently one of the more powerful techniques available for analyzing gene expression levels in tissues derived from living organisms. In comparison to classic transcript analysis tools, such as Northern blotting, RNase protection analysis, in situ hybridization, and semi-quantitative RT-PCR (RT-PCR), qPCR reactions are more sensitive and more specific, have broader quantification ranges, and are easy to perform (Ginzinger, 2002; Kubista et al., 2006). In order to maximize the accuracy of qPCR assays, several steps related to the quality of the RNA samples, the RNA input used in the reverse-transcription reaction, and the enzymatic efficiencies in the reverse transcription reaction should be taken into consideration. Above all, the most important concern is the need for appropriate normalization of the data with a suitable reference gene to overcome differences between tissues, cells, and treatments (Vandesompele et al., 2002).
It has been shown that real-time RT-PCR results are highly dependent on the reference genes chosen (e.g. Dheda et al., 2005). Useful reference genes must be present in all tested samples and should be relatively highly expressed (Pfaffl et al., 2004). In addition, the expression levels of a reference gene need to remain constant relative to experimental variables such as treatments, organs and experimental pressures introduced; otherwise, it may lead to invalid results (Suzuki et al., 2000; Bustin, 2002; Bustin and Nolan, 2004). The reference genes that are usually used for this purpose are “housekeeping genes”, which are involved in basic cellular processes, and supposed to have constant levels of expression across different treatments, organs, and developmental stages. Previously, these have been found to be reasonable internal reference genes for normalizing real-time data (e.g. Remans et al., 2008). In Arachis, for example, the ubiquitin gene is sometimes used in qPCR studies as a stable housekeeping gene (e.g., Luo et al., 2005; Nobile et al., 2007). However, several reports have shown that the expression levels of these so-called housekeeping genes differ among different plant tissues and organs (Jain et al., 2006; Remans et al., 2008; Barsalobres-Cavallari et al., 2009). Furthermore, normalization to a single housekeeping gene can lead to invalid normalization, because many housekeeping genes, are not only important for basal cell metabolism, but also participate in other cellular functions that may vary under changing conditions (Singh and Green, 1993). Consequently, some of these genes are unsuitable for use as transcriptional inner controls, and using them for the normalization of qPCR data corresponding to different tissues may lead to significant experimental errors that could result in the inappropriate interpretation of biological data (e.g. Schmittgen et al., 2000).
In recognition of the importance of reference genes for the normalization of qPCR data, the stability of various housekeeping genes has been evaluated under specific conditions in different organisms. In plants, a growing number of studies have evaluated reference genes in detail, including studies in rice (Jain et al., 2006; Mukesh et al., 2006), poplar (Brunner et al., 2004), potato (Nicot et al., 2005), soybean (Jian et al., 2008), Arabidopsis (Remans et al., 2008), grapevine (Reid et al., 2006), and Coffea arabica (Barsalobres-Cavallari et al., 2009). The search for the most stable reference gene usually involves scoring the stability of a series of 8–15 candidate genes for a set of different tissues and/or conditions. An alternative approach is to seek out genes that show little variation across a series of many microarray experiments resembling many tissue types and/or experimental conditions (Hovav et al., 2008; Libault et al., 2008). To date, appropriate internal controls have not been identified for gene expression studies in Arachis hypogaea.
In this study, we used real-time qRT-PCR to examine the expression of 10 housekeeping genes in a variety of peanut tissues, as well as in peanut kernels collected at different developmental stages. The goal of the study was to detect changes associated with kernel development for several oil regulatory genes. Therefore, an assay was undertaken to compare the stability of several potential control genes. Following the current literature (Jian et al., 2008; Remans et al., 2008; Barsalobres-Cavallari et al., 2009), nine candidate reference genes, namely alcohol dehydrogenase (adh3), polyubiquitin10 (ubq10), 60s, ubiquitin c (ubc), 14-3-3, β-actin (act7), glyceraldehyde-3-phosphate dehydrogenase (gapdh), yellow leaf specific (yls8), and elongation factor 1α (ef-1α) were selected. In addition, another gene coding for RNA helicase 1 (hel1), which was previously found to be stable in a cotton qPCR system (Hovav et al., 2008), was included in this analysis. These potential reference genes were ranked according to their expression profiles and stability in our experimental system using the GeNorm, NormFinder, and BestKeeper software programs. Different degrees of variation in the expression of the examined genes in the different tissue samples were found. Integrating the results from all three assays, we found the combination of adh3 and yls8 to be the most stable. The results were validated using unpooled tissue samples from five peanut kernel developmental stages and the effects of using one or more reference genes on the relative expression levels of a diacylglycerol acyltransferase gene (Dgat1) during kernel development are demonstrated.
Materials and Methods
Plant Material and Growing Conditions
Samples of seeds at different developmental stages, as well as five different tissue types (full pod, mature seed, leaf, gynophore, and root) were obtained from four-month-old Virginia-type peanut plants (Arachis hypogea var. Shulamit) grown under greenhouse stable conditions (25–30 C, 30% RH; 14 h photoperiod with fluorescent light of intensity 35 µmol m−2 s−1) in Bet-Dagan, Israel. Three replications of three plants each were sampled and bulked, and the fresh tissue samples were frozen immediately in liquid nitrogen until RNA extraction.
RNA Isolation and Quality Controls
Total RNA was extracted using the hot borate (sodium borate decahydrate) method, as previously described for cotton ovules by Hovav et al. (2008) with some adjustments. Tissue samples of 0.25 to 1.2 g were weighed and ground to fine powder in liquid nitrogen using a pre-cooled mortar and pestle. The frozen ground samples were combined with 8 mL of borate buffer (0.2 M sodium borate decahydrate; 30 mM EGTA; 1% (w/v) SDS; 1% sodium deoxycholic acid; 10 mM DTT; 1% IGEPAL CA-630 (Nonidet P-40, NP-40); 2% (w/v) PVP-40) at 65 C and ground thoroughly until the suspension was evenly dispersed. The homogenate was transferred to a 50 mL tube and incubated for 1.5 hours in a 42 C incubator/shaker. The samples were supplemented with 1 mL of 2 M potassium chloride and placed on ice for 1 hour.
Subsequent to centrifugation, the supernatant of each sample was transferred to a 50 mL tube containing 8 M lithium chloride (for a final concentration of 2 M) and incubated on ice overnight. Following a second centrifugation, the supernatant was discarded and the pellet was washed with 1.5 mL of 2 M lithium chloride. This was repeated two to three times. The pellet was then suspended in 250 µL 1× TE and warmed to room temperature. Each sample was centrifuged again and the supernatant was transferred to a 1.5-mL tube containing 2 M potassium acetate and incubated on ice for 15 min. After an additional centrifugation, the pellet was discarded and the supernatant was transferred to a tube containing 3 M sodium acetate and 2.5× cold 100% ethanol. The samples were stored at −80 C for 1–2 h. Following centrifugation, the supernatant was discarded and the pellet was washed with 1 mL of 70% ethanol. After an additional centrifugation, the ethanol was discarded and the RNA was resuspended in 100 µL of DEPC-treated water and stored at −80 C. Only RNA samples with a 260/280 ratio between 1.9 and 2.1 and a 260/230 ratio greater than 2.0 were used for subsequent analyses. The integrity of the RNA samples was also assessed using 1.0% agarose/formaldehyde gel electrophoresis.
Reverse Transcription
From each sample, two micrograms of total RNA were treated with RNase-free DNase (Promega, Madison, WI). Treated RNA was reverse-transcribed using the AccuPower RT PreMix kit for RT-PCR (Bioneer, Alameda, CA) with oligo-(dT)-15 primer and hexameric primer (both from Promega). The concentration of cDNA for each sample was determined using the Nano-Drop ND-1000 spectrophotometer (NanoDrop Technologies, Wilmington, DE).
Primer Design
Primers were designed for Arachis hypogea based on commonly used housekeeping genes representing distinct functional classes, identified by BLAST searches in the Arachis hypogaea EST database available at the Legume Information System website (http://www.comparative-legumes.org/index.php/Home) and at the GeneBank (http://www.ncbi.nlm.nih.gov/Genbank/). The name, descriptor, primer sequence, primers efficiency and amplicon length of each reference gene are presented in Table 1. Each of the primer pairs produced a single product and amplified the target transcript with efficiency values ranging between 0.89 and 1.04 (Table 1) over a 1000-fold range of input material.
Quantitative PCR
The PCR mixture contained 3 µL of a 1∶100 dilution of the synthesized cDNA (equivalent to 20 ng input RNA), primers to a final concentration of 0.4 µM each, 5 µL of the SYBR Green PCR Master Mix (Takara, Saint-Germain-en-Laye, France), and PCR-grade water to a total volume of 10 µL. Each reaction was performed in triplicate. PCR reactions were also performed in the absence of template; these reactions served as negative controls for each primer pair. Equimolar pools of cDNA samples of five Arachis tissue types (full pod, mature seed, leaf, gynophore and root), as well as five seed developmental stages, were prepared to be used as a common reference for all qPCR analyses. The quantitative PCRs were performed in the Rotor Gene 6000 Real-Time PCR cycler (Qiagen, Valencia, CA). All PCR reactions were performed in a 72-well rotor (Qiagen, Valencia, CA) under the following conditions: 10 min at 95 C, 40 cycles of 5 s at 95 C, 15 s at 55 C, 10 s at 60 C, and 20 s at 72 C. Confirmation of amplicon specificity was based on the dissociation curve produced at the end of each run and on visualization of the products following electrophoresis on an 8% polyacrylamide gel.
Analysis of Candidate Reference Genes
The BestKeeper descriptive statistical method (Pfaffl et al., 2004) was used to estimate the variation in the expression levels of the 10 candidate genes over a set of five peanut tissue/organ samples. To estimate the variance of expression in the system, two different approaches were used. The first involved model-based variance estimation performed using a Visual Basic application for Microsoft Excel called NormFinder (http://www.mdl.dk/publicationsnormfinder.htm). This approach enables estimation not only of the overall variation of the candidate normalization genes, but also of the variation between subgroups of the sample set (Andersen et al., 2004). It provides a direct measure for the estimated expression variation, enabling the user to evaluate the systematic error introduced when the gene is used.
The second approach used the GeNorm software program (http://medgen.ugent.be/∼jvdesomp/genorm). This method is based on the fact that the expression ratio of two ideal internal control genes is identical in all samples and the variation of the expression ratios of two “real” housekeeping genes reflects the fact that one (or both) is/are not constantly expressed, with increasing variation in the ratio corresponding to decreasing expression stability. The pair-wise variation for each control gene as compared to each of the other control genes is determined, and the internal control gene-stability measure M is defined as the average pair-wise variation of a particular gene with all of the other control genes (Vandesompele et al., 2002). Genes with the lowest M values have the most stable expression.
Results and Discussion
Descriptive Statistics of the Reference Candidate Genes
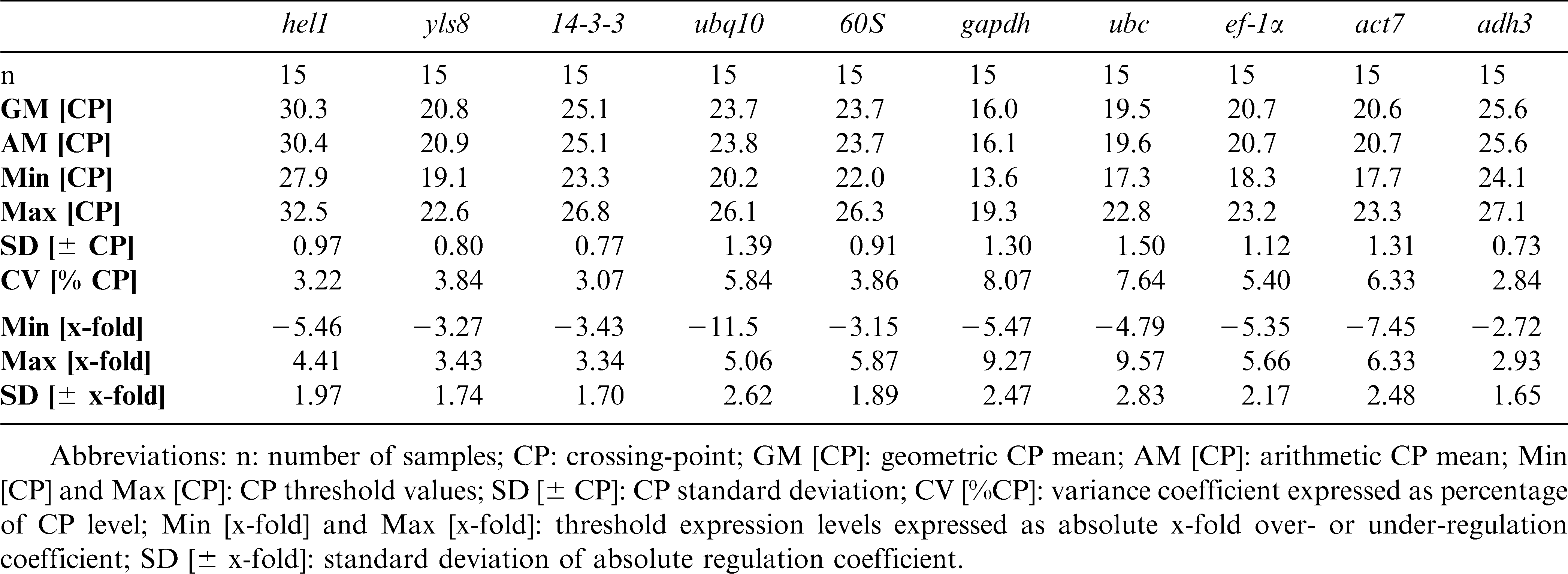
The expression profiles of 10 candidate reference genes (hel1, yls8, 14-3-3, ubq10, 60S. gapdh, ubc, ef-1α, act7, and adh3) were initially assessed for a group of five peanut organ samples (full pod, mature seed, leaf, gynophore, and root) using qPCR. Descriptive statistics of the derived crossing points (CP) were calculated using the BestKeeper program (http://www.gene-quantification.de/bestkeeper.html). This program examines the level of variation in the expression of each candidate gene, as described by Pfaffl et al. (2004).
According to this analysis, the gene with the lowest level of expression across all 15 samples (five tissues × three replications) was hel1 (RNA helicase 1), for which CP values were obtained around cycles 28–33 (Table 2). The gene with the highest level of expression was gapdh, whose CP values were obtained around cycles 14–19. The expression variation levels of the entire gene set could be divided into two groups, while adh3, 60S, 14-3-3, hel1 and yls8 showed ± 1.79 x-fold (1.65 x-fold < SD < 1.97 x-fold) variation, the expression of ubq10, act7, gapdh, ubc, and ef-1α exhibited higher ranges of CP variation (SD = ± 2.5 cycles). The coefficient of variation (CV) for the overall assay was 5.35% (total assay variability), which is within the range (from 3.4% to 11.6%) of values reported in previously for qPCR (Pfaffl et al., 2004). Overall, the X-fold variance values in this study are somewhat higher than the values that have been observed in other plant systems (e.g., Mukesh et al., 2006; Remans et al., 2008, Barsalobres-Cavallari et al., 2009). Besides laboratory working errors these higher values may indicate high levels of variability across peanut organs and tissues, and this variability may be partially explained by the fact that many housekeeping genes, like gapdh, are not only important for basal cell metabolism, but also participate in other cellular functions (Singh and Green, 1993; Ishitani et al., 1996). This notion corresponds well with the results of other qRT-PCR normalization assays in plants that have shown similar ranges of variance (e.g., Nicot et al., 2005; Jian et al., 2008).
Ranking the Candidate Reference Genes
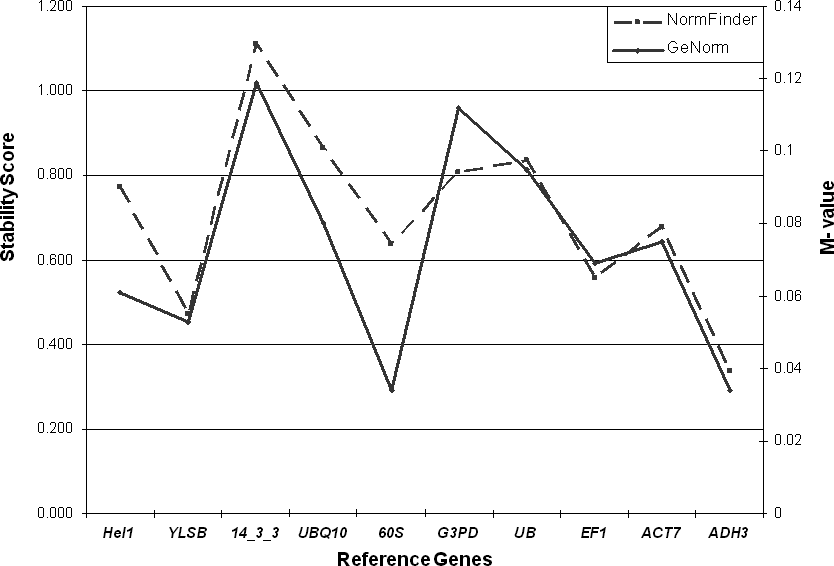
Two different methods, the NormFinder program (http://www.mdl.dk/publicationsnormfinder.htm) and the GeNorm program (http://medgen.ugent.be/∼jvdesomp/genorm), were used to estimate the variance of expression in our system. As shown in Figure 1, both methods for estimating stability produced parallel results. Overall, the gene with most stable expression was adh3, which had minimal estimated intra- and inter-tissue variation (stability = 0.334) according to NormFinder, and the lowest M value as calculated using GeNorm, alongside with 60s (M = 0.041). The gene with the least stable expression was 14-3-3 (stability = 1.11; M value = 0.124), followed by ubq10 (according to NormFinder, stability = 0.865) and gapdh (according to GeNorm, M value = 0.115). Interestingly, the ubq10 (ubiquitin) gene, which has been used in several qPCR studies in Arachis (Luo et al., 2005; Nobile et al., 2007), showed quite a low level of stability in this study.
Another function of the NormFinder program allows the user to test the best combination of two genes. Based on this analysis, the best gene combination is yls8 and adh3; this combination had a stability value of 0.261(data not shown). In the GeNorm program, that ranks reference genes from the least stable down to the two most stable genes (and beyond that the best two cannot be discerned), adh3 and 60s were found to be the most stable. Pairwise variation analyses from the geNorm program, which evaluates the optimal number of reference genes to be used in a study, showed that the first variation value (V2/V3) was under the 0.15 suggested value (Vandesompele et al., 2002). This means that the inclusion of additional reference genes besides adh3 and 60s is not required in this study. Based on these results, the use of the combination of yls8 with adh3 or 60s with adh3 is recommended in qPCR expression studies in Arachis.
Validation of Results in Peanut Kernels at Different Developmental Stages
An additional step in the validation of the expression levels of two highly stable (adh3, yls8) and two less stable (14-3-3 and ubq10) reference genes was performed using unpooled tissue samples from five peanut-kernel developmental stages. Developmental stages were defined according to the established classification system (Boote, 1982), with R4, R5, R6, R7, and R8 representing the ovary from full pod, beginning seed, full seed, beginning maturity, and harvest maturity developmental stages, respectively.
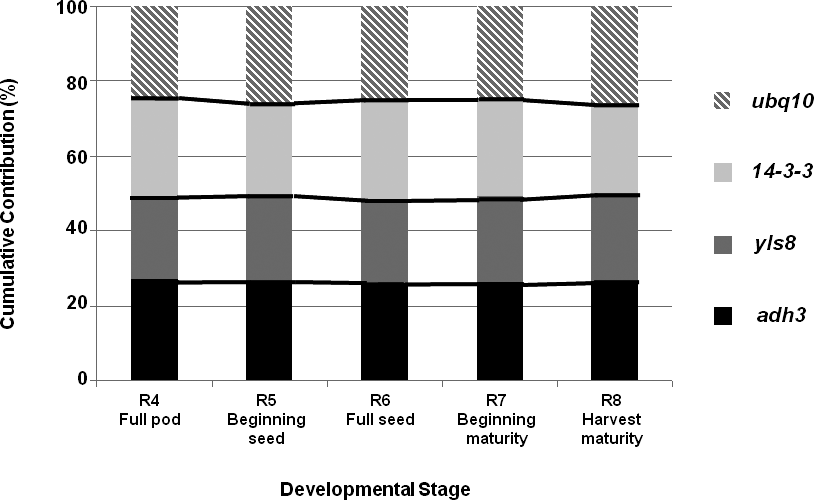
In this analysis, the most highly expressed gene was yls8 (21 cycles), followed by ubq10 (24 cycles); while 14-3-3 and adh3 presented the same mean CP (25 cycles; data not shown). A comparison of gene contributions is presented in Figure 2. As observed earlier, adh3 had the most stable expression across all of the examined peanut-kernel developmental stages; while the highest levels of variation were observed for ubq10 and 14-3-3.
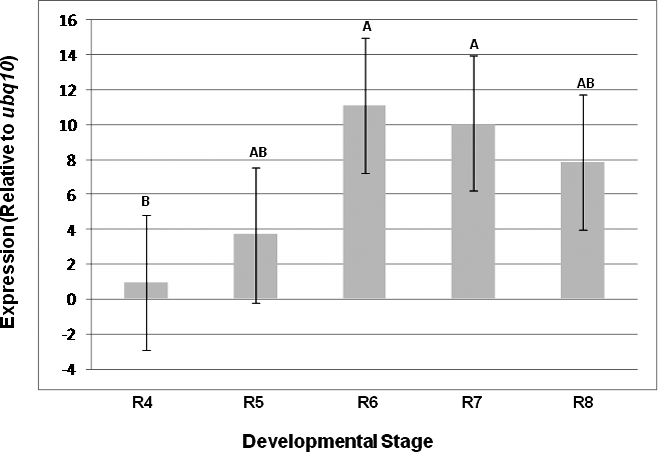
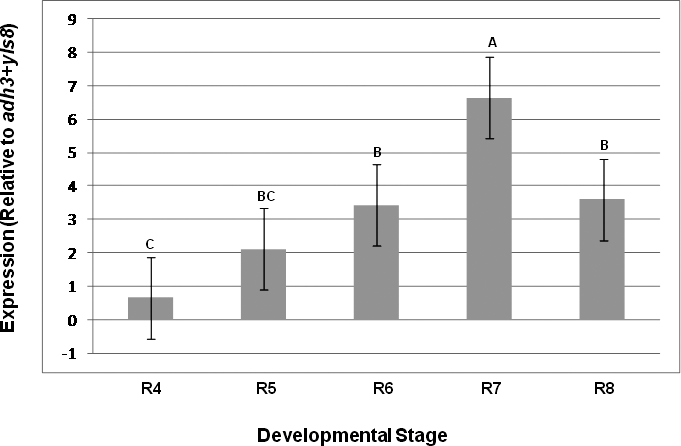
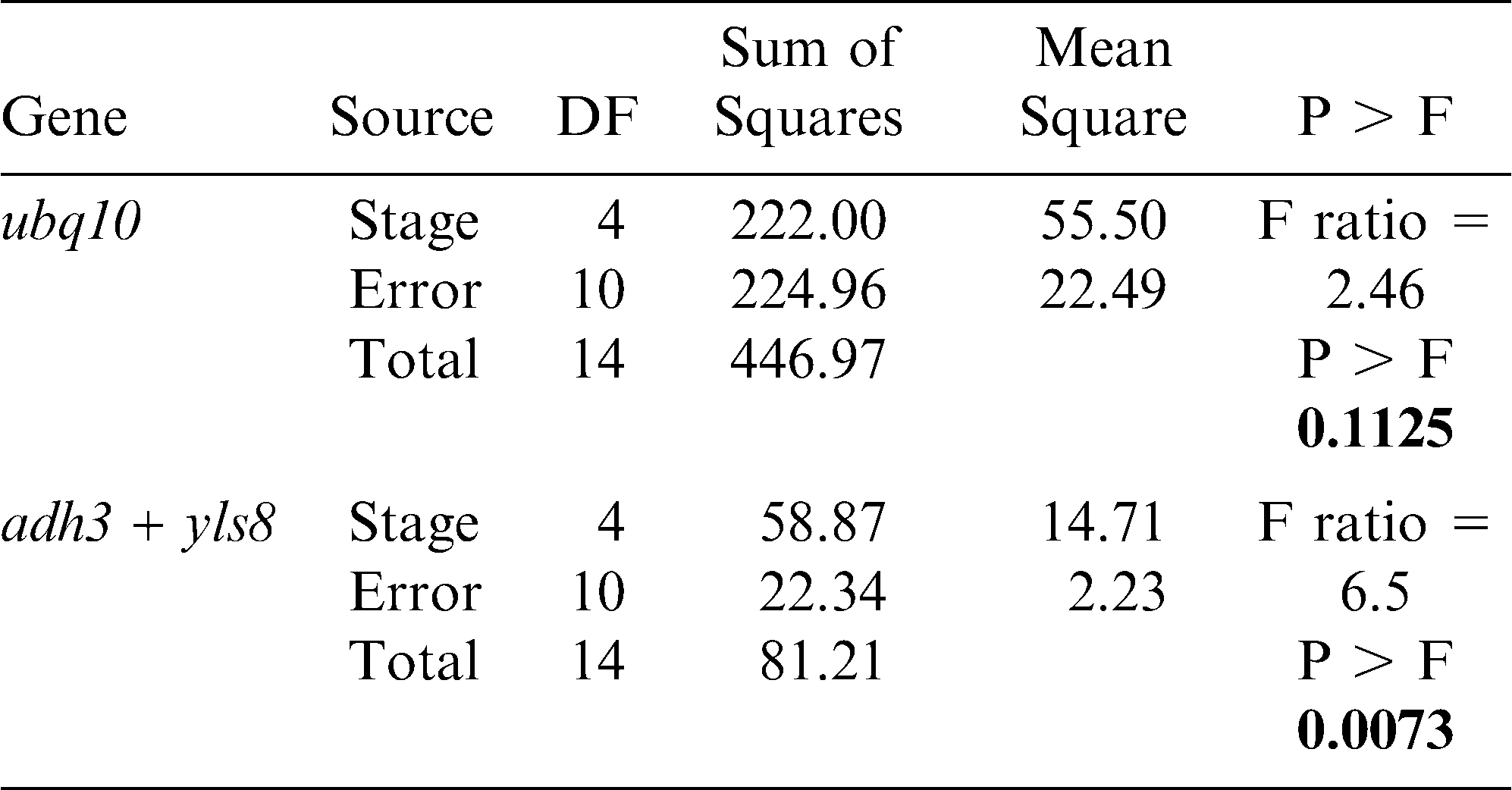
The importance of this assay was demonstrated by comparing the expression results of a seed oil-related gene (Dgat1) as evaluated using the yls8 + adh3 combination as the reference gene with the observed expression of this gene when ubq10 was used as the reference gene for normalization. Dgat1 encodes for a diacylglycerol acyltransferase (DGAT; EC 2.3.1.20) that is involved in the terminal step in TAG formation in plants (Lung and Weselake, 2006). The comparison was conducted by SAS program using a variance and least square means analysis of the CP differences (ΔCP) obtained from the assayed expression levels of the tested genes at different A. hypogaea seed developmental stages. The average CP (N = 3) was calculated for each gene and the ΔCP (CPdgat1 – CP reference gene) was determined for each stage. As shown in Table 3 and in Figures 3 and 4, the analysis of variance showed different outcomes for each trial. Using yls8 + adh3 as a reference gene combination resulted in observations of more significant differences between developmental stages (P > F = 0.0073), as compared to when ubq10 was used as the reference gene (P > F = 0.1125). Accordingly, the Student's t analysis of the differences between the least square means showed almost no significant differences in Dgat1 at the different developmental stages when ubq10 was used as the reference gene (Fig. 3). However, when the yls8 + adh3 combination was used as the reference gene, Dgat1 expression was significantly lower at stage R4 (full pod) than at R6, R7, and R8 (Figure 4). Also, when the yls8 + adh3 combination was used as the reference gene, Dgat1 expression at the R7 (beginning maturity) stage was significantly higher than it was at any of the other stages. The use of only adh3 as a reference gene produced results similar to those observed for the yls8 + adh3 combination (data not shown).
Quantitative real-time PCR analyses of the diacylglycerol acyltransferase 1 gene using ubq10 as reference gene. R4, R5, R6, R7, and R8 represent different developmental stages: ovary from full pod, beginning seed, full seed, beginning maturity, and harvest maturity, respectively. Significantly different Student's t-test groups are indicated as well.
Quantitative real-time PCR analyses of the diacylglycerol acyltransferase 1 gene using a combination of adh3 + yls8 as reference genes. R4, R5, R6, R7, and R8 represent different developmental stages: ovary from full pod, beginning seed, full seed, beginning maturity, and harvest maturity, respectively. Significantly different Student's t-test groups are indicated as well.
General Remarks about the Selected Reference Genes
According to the results of this study, the reference genes with the most stable expression were adh3, yls8 and 60s. Therefore, these genes should be considered suitable reference genes for Arachis, particularly in studies involving seed development. Different samples or treatments, though, may require the reevaluation of a suitable reference gene, since changing conditions can sometimes cause a suitable reference gene to become unstable (e.g. Remans et al., 2008). Adh3 encodes for alcohol dehydrogenase class III enzyme, which catalyzes the interconversion of alcohols and aldehydes or ketones with the reduction of NAD+ to NADH, and also plays an important role in lowering the toxicity of the cell (Yokoyama and Harry, 1993). Interestingly, adh3 is not frequently used as a qPCR reference gene in quantitative RT-PCR assays. Our observation, though, corroborates the results of the quantification of adh3 expression in different tissues of Coffea arabica, where it showed similar stability values, although this work concluded that gapdh was the most appropriate reference gene (Barsalobres-Cavallari et al., 2009). In the same study the 60s gene was also found to have acceptable low variation among tissue/organ types. 60s encodes for the L7 protein, a transcriptional regulator and structural constituent of the 60s subunit of the cytosolic ribosome. It was also reported to have minimum variation among Serial Analysis of Gene Expression (SAGE) libraries in sugarcane (Calsa et al., 2007). Yls8 encodes for the mitosis protein YLS8. While the actual function of this gene is not known, it was recently shown to be highly stable in a similar real-time stability assay conducted in Arabidopsis exposed to increased metal concentrations (Remans et al., 2008).
Another outcome of this analysis is that ubq10 and gapdh, two of the reference genes most commonly used in plants, which have been found to be stable in many qPCR systems, (e.g., Jain et al., 2006; Reid et al., 2006), were two of the most unstable genes in the current study. Also, assays conducted in rice (Jain et al., 2006) and potato (Nicot et al., 2005) showed ef-1α to be the most stable gene. This is not surprising and demonstrates the importance of empirically choosing a suitable reference gene for not only particular species, but also within the same species when different treatments/organs are chosen. This also emphasizes the need to work with more than one reference gene to avoid invalid normalizations. Interestingly, in one study conducted in soybean, a close relative of peanut, an examination of 10 reference genes in different tissues, at developmental stages, in plants exposed to different photoperiodic treatments, and at different times of day found these two reference genes to be the most unstable and undesirable for quantitative RT-PCR studies (Jian et al., 2008). Similarly, the findings of the current study demonstrate that ubq10 and gapdh should be avoided in quantitative mRNA expression analyses in peanut.
Summary and Conclusions
The aim of this study was to select suitable reference gene/s for quantitative RT-PCR in Arachis. The expression stabilities of 10 commonly used housekeeping genes were assessed in a set of five diverse peanut tissue samples, as well as unpooled samples from five peanut kernel developmental stages by using GeNorm and NormFinder programs. Overall, the gene with the most stable expression and the minimal estimated intra- and inter-tissue variation across all of the examined tissues and both programs was adh3, followed by 60s and yls8. The importance of the use of specific reference gene and the number of reference genes on the observed relative expression levels of the Dgat1 gene during kernel development was demonstrated as well. Based on findings, the suggestion is that adh3, or a combination of this gene with 60s and yls8 should be considered for use in quantitative mRNA expression analyses in Arachis, particularly in studies involving seed development; whereas ubq10 and gapdh should be avoided.
Literature Cited
Andersen C. L. , Jensen J. L. , and Orntoft T. F. 2004 Normalization of real-time quantitative reverse transcription-PCR data: A model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 64 : 5245 – 5250 .
Barsalobres-Cavallari C. F. , Severino F. E. , Maluf M. P. , and Maia I. G. 2009 Identification of suitable internal control genes for expression studies in Coffea arabica under different experimental conditions. BMC Mol. Biol 6 10 : 1 .
Boote K. G. 1982 Growth stages of peanuts (Arachis hypogaea L.). Peanut Sci 9 : 35 – 40 .
Brunner A. M. , Yakovlev I. A. , and Strauss S. H. 2004 Validating internal controls for quantitative plant gene expression studies. BMC Plant Biol l4 : 14 .
Bustin S. A. 2002 Quantification of mRNA using real-time reverse transcription PCR (RT-PCR): trends and problems. J. Mol. Endocrinol 29 : 23 – 39 .
Bustin S. A. and Nolan T. 2004 Pitfalls of quantitative real-time reverse-transcription polymerase chain reaction. J. Biomol. Tech 15 : 155 – 166 .
Calsa T. and Figueira A. 2007 Serial analysis of gene expression in sugarcane (Saccharum spp.) leaves revealed alternative C4 metabolism and putative antisense transcripts. Plant Mol. Biol 63 : 745 – 726 .
Dheda K. , Huggett J. F. , Chang J. S. , Kim L. U. , Bustin S. A. , Johnson M. A. , Rook G. A. W. , and Zumla A. 2005 The implications of using an inappropriate reference gene for real-time reverse transcription PCR data normalization. Anal. Biochem 344 : 141 – 143 .
Ginzinger D. G. 2002 Gene quantification using real-time quantitative PCR: An emerging technology hits the mainstream. Exp. Hematol 30 : 503 – 512 .
Hovav R. , Udall J. A. , Flagel L. , Rapp R. A. , Hovav E. , and Wendel J. F. 2008 A majority of cotton genes are expressed in single-celled fiber. Planta 227 : 319 – 329 .
Ishitani R. , Sunaga K. , Hirano A. , Saunders P. , Katsube N. , and Chuang D. M. 1996 Evidence that glyceraldehyde-3-phosphate dehydrogenase is involved in age-induced apoptosis in mature cerebellar neurons in culture. J. Neurochem 66 : 928 – 935 .
Jain M. , Nijhawan A. , Tyagi A. K. , and Khurana J. P. 2006 Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative real-time PCR. Biochem. Biophys. Res. Commun 345 : 646 – 651 .
Jian B. , Liu B. , Bi Y. , Hou W. , Wu C. , and Han T. 2008 Validation of internal control for gene expression study in soybean by quantitative real-time PCR. BMC Mol. Biol 9 : 59 .
Kubista M. , Andrade J. M. , Bengtsson M. , Forootan A. , Jonak J. , Lind K. , Sindelka R. , Sjoback R. , Sjogreen B. , and Strombom L. 2006 The real-time polymerase chain reaction. Mol. Aspects. Med 27 : 95 – 125 .
Libault M. , Thibivilliers S. , Bilgin D. D. , Radwan O. , Benitez M. , Clough S. J. , and Stacey G. 2008 Identification of four soybean reference genes for gene expression normalization. Plant Genome 1 : 44 – 54 .
Lung S. C. and Weselake R. J. 2006 Diacylglycerol acyltransferase: A key mediator of plant triacylglycerol synthesis. Lipids 41 : 1073 – 1088 .
Luo M. , Liang X. Q. , Dang P. , Holbrook C. C. , Bausher M. G. , Lee R. D. , and Guo B. Z. 2005 Microarray-based screening of differentially expressed genes in peanut in response to Aspergillus parasiticus infection and drought stress. Plant Sci 169 : 695 – 703 .
Mukesh J. , Aashima N. , Akhilesh K. T. , and Jitendra P. K. 2006 Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative real-time PCR. Biochem. Biophys. Res. Commun 345 : 646 – 651 .
Nicot N. , Hausman J. F. , Hoffmann L. , and Evers D. 2005 Housekeeping gene selection for real-time RT-PCR normalization in potato during biotic and abiotic stress. J. Exp. Bot 56 : 2907 – 2914 .
Nobile P. M. , Lopes C. R. , Barsalobres-Cavallari C. , Quecini V. , Coutinho L. L. , Hoshino A. A. , and Gimenes M. A. 2007 Peanut genes identified during initial phase of Cercosporidium personatum infection. Plant Sci 174 : 78 – 87 .
Papini-Terzi F. S. , Rocha F. R. , Vêncio R. Z. , Oliveira K. C. , Felix Jde M. , Vicentini R. , Rocha Cde S. , Simões A. C. , Ulian E. C. , di Mauro S. M. , da Silva A. M. , Pereira C. A. , Menossi M. , and Souza G. M. 2005 Transcription profiling of signal transduction-related genes in sugarcane tissues. DNA Res 12 : 27 – 38. doi: 10.1093/dnares/12.
Pfaffl M. W. , Tichopad A. , Prgomet C. , and Neuvians T. P. 2004 Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper - Excel-based tool using pair-wise correlations. Biotechnol. Lett 26 : 509 – 515.
Reid K. E. , Olsson N. , Schlosser J. , Peng F. , and Lund S. T. 2006 An optimized grapevine RNA isolation procedure and statistical determination of reference genes for real-time RT-PCR during berry development. BMC Plant Biol 14 6 : 27 .
Remans T. , Smeets K. , Opdenakker K. , Mathijsen D. , Vangronsveld J. , and Cuypers A. 2008 Normalisation of real-time RT-PCR gene expression measurements in Arabidopsis thaliana exposed to increased metal concentrations. Planta 227 : 1343 – 1349 .
Schmittgen T. D. and Zakrajsek B. A. 2000 Effect of experimental treatment on housekeeping gene expression: validation by real-time, quantitative RT-PCR. J. Biochem. Biophys. Methods 46 : 69 .
Singh R. and Green M. R. 1993 Sequence-specific binding of transfer RNA by glyceraldehyde-3-phosphate dehydrogenase. Science 259 : 365 – 368 .
Suzuki T. , Higgins P. J. , and Crawford D. R. 2000 Control selection for RNA quantitation. BioTechniques 29 : 332 – 337 .
Vandesompele J. , De Preter K. , Pattyn F. , Poppe B. , Van Roy N. , De Paepe A. , and Speleman F. 2002 Accurate normalization of real-time quantitative RT-PCR by geometric averaging of multiple internal control genes. Genome Biol 3 : 34 .
Yokoyama S. and Harry D. E. 1993 Molecular phylogeny and evolutionary rates of alcohol dehydrogenases in vertebrates and plants. Mol. Biol. Evol 10 : 1215 – 1226 .
Notes
Author Affiliations
1 Dept. of Crop Science and Natural Resources, Volcani Center, ARO, Bet-Dagan, 50250, Israel.
*Corresponding author (email: ranh@agri.gov.il)